Abstract
Polyaromatic hydrocarbons (PAHs) are hazardous organic compounds with established toxicity, carcinogenicity, and mutagenicity, ubiquitous distribution, and persistence in different environmental matrices. In the present study, degradation of the mixture of PAHs (phenanthrene, anthracene, fluorene, and pyrene) by Kocuria flava and Rhodococcus pyridinivorans was investigated. The individual strains and consortium of both degraded 55.6%, 59.5%, and 59.1% of 10 mg L−1 of mixed PAHs, respectively, within 15 days. The participation of catabolic enzymes [catechol 2,3-dioxygenase (C23O), dehydrogenase (DH), and peroxidase (POD)] was confirmed during catalytic oxidation through meta-cleavage of mixed PAHs in this study. The catabolic gene expression of naphthalene dioxygenase (NAH) and catechol 2,3-dioxygenase (C23O) during degradation was confirmed using RT-qPCR in the present study. This is the first study that shows significant gene expression of the catabolic genes during degradation of mixed PAHs by selected bacterial strains. The C23O gene showed a 6.02 log fold higher expression in Kocuria flava in comparison to Rhodococcus pyridinivorans whereas NAH gene exhibited a 7.9 log fold higher expression in Rhodococcus pyridinivorans in comparison to Kocuria flava. Hence it is likely to conclude that combination of Kocuria flava and Rhodococcus pyridinivorans can effectively remove hazardous mixture of PAHs from the contaminated environmental matrix.
Graphical abstract








Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this article.
References
Afzal M, Yousaf S, Reichenauer TG, Kuffner M, Sessitsch A (2011) Soil type affects plant colonization, activity and catabolic gene expression of inoculated bacterial strains during phytoremediation of diesel. J Hazard Mater 186(2–3):1568–1575. https://doi.org/10.1016/j.jhazmat.2010.12.040
Agency for Toxic Substances and Disease Registry (ATSDR) (1990) Toxicological profile for polycyclic aromatic hydrocarbons. Acenaphthene, Acenaphthylene, Anthracene, Benzo[a]anthracene, Benzo[a]pyrene, Benzo[b]fluoranthene, Benzo[g,i,h]perylene, Benzo[k]fluoranthene, Chrysene, Dibenzo[a,h]anthracene, Fluoranthene, Fluorene, Indeno[1,2,3-c,d]pyrene, Phenanthrene, Pyrene. Atlanta, GA:Agency for Toxic Substances and Disease Registry.
Alegbeleye OO, Opeolu BO, Jackson V (2017) Bioremediation of polycyclic aromatic hydrocarbon (PAH) compounds: (acenaphthene and fluorene) in water using indigenous bacterial species isolated from the Diep and Plankenburg rivers, Western Cape, South Africa. Braz J Microbiol 48(2):10. https://doi.org/10.1016/j.bjm.2016.07.027
Badejo AC, Badejo AO, Shin KH, Chai YG (2013) A gene expression study of the activities of aromatic ring-cleavage dioxygenases in Mycobacterium gilvum PYR-GCK to changes in salinity and pH during pyrene degradation. PLoS ONE 8(2):e58066. https://doi.org/10.1371/journal.pone.0058066
Bamforth SM, Singleton I (2005) Bioremediation of polycyclic aromatic hydrocarbons: current knowledge and future directions. J Chem Technol Biotechnol 80(7):723–736. https://doi.org/10.1002/jctb.1276
Cameselle C, Gouveia S (2019) Phytoremediation of mixed contaminated soil enhanced with electric current. J Hazard Mater 361:95–102
