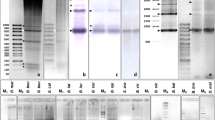
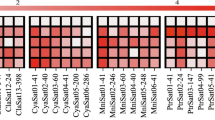
Abstract
The composition and orientation of the house mouse satellite DNA sequences (minor, major, TLC) were investigated by a FISH and CO-FISH approach in 11 taxa belonging to three clades of the subgenus Mus. Using a phylogenetic framework, our results highlighted two distribution patterns. The TLC satellite, the most recently discovered satellite, was present in all clades but varied quantitatively among species. This distribution supported its appearance in the ancestor of the subgenus followed by independent evolution in species of each clade. In contrast, the minor and major satellites occurred in only two clades of the subgenus indicating the simultaneous and recent amplification of these sequences. In addition, although qualitative differences in the composition and orientation of the satellite sequences were observed among the taxa, none of the features studied were unique to the house mouse and could account for the extensive chromosomal plasticity evidenced in Mus musculus domesticus.




Similar content being viewed by others
References
Acosta M, Marchal J, Fernández-Espartero C, Romero-Fernández I, Rovatsos M, Giagia-Athanasopoulou E, Gornung E, Castiglia R, Sánchez A (2010) Characterization of the satellite DNA Msat-160 from species of Terricola (Microtus) and Arvicola (Rodentia, Arvicolinae). Genetica 138(9):1085–1098
Adega F, Guedes-Pinto H, Chaves R (2009) Satellite DNA in the karyotype evolution of domestic animals—clinical considerations. Cytogenet Genome Res 126:12–20
Bandyopadhyay R, Berend SA, Page SL, Choo KHA, Shaffer LG (2001) Satellite III sequences on 14p and their relevance to Robertsonian translocation formation. Chromosom Res 9(3):235–242
Boursot P, Jacquart T, Bonhomme F, Britton-Davidian J, Thaler L (1985) Différenciation géographique du génome mitochondrial chez Mus spretus Lataste. C R Acad Sci 301:161–165
Brown JD, Piccuillo V, O'Neill RJ (2012) Retroelement demethylation associated with abnormal placentation in Mus musculus × Mus caroli hybrids. Biol Reprod 86:88
Carbone L, Harris RA, Vessere GM, Mootnick AR, Humphray S, Rogers J, Kim SK, Wall JD, Martin D, Jurka J, Milosavljevic A, de Jong PJ (2009) Evolutionary breakpoints in the gibbon suggest association between cytosine methylation and karyotype evolution. PLoS Genet 5(6):e1000538
Cazaux B, Catalan J, Veyrunes F, Douzery EJP, Britton-Davidian J (2011) Are ribosomal DNA clusters rearrangement hotspots? A case study in the genus Mus (Rodentia, Muridae). BMC Evol Biol 11:124
Chakrabarti S, Chakrabarti A (1977) Spontaneous Robertsonian fusion leading to karyotype variation in the house mouse—first report from Asia. Experientia 33:175–177
Chaves R, Adega F, Heslop-Harrison J, Guedes-Pinto H, Wienberg J (2003a) Complex satellite DNA reshuffling in the polymorphic t(1;29) Robertsonian translocation and evolutionarily derived chromosomes in cattle. Chromosom Res 11(7):641–648
Chaves R, Adega F, Wienberg J, Guedes-Pinto H, Heslop-Harrison JS (2003b) Molecular cytogenetic analysis and centromeric satellite organization of a novel 8; 11 translocation in sheep: a possible intermediate in biarmed chromosome evolution. Mamm Genome 14(10):706–710
Chevret P, Jenkins P, Catzeflis F (2003) Evolutionary systematics of the indian mouse Mus famulus Bonhote, 1898: molecular (DNA/DNA hybridization and 12S rRNA sequence) and morphological evidence. Zool J Linnean Soc 137:385–401
Chevret P, Veyrunes F, Britton-Davidian J (2005) Molecular phylogeny of the genus Mus (Rodentia: Murinae) based on mitochondrial and nuclear data. Biol J Linn Soc 84:417–427
Coghlan A, Eichler EE, Olivier SG, Paterson AH, Stein L (2005) Chromosome evolution in eukaryotes: a multi-kingdom perspective. Trends Genet 21:673–682
Cowell JK (1984) A photographic representation of the variability of G-banded structure of the chromosomes of the mouse karyotype. Chromosoma 89:294–320
