Abstract
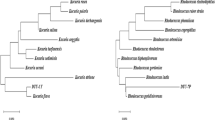
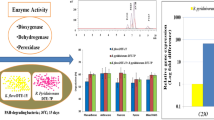
Polycyclic aromatic hydrocarbons (PAHs) are a group of environmental pollutant that are given top priority to maintain water and soil quality to the most amenable standard. Biodegradation of PAHs by bacteria is the convenient option for decontamination on site or off site. The aim of the present study was to isolate and identify naturally occurring bacteria having mixed PAHs biodegradation ability. The newly isolated Pseudomonas putida strain KD6 was found to efficiently degrade 97.729% of 1500 mg L−1 mixed PAHs within 12 days in carbon-deficient minimal medium (CSM). The half-life (t 1/2) and degradation rate constant (k) were estimated to be 3.2 and 0.2165 days, respectively. The first-order kinetic parameters in soil by strain KD6 had shown efficient biodegradation potency with the higher concentration of total PAHs (1500 mg kg−1 soil), t 1/2 = 10.44 days−1. However, the biodegradation by un-inoculated control soil was found slower (t 1/2 = 140 days−1) than the soil inoculated with P. putida strain KD6. The enzyme kinetic constants are also in agreement with chemical data obtained from the HPLC analysis. In addition, the sequence analysis and molecular docking studies showed that the strain KD6 encodes a mutant version of naphthalene 1,2-dioxygenase which have better Benzpyrene binding energy (−9.90 kcal mol−1) than wild type (−8.18 kcal mol−1) enzyme (chain A, 1NDO), respectively, with 0.00 and 0.08 RMSD values. The mutated naphthalene 1,2-dioxygenase nahAc has six altered amino acid residues near to the ligand binding site. The strain KD6 could be a good bioresource for in situ or ex situ biodegradation of polycyclic aromatic hydrocarbon.





Similar content being viewed by others
References
Abdel-Shafy HI, Mansour MSM (2015) A review on polycyclic aromatic hydrocarbons: source, environmental impact, effect on human health and remediation. Egypt J Pet 25:107–123. doi:10.1016/j.ejpe.2015.03.011
Akbar S, Sultan S, Kertesz M (2014) Determination of cypermethrin degradation potential of soil bacteria along with plant growth-promoting characteristics. Curr Microbiol 70:75–84. doi:10.1007/s00284-014-0684-7
Albuquerque M, Coutinho M, Borrego C (2016) Long-term monitoring and seasonal analysis of polycyclic aromatic hydrocarbons (PAHs) measured over a decade in the ambient air of Porto, Portugal. Sci Total Environ 543:439–448. doi:10.1016/j.scitotenv.2015.11.064
Arvanitis N, Katsifas EA, Chalkou KI et al (2008) A refinery sludge deposition site: presence of nahH and alkJ genes and crude oil biodegradation ability of bacterial isolates. Biotechnol Lett 30:2105–2110. doi:10.1007/s10529-008-9816-0
Baird WM, Hooven LA, Mahadevan B (2005) Carcinogenic polycyclic aromatic hydrocarbon-DNA adducts and mechanism of action. Environ Mol Mutagen 45:106–114
Biasini M, Bienert S, Waterhouse A et al (2014) SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. doi:10.1093/nar/gku340
Bonilla M, Olivaro C, Corona M et al (2005) Production and characterization of a new bioemulsifier from Pseudomonas putida ML2. J Appl Microbiol 98:456–463. doi:10.1111/j.1365-2672.2004.02480.x
Carugo O, Pongor S (2008) A normalized root-mean-spuare distance for comparing protein three-dimensional structures. Protein Sci 10:1470–1473. doi:10.1110/ps.690101