Cao L, Wang Q, Zhang J, Li C, Yan X, Lou X, Xia Y, Hong Q, Li S (2012) Construction of a stable genetically engineered rhamnolipid-producing microorganism for remediation of pyrene-contaminated soil. World J Microbiol Biotechnol 28(9):2783–2790
Cerniglia CE (1993) Biodegradation of polycyclic aromatic hydrocarbons. Curr Opin Biotechnol 4:331–338
Chen M, Xu P, Zeng G, Yang C, Huang D, Zhang J (2015) Bioremediation of soils contaminated with polycyclic aromatic hydrocarbons, petroleum, pesticides, chlorophenols and heavy metals by composting: applications, microbes and future research needs. Biotechnol Adv 33(6):745–755. https://doi.org/10.1016/j.biotechadv.2015.05.003
Dhanya MS, Kalia A (2020) Bioremediation: an eco-friendly cleanup strategy for polyaromatic hydrocarbons from petroleum industry waste. In: Saxena G, Bharagava R (eds) Bioremediation of industrial waste for environmental safety. Springer, Singapore. https://doi.org/10.1007/978-981-13-1891-7_18
Dutta K, Shityakov S, Das PP, Ghosh C (2017) Enhanced biodegradation of mixed PAHs by mutated naphthalene 1,2-dioxygenase encoded by Pseudomonas putida strain KD6 isolated from petroleum refinery waste. 3 Biotech 7(6):365. https://doi.org/10.1007/s13205-017-0940-1
Elyamine AM, Kan J, Meng S, Tao P, Wang H, Hu Z (2021) Aerobic and anaerobic bacterial and fungal degradation of pyrene: mechanism pathway including biochemical reaction and catabolic genes. Int J Mol Sci 22:8202. https://doi.org/10.3390/ijms22158202
Gennaro PD, Moren B, Annoni E, García-Rodríguez S, Bestetti G, Benitez E (2009) Dynamic changes in bacterial community structure and in naphthalene dioxygenase expression in vermicompost-amended PAH-contaminated soils. J Hazard Mater 172(2–3):14640–21469. https://doi.org/10.1016/j.jhazmat.2009.08.013
Goyal AK, Zylstra GJ (1996) Molecular cloning of novel genes for polycyclic aromatic hydrocarbon degradation from Comamonas testosteroni GZ39. Appl Environ Microbiol 62:230–236
Haritash AK, Kaushik CP (2009) Biodegradation aspects of polycyclic aromatic hydrocarbons: a review. J Hazard Mater 169:1–15
Haritash AK, Kaushik CP (2016) Degradation of low molecular weight polycyclic aromatic hydrocarbons by microorganisms isolated from contaminated soil. Int J Environ Sci 6(5):646–656
Hennessee CT, Li QX (2016) Effects of polycyclic aromatic hydrocarbon mixtures on degradation, gene expression, and metabolite production in four Mycobacterium species. Appl Environ Microbiol 82(11):3357–3369. https://doi.org/10.1128/AEM.00100-16
Hesham AEL, Mawad AMM, Mostafa YM, Shoreit A (2014) Biodegradation ability and catabolic genes of petroleum-degrading Sphingomonas koreensis strain ASU-06 isolated from Egyptian oily soil. BioMed Res Int 2014:1–10. https://doi.org/10.1155/2014/127674
Institute for Health Metrics and Evaluation (IHME), Global Burden of Disease (2019)
Juhasz AL, Britz ML, Stanley GA (1996) Degradation of high molecular weight polycyclic aromatic hydrocarbons by Pseudomonas cepacia. Biotechnol Lett 18:577–582
Kachieng’a L, Momba M (2017) Kinetics of petroleum oil biodegradation by a consortium of three protozoan isolates (Aspidisca sp., Trachelophyllum sp. and Peranema sp.). Biotechnol Rep (Amsterdam, Netherlands) 15:125–131. https://doi.org/10.1016/j.btre.2017.07.001
Kelley I, Cerniglia CE (1995) Degradation of a mixture of high-molecular-weight polycyclic aromatic hydrocarbons by a Mycobacterium strain PYR-1. J Soil Contam 4:77–91