Cucchi T, Orth A, Auffray J-C, Renaud S, Fabre L, Catalan J, Hadjisterkotis E, Bonhomme F, Vigne J-D (2006) A new endemic species of the subgenus Mus (Rodentia, Mammalia) on the island of Cyprus. Zootaxa 1241:1–36
Di Meo G, Perucatti A, Chaves R, Adega F, De Lorenzi L, Molteni L, De Giovanni A, Incarnato D, Guedes-Pinto H, Eggen A, Iannuzzi L (2006) Cattle rob(1;29) originating from complex chromosome rearrangements as revealed by both banding and FISH-mapping techniques. Chromosom Res 14(6):649–655
Dobigny G, Waters PD, Robinson TJ (2006) Absence of hypomethylation and LINE-1 amplification in a white × black rhinoceros hybrid. Genetica 127:81–86
Dod B, Mottez E, Desmarais E, Bonhomme F, Roizes G (1989) Concerted evolution of light satellite DNA in genus Mus implies amplification and homogenization of large blocks of repeats. Mol Biol Evol 6:478–491
Eichler EE, Sankoff D (2003) Structural dynamics of eukaryotic chromosome evolution. Science 301:793–797
Ellingsen A, Slamovits CH, Rossi MS (2007) Sequence evolution of the major satellite DNA of the genus Ctenomys (octodontidae, Rodentia). Gene 392:283–290
Falconer E, Chavez E, Henderson A, Lansdorp P (2010) Chromosome orientation fluorescence in situ hybridization to study sister chromatid segregation in vivo. Nat Protoc 5:1362–1377
Farre M, Montserrat B, Lopez-Giraldez F, Ponsa M, Ruiz-Herrera A (2011) Assessing the role of tandem repeats in shaping the genomic architecture of great apes. PLoS One 6:e27239
Frazer KA, Eskin E, Kang HM, Bogue MA, Hinds DA, Beilharz EJ, Gupta RV, Montgomery J, Morenzoni MM, Nilsen GB, Pethiyagoda CL, Stuve LL, Johnson FM, Daly MJ, Wade CM, Cox DR (2007) A sequence-based variation map of 8.27 million SNPs in inbred mouse strains. Nature 448:1050–1056
Froenicke L, Lyons AL (2008) Hotspots of mammalian chromosome evolution. In: eLS. John Wiley & Sons Ltd, Chichester. http://www.els.net. doi:10.1002/9780470015902.a0020750
Garagna S, Redi C, Capanna E, Andayani N, Alfano RM, Doi P, Viale G (1993) Genome distribution, chromosomal allocation, and organisation of the major and minor satellite DNAs in 11 species and subspecies of genus Mus. Cytogenet Cell Genet 64:247–255
Garagna S, Broccoli D, Redi C, Searle JB, Cooke HJ, Capanna E (1995) Robertsonian metacentrics of the house mouse lose telomeric sequences but retain some minor satellite DNA in the pereicentromeric area. Chromosoma 103:685–692
Garagna S, Marziliano N, Zuccotti M, Searle JB, Capanna E, Redi CA (2001) Pericentromeric organization at the fusion point of mouse Robertsonian translocation chromosomes. Proc Natl Acad Sci U S A 98(1):171–175
Gauthier P, Hima K, Dobigny G (2010) Robertsonian fusions, pericentromeric repeat organization and evolution: a case study within a highly polymorphic rodent species, Gerbillus nigeriae. Chromosom Res 18:473–486
Goodwin EH, Meyne J (1993) Strand-specific FISH reveals orientation of chromosome 18 alphoid DNA. Cytogenet Cell Genet 63:126–127
Halkka L, Vakula N, Kaikusalo A (1994) Polymorphic and stable chromosomes of Sorex araneaus L. differ in centromere constitution (R-banding): evolutionary aspects. Ann Zool Fenn 31:289–296
Henikoff S, Ahmad K, Malik HS (2001) The centromere paradox: stable inheritance with rapidly evolving DNA. Science 293(5532):1098–1102. doi:10.1126/science.1062939
Kalitsis P, Fowler KJ, Earle E, Griffiths B, Howman E, Newson AJ, Choo KHA (2003) Partially functional Cenpa-GFP fusion protein causes increased chromosome missegregation and apoptosis during mouse embryogenesis. Chromosom Res 11:345–357
Kalitsis P, Griffiths B, Choo KHA (2006) Mouse telocentric sequences reveal a high rate of homogenization and possible role in Robertsonian translocation. Proc Natl Acad Sci 103(23):8786–8791. doi:10.1073/pnas.0600250103
Kemkemer C, Kohn M, Cooper DN, Froenicke L, Högel J, Hameister H, Kehrer-Sawatzki H (2009) Gene synteny comparisons between different vertebrates provide new insights into breakage and fusion events during mammalian karyotype evolution. BMC Evol Biol 9:84
Kipling D, Warburton PE (1997) Centromeres, CENP-B and Tigger too. Trends Genet 13(4):141–145