Chemical Computing Group Inc (2011) Molecular operating environment (MOE), 10th edn. Chemical Computing Group Inc, Montreal
Chen JE, Huang CC, Ferrin TE (2015) RRDistMaps: a UCSF chimera tool for viewing and comparing protein distance maps. Bioinformatics 31:1484–1486. doi:10.1093/bioinformatics/btu841
Chou C-C, Wang AH-J (2015) Structural D/E-rich repeats play multiple roles especially in gene regulation through DNA/RNA mimicry. Mol BioSyst 11:2144–2151. doi:10.1039/C5MB00206K
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genom Res 14:1188–1190. doi:10.1101/gr.849004
Diggs DL, Huderson AC, Harris KL et al (2011) Polycyclic aromatic hydrocarbons and digestive tract cancers: a perspective. J Environ Sci Heal Part C 29:324–357. doi:10.1080/10590501.2011.629974
Duran R, Cravo-Laureau C (2016) Role of environmental factors and microorganisms in determining the fate of polycyclic aromatic hydrocarbons in the marine environment. FEMS Microbiol Rev 40:814–830. doi:10.1093/femsre/fuw031
Eisenthal R, Danson MJ, Hough DW (2007) Catalytic efficiency and kcat/KM: a useful comparator? Trends Biotechnol 25:247–249. doi:10.1016/j.tibtech.2007.03.010
Gallego JLR, García-Martínez MJ, Llamas JF et al (2007) Biodegradation of oil tank bottom sludge using microbial consortia. Biodegradation 18:269–281. doi:10.1007/s10532-006-9061-y
Govarthanan M, Mythili R, Selvankumar T et al (2016) Bioremediation of heavy metals using an endophytic bacterium Paenibacillus sp. RM isolated from the roots of Tridax procumbens. 3 Biotech 6:1–7. doi:10.1007/s13205-016-0560-1
Guo W, He M, Yang Z et al (2011) Aliphatic and polycyclic aromatic hydrocarbons in the Xihe River, an urban river in China’s Shenyang City: distribution and risk assessment. J Hazard Mater 186:1193–1199. doi:10.1016/j.jhazmat.2010.11.122
Ifegwu C, Osunjaye K, Fashogbon F, Oke K, Adeniyi A, Anyakora C, Ifegwu C, Osunjaye K et al (2012) Urinary 1-hydroxypyrene as a biomarker to carcinogenic polycyclic aromatic hydrocarbon exposure. Biomark Cancer 4:7. doi:10.4137/BIC.S10065
Iyer R, Stepanov VG, Iken B (2013) Isolation and molecular characterization of a novel Pseudomonas putida strain capable of degrading organophosphate and aromatic compounds. Adv Biol Chem 3:564–578. doi:10.4236/abc.2013.36065
Jacques RJS, Okeke BC, Bento FM et al (2008) Microbial consortium bioaugmentation of a polycyclic aromatic hydrocarbons contaminated soil. Bioresour Technol 99:2637–2643. doi:10.1016/j.biortech.2007.04.047
Keshavarzifard M, Zakaria MP, Shau Hwai T et al (2014) Baseline distributions and sources of Polycyclic Aromatic Hydrocarbons (PAHs) in the surface sediments from the Prai and Malacca Rivers, Peninsular Malaysia. Mar Pollut Bull 88:366–372. doi:10.1016/j.marpolbul.2014.08.014
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol. doi:10.1093/molbev/msw054
Lee K (1999) Benzene-induced uncoupling of naphthalene dioxygenase activity and enzyme inactivation by production of hydrogen peroxide. J Bacteriol 181:2719–2725
Lee B-K, Vu VT (2010) Sources, distribution and toxicity of polycyclic aromatic hydrocarbons (PAHs) in particulate matter. In: Air pollution. pp 22–25
Lin QS, Chen SH, Hu MY et al (2011) Biodegradation of cypermethrin by a newly isolated actinomycetes HU-S-01 from wastewater sludge. Int J Environ Sci Technol 8:45–56. doi:10.1007/BF03326194
Lodish H, Berk A, Zipursky SL et al (2000) Molecular cell biology, 4th edn. WH Freeman, New York