Keum YS, Seo JS, Li QX, Kim JH (2008) Comparative metabolomic analysis of Sinorhizobium sp C4 during the degradation of phenanthrene. Appl Microbiol Biotechnol 80:863–872
Ławniczak Ł, Woźniak-Karczewska M, Loibner AP, Heipieper HJ, Chrzanowski Ł (2020) Microbial degradation of hydrocarbons—basic principles for bioremediation: a review. Molecules 25(4):856. https://doi.org/10.3390/molecules25040856
Liu K, Han W, Pan WP, Riley JT (2001) Polycyclicaromatic hydrocarbon (PAH) emissions from a coal fired pilot FBC system. J Hazard Mater 84:175–188. https://doi.org/10.1016/S0304-3894(01)00196-0
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta C(T)) method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Lu XY, Zhang T, Fang HHP (2011) Bacteria-mediated PAH degradation in soil and sediment. Appl Microbiol Biotechnol 89:1357–1371. https://doi.org/10.1007/s00253-010-3072-7
Lyu Y, Zheng W, Zheng T, Tian Y (2014) Biodegradation of polycyclic aromatic hydrocarbons by Novosphingobium pentaromativorans US6-1. PLoS ONE 9(7):e101438
Margesin R, Zimmerbauer A, Schinner F (2000) Monitoring of bioremediation by soil biological activities. Chemosphere 40:339–346
Mawad AMM, Abdel-Mageed WS, Hesham AEL (2020) Quantification of naphthalene dioxygenase (NahAC) and catechol dioxygenase (C23O) catabolic genes produced by phenanthrene-degrading Pseudomonas fluorescens AH-40. Curr Genomics 21(2):111–118. https://doi.org/10.2174/1389202921666200224101742
Meyer S, Moser R, Neef A, Stahl U, Kämpfer P (1999) Differential detection of key enzymes of polyaromatic-hydrocarbon-degrading bacteria using PCR and gene probes. Microbiology 145:1731–41
Naether D, Slawtschew S, Stasik S, Engel M, Olzog M, Wick LY, Timmis KN, Heipieper HJ (2013) Adaptation of hydrocarbonoclastic Alcanivorax borkumensis SK2 to alkanes and toxic organic compounds—a physiological and transcriptomic approach. Appl Environ Microb 79(14):4282–4293. https://doi.org/10.1128/AEM.00694-13
Olowomofe TO, Oluyege JO, Aderiye BI, Oluwole OA (2019) Degradation of poly aromatic fractions of crude oil and detection of catabolic genes in hydrocarbon-degrading bacteria isolated from Agbabu bitumen sediments in Ondo State. AIMS Microbiol 5(4):308–323. https://doi.org/10.3934/microbiol.2019.4.308
Pandey AK, Chaudhary P, Singh SB, Arora A, Kumar K, Chaudhry S, Nain L (2012) Deciphering the traits associated with PAH degradation by a novel Serratia marcesencs L-11 strain. J Environ Sci Health A 47(5):755–765
Peng T, Luo A, Kan J, Liang L, Huang T, Hu Z (2018) Identification of a ring-hydroxylating dioxygenases capable of anthracene and benz[a]anthracene oxidization from Rhodococcus sp. P14. J Mol Microbiol Biotechnol 28:183–189. https://doi.org/10.1159/000494384
Pinyakong O, Habe H, Supaka N, Pinpanichkarn P, Juntongjin K, Yoshida T, Furihata K, Nojiri H, Yamane H, Omori T (2000) Identification of novel metabolites in the degradation of phenanthrene by Sphingomonas sp. Strain P2. FEMS Microbiol Lett 191:115–121. https://doi.org/10.1111/j.1574-6968.2000.tb09327.x
Reddy KR, Adams JA (2015) Sustainable remediation of contaminated sites. Momentum Press, New York
Rengarajan T, Rajendran P, Nandakumar N, Lokeshkumar B, Rajendran P, Nishigaki I (2015) Exposure to polycyclic aromatic hydrocarbons with special focus on cancer. Asian Pac J Trop Biomed 5(3):182–189. https://doi.org/10.1016/S2221-1691(15)30003-4