Kipling D, Mitchell AR, Masumoto H, Wilson HE, Nicol L, Cooke HJ (1995) CENP-B binds a novel centromeric sequence in the Asian mouse Mus caroli. Mol Cell Biol 15:4009–4020
Kunze B, Traut W, Garagna S, Weichenham D, Redi CA, Winking H (1999) Pericentric satellite DNA and molecular phylogeny in Acomys (Rodentia). Chromosom Res 7:131–141
Lee MR, Elder FFB (1980) Yeast simulation of bone marrow mitosis for cytogenetic investigations. Cytogenet Cel Genet 26:36–40
Metcalfe CJ, Bulazel KV, Ferreri GC, Schroeder-Reiter E, Wanner G, Rens W, Obergfell C, Eldridge MDB, O'Neill RJ (2007) Genomic instability within centromeres of interspecific Marsupial hybrids. Genetics 177:2507–2517
Mitchell AR (1996) The mammalian centromere: its molecular architecture. Mutat Res 372:153–162
Mlynarski EE, Obergfell CJ, O’Neill MJ, O’Neill RJ (2010) Divergent patterns of breakpoint reuse in Muroid rodents. Mamm Genome 21:77–87
Nanda I, Schneider-Rasp S, Winking H, Schmid M (1995) Loss of telomeric sites in the chromosomes of Mus musculus domesticus (Rodentia: Muridae) during Robertsonian rearrangements. Chromosom Res 3:399–409
Narayanswami S, Doggett NA, Clark LM, Hildebrand CE, Weier HU, Hamkalo BA (1992) Cytological and molecular characterization of centromeres in Mus domesticus and Mus spretus. Mamm Genome 2(3):186–194. doi:10.1007/bf00302876
Nguyen TT, Aniskin VM, Gerbault-Seureau M, Planton H, Renard JP, Nguyen BX, Hassanin A, Volobouev VT (2008) Phylogenetic position of the saola (Pseudoryx nghetinhensis) inferred from cytogenetic analysis of eleven species of Bovidae. Cytogenet Genome Res 122:41–54
Nijman IJ, Lenstra JA (2001) Mutation and recombination in cattle satellite DNA: a feedback model for the evolution of satellite DNA repeats. J Mol Evol 52(4):361–371
O’Neill RJW, O’Neill MJ, Graves JAM (1998) Undermethylation associated with retroelement activation and chromosome remodeling in an interspecific mammalian hybrid. Nature 393:68–72
Pertile MD, Graham AN, Choo AKH, Kalitsis P (2009) Rapid evolution of mouse Y centromere repeat DNA belies recent sequence stability. Genome Res 19
Piálek J, Hauffe HC, Searle JB (2005) Chromosomal variation in the house mouse. Biol J Linn Soc 84:535–563
Pietras DF, Bennett KL, Siracusa LD, Woodworth-Gutai M, Chapman VM, Gross KW, Kane-Haas C, Hastie ND (1983) Construction of small Mus musculus repetitive DNA library: identification of a new satellite sequences in Mus musculus. Nucleic Acids Res 11:6976–6983
Plohl M, Luchetti A, Mestrovic N, Mantovani B (2008) Satellite DNAs between selfishness and functionality: structure, genomics and evolution of tandem repeats in centromeric (hetero)chromatin. Gene 409:72–82
Rebuzzini P, Castiglia R, Nergadze SG, Mitsainas GP, Munclinger P, Zuccotti M, Capanna E, Redi C, Garagna S (2009) Quantitative variation of Line-1 sequences in five species and three subspecies of the subgenus Mus and in five Robertsonian races of Mus musculus domesticus. Chromosome Res 17(1):65–76
Redi C, Garagna S, Zuccotti M (1990a) Robertsonian chromosome formation and fixation: the genomic scenario. Biol J Linn Soc 41:235–255
Redi CA, Garagna S, Della Valle G, Bottiroli G, Dell'Orto P, Viale G, Peverali FA, Raimondi E, Forejt J (1990b) Differences in the organization and chromosomal allocation of satellite DNA between the European long tailed house mice Mus domesticus and Mus musculus. Chromosoma 99:11–17
Robertson W (1916) Chromosome studies. I. Taxonomic relationships shown in the chromosomes of Tettigidae and Acrididae. V-shaped chromosomes and their significance in Acrididae, Locustidae and Gryllidae: chromosomes and variation. J Morphol 27:179–331
Roeder GS (1997) Meiotic chromosomes: it takes two to tango. Genes Dev 11(20):2600–2621. doi:10.1101/gad.11.20.2600
Sasaki N, Yamauchi H, Tomohiro N, Agui T (2012) The telocentric tandem repeat at the p-arm is not conserved in Mus musculus subspecies. Gene. doi:10.1016/j.gene.2012.10.050
Schueler MG, Dunn JM, Bird CP, Ross MT, Viggiano L, Program NCS, Rocchi M, Willard HF, Green ED (2005) Progressive proximal expansion of the primate X chromosome centromere. Proc Natl Acad Sci USA 102:10563–11568