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193:265–275. doi:10.1016/0304-3894(92)87011-4
Mendoza VL, Vachet RW (2009) Probing protein structure by amino acid-specific covalent labeling and mass spectrometry. Mass Spectrom Rev 28:785–815
Pacwa-Płociniczak M, Płaza GA, Piotrowska-Seget Z, Cameotra SS (2011) Environmental applications of biosurfactants: recent advances. Int J Mol Sci 12:633–654
Peng N, Li Y, Liu Z et al (2016) Emission, distribution and toxicity of polycyclic aromatic hydrocarbons (PAHs) during municipal solid waste (MSW) and coal co-combustion. Sci Total Environ 565:1201–1207. doi:10.1016/j.scitotenv.2016.05.188
Pérez-Pantoja D, Donoso R, Agulló L et al (2012) Genomic analysis of the potential for aromatic compounds biodegradation in Burkholderiales. Environ Microbiol 14:1091–1117
Pettersen EF, Goddard TD, Huang CC et al (2004) UCSF chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612. doi:10.1002/jcc.20084
Qi J, Wang B, Li J et al (2015) Genetic determinants involved in the biodegradation of naphthalene and phenanthrene in Pseudomonas aeruginosa PAO1. Environ Sci Pollut Res 22:6743–6755. doi:10.1007/s11356-014-3833-4
Rengarajan T, Rajendran P, Nandakumar N et al (2015) Exposure to polycyclic aromatic hydrocarbons with special focus on cancer. Asian Pac J Trop Biomed 5:182–189. doi:10.1016/S2221-1691(15)30003-4
Samanta SK, Singh OV, Jain RK (2002) Polycyclic aromatic hydrocarbons: environmental pollution and bioremediation. Trends Biotechnol 20:243–248. doi:10.1016/S0167-7799(02)01943-1
Sanner MF (1999) News and views. J Mol Graph Model 17:55–84. doi:10.1016/S1093-3263(99)99999-0
Tobiszewski M, Namieśnik J (2012) PAH diagnostic ratios for the identification of pollution emission sources. Environ Pollut 162:110–119
Valavanidis A, Vlahogianni T, Dassenakis M, Scoullos M (2006) Molecular biomarkers of oxidative stress in aquatic organisms in relation to toxic environmental pollutants. Ecotoxicol Environ Saf 64:178–189. doi:10.1016/j.ecoenv.2005.03.013
Warwicker J, Charonis S, Curtis RA (2014) Lysine and arginine content of proteins: computational analysis suggests a new tool for solubility design. Mol Pharm 11:294–303. doi:10.1021/mp4004749
World Health Organization (2010) WHO guidelines for indoor air quality: selected pollutants. Bonn, Ger puncto druck + Medien GmbH 484. doi: 10.1186/2041-1480-2-S2-I1
Yang Q, Chen H, Li B (2015) Polycyclic aromatic hydrocarbons (PAHs) in indoor dusts of guizhou, southwest of china: status, sources and potential human health risk. PLoS ONE. doi:10.1371/journal.pone.0118141
Zhu X, Ni X, Gatheruwaigi M et al (2016) Biodegradation of mixed PAHS by PAH-degrading endophytic bacteria. Int J Environ Res Public Health 13:805. doi:10.3390/ijerph13080805
Acknowledgements
This study was supported by University Grands Commission (VU/Innovative/Sc/17/2015), Govt. of India. We are thankful to Mr. Dipankar Mandal, USIC, Vidyasagar University for the HPLC analysis. We are grateful to Mr. Smriti Ranjan Majhi, Bose Institute, for GCMS analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dutta, K., Shityakov, S., Das, P.P. et al. Enhanced biodegradation of mixed PAHs by mutated naphthalene 1,2-dioxygenase encoded by Pseudomonas putida strain KD6 isolated from petroleum refinery waste. 3 Biotech 7, 365 (2017). https://doi.org/10.1007/s13205-017-0940-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-017-0940-1