Rio DC, Ares M Jr, Hannon GJ, Nilsen TW (2010) Purification of RNA using TRIzol (TRI reagent). Cold Spring Harb Protoc 6:prot5439. https://doi.org/10.1101/pdb.prot5439
Rogers SW, Ong SK, Kjartanson BH, Golchin J, Stenback GA (2002) Natural attenuation of polycyclic aromatic hydrocarbon-contaminated sites: review. Pract Period Hazard Toxic Radioact Waste Manag 6(3):141–155. https://doi.org/10.1061/(asce)1090-025x(2002
Sakshi HAK (2020) A comprehensive review of metabolic and genomic aspects of PAH- degradation. Arch Microbiol 202(8):2033–2058. https://doi.org/10.1007/s00203-020-01929-5
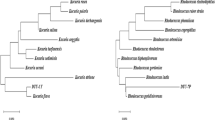
Sakshi SSK, Haritash AK (2020) Evolutionary relationship of polycyclic aromatic hydrocarbons degrading bacteria with strains isolated from petroleum contaminated soil based on 16S rRNA diversity. Polycycl Aromat Compd. https://doi.org/10.1080/10406638.2020.1825003
Sakshi SSK, Haritash AK (2021a) Catabolic enzyme activities during biodegradation of three-ring PAHs by novel DTU-1Y and DTU-7P strains isolated from petroleum-contaminated soil. Arch Microbiol 203(6):3101–3110. https://doi.org/10.1007/s00203-021-02297-4
Sakshi SSK, Haritash AK (2021b) Catabolic enzyme activity and kinetics of pyrene degradation by novel bacterial strains isolated from contaminated soil. Environ Technol Innov. https://doi.org/10.1016/j.eti.2021.101744
Sakshi, Singh SK, Haritash AK (2019) Polycyclic aromatic hydrocarbons: soil pollution and remediation. Int J Environ Sci Technol 16:6489–6512. https://doi.org/10.1007/s13762-019-02414-3
Sangkharak K, Choonut A, Rakkan T, Prasertsan P (2020) The degradation of phenanthrene, pyrene, and fluoranthene and its conversion into medium-chain-length polyhydroxyalkanoate by novel polycyclic aromatic hydrocarbon-degrading bacteria. Curr Microbiol. https://doi.org/10.1007/s00284-020-01883-x
Sayler GS, Perkins RE, Sherrill TW, Perkins BK, Reid MC, Shields MS, Kong HL, Davis JW (1983) Microcosm and experimental pond evaluation of microbial community response to synthetic oil contamination in freshwater sediments. Appl Environ Microbiol 46:211–219
Van Hamme JD, Singh A, Ward OP (2003) Recent advances in petroleum microbiology. Microbiol Mol Biol Rev 67(4):503–549
Wang Z, Li J, Hesham AE, He S, Zhang Y, Wang Z, Yang M (2007) Co-variations of bacterial composition and catabolic genes related to PAH degradation in a produced water treatment system consisting of successive anoxic and aerobic units. Sci Total Environ 373(1):356–362
Acknowledgements
The authors acknowledge the help of Superworth Biodiscoveries, Delhi, India, for analysis of gene expression.
Funding
The authors declare that no funds, grants or other support was received during the preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
The study was conceptualised and executed by Sakshi and Anil Kumar Haritash; and drafted by all the authors together.
Corresponding author
Ethics declarations
Competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sakshi, Singh, S.K. & Haritash, A.K. Bacterial degradation of mixed-PAHs and expression of PAH-catabolic genes. World J Microbiol Biotechnol 39, 47 (2023). https://doi.org/10.1007/s11274-022-03489-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-022-03489-w