Searle JB, Wójcik JM (1998) Chromosomal evolution: the case of Sorex araneus. In: Wójcik JM, Wolsan M (eds) Evolution of shrews. Mammal Research Institute, Polish Academy of Sciences, Bialowieza, pp 219–268
Sen S, Sharma T (1983) Role of constitutive heterochromatin in evolutionary divergence: results of chromosome banding and condensation inhibition studies in Mus musculus, Mus booduga and Mus dunni. Evolution 37:628–636
Shimada T, Aplin KP, Suzuki H (2010) Mus lepidoides (Muridae, Rodentia) of central Burma is a distinct species of potentially great evolutionary and biogeographic significance. Zool Sci 27:449–459
Slamovits CH, Rossi MS (2002) Satellite DNA: agent of chromosomal evolution in mammals. A review. J Neotropical Mamm 9:297–308
Song Y, Endepols S, Klemann N, Richter D, Matuschka F-R, Shih C-H, Nachman MW, Koh MH (2011) Adaptive introgression of anticoagulant rodent poison resistance by hybridization between old world mice. Curr Biol 21:1296–1301
Suzuki H, Shimada T, Terashima M, Tsuchiya K, Aplin K (2004) Temporal, spatial, and ecological modes of evolution of Eurasian Mus based on mitochondrial and nuclear gene sequences. Mol Phylogenet Evol 33:626–646
Thomas JW, Schueler MG, Summers TJ, Blakesley RW, McDowell JC, Thomas PJ, Idol JR, Maduro VVB, Lee-Lin S-Q, Touchman JW, Bouffard GG, Beckstrom-Sternberg SM, Program NCS, Green ED (2003) Pericentromeric duplications in the laboratory mouse. Genome Res 13:55–63
Trifonov VA, Kosyakova N, Romanenko SA, Stanyon R, Graphodatsky AS, Lieh T (2010) New insights into the karyotypic evolution in muroid rodents revealed by multicolor banding applying murine probes. Chromosom Res. doi:10.1007/s10577-010-9110-6
Van Vuuren BJ, Robinson TJ (2001) Retrieval of four adaptive lineage in duiker antelope: evidence from mitochondrial DNA sequences and fluorescence in situ hybridization. Mol Phylogenet Evol 20:409–425
Veyrunes F, Dobigny G, Yang F, O'Brien PCM, Catalan J, Robinson TJ, Britton-Davidian J (2006) Phylogenomics of the genus Mus (Rodentia; Muridae): extensive genome repatterning is not restricted to the house mouse. Proc R Soc Lond B 273:2925–2934
Vidal-Rioja L, Semorile L, Bianchi NO, Padron J (1987) DNA composition in South American camelids. I. Characterization and in situ hybridization of satellite DNA fraction. Genetica 72:137–146
Vissel B, Choo KH (1989) Mouse major (gamma) satellite DNA is highly conserved and organized into extremely long tandem arrays—implications for recombination between nonhomologous chromosomes. Genomics 5(3):407–414. doi:10.1016/0888-7543(89)90003-7
Acknowledgments
We are grateful to A. Orth for animal samples, C. Desmaze for her technical assistance in CO-FISH and P. Kalitsis for providing the major, minor and TLC probes. The Southern blot complement could not have been performed without the enlightened and much appreciated assistance of C. Mari, P. Caminade, E. Douzery, B. de Massy, J.-C. Davidian and C. Benoist. We are grateful to Gunilla Rosenqvist for assistance in the completion of this work. The GDR 3047 (Cytogénomique Structurale et Evolutive) financed a travel grant for training in CO-FISH procedures. All cytogenetic analyses were performed in the Plateforme de Cytogénomique Evolutive of the SFR MEB in Montpellier. This study was supported by recurrent funding from the CNRS and UM2.This is publication ISEM no. 2013-008.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Hiroshi Masumoto
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(JPEG 149 kb)
High resolution
(TIFF 22668 kb)
ESM 2
(JPEG 164 kb)
High resolution
(TIFF 2486 kb)
Rights and permissions
About this article
Cite this article
Cazaux, B., Catalan, J., Justy, F. et al. Evolution of the structure and composition of house mouse satellite DNA sequences in the subgenus Mus (Rodentia: Muridea): a cytogenomic approach. Chromosoma 122, 209–220 (2013). https://doi.org/10.1007/s00412-013-0402-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-013-0402-4