Abstract
Ochratoxin A (OTA) is a mycotoxin produced by Aspergillus spp. and Penicillium spp. that causes a threat to food safety and human health. Fungal biodegradation might be a promising strategy for reducing the OTA contamination in the future. In this study, the ability of Trichoderma koningii strains to degrade OTA produced by Aspergillus niger T2 (MW513392.1) isolated from tomato seeds was investigated. Among T. koningii strains tested, three strains; AUMC11519, AUMC11520 and AUMC11521 completely eliminated OTA from the culture medium, while AUMC11522 strain eliminated only 41.82% of OTA. OTα-amide, 3-phenylpropionic acid, OTα and phenylalanine were assayed as degradation products by FTIR analysis and LC–MS/MS spectra. Carboxypeptidase A (CPA) was found responsible for OTA degradation when a metal ion chelator, EDTA, was added to cell free supernatants of the three effective strains. OTA detoxification by T. koningii could present new prospective strategies for a possible application in food commodities intoxicated with ochratoxin.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ochratoxin A (L-phenylalanine-N-[(5-chloro-3,4-dihydro-8- hydroxy-3-methyl-1-oxo-1H-2-benzopyran-7-yl)carbonyl]-(R)-isocoumarin) (OTA, Supplementary Fig. S1), is a mycotoxin, which was first discovered and chemically characterized by van der Merwe et al. (1965a, b) in a South African culture of Aspergillus ochraceus inoculated in a corn meal. Subsequently, it was found to be produced by different species of Aspergillus and Penicillium, particularly A. ochraceus, A. niger and P. verrucosum, respectively (Ali et al. 2013; García-Cela et al. 2014; Ismaiel and Papenbrock 2015).
OTA contaminates a wide range of food and feed commodities in many countries, such as cereals, cereal products, grapes, coffee, dried vine fruits, grape juice, nuts, spices and animal-derived food stuff (Duarte et al. 2012). OTA causes economic losses when synthesized in pre- and post-harvest plants as well as the losses occurred when synthesized in stored products, it also causes chlorosis of leaves, increased oxidative stress and even causes cell death of plants (Hao et al. 2015). OTA phytotoxicity is associated with damage of DNA, inhibition of protein synthesis, generation of reactive oxygen species, dysfunction of mitochondria, and the calcium homeostasis disruption (Wang et al. 2014). It also enhances free radical production, which may affect the redox-regulated antioxidant activity of antioxidant enzymes including catalase, glutamate cysteine ligase, superoxide dismutase, glutathione peroxidase, glutathione and glutathione-S-transferases (Preuss et al. 2014).
Previous studies showed that OTA has nephrotoxic, hepatotoxic, neurotoxic, teratogenic, mutagenic and immunotoxic characters and it is capable of causing kidney and liver tumors in mice and rats, its toxicity is dependent upon sex, the cellular type and the species of the tested animals (Abrunhosa et al. 2010). It was reported that OTA causes the formation of DNA-adducts after chronic exposure of OTA to rat and sub-acute exposure to pig (Faucet et al. 2004). OTA is also classified as possibly carcinogenic (group 2B) to humans as there is evidence of its carcinogenicity on experimental animals not on humans (IARC 1993). Due to all these health hazardous effects, OTA was subjected to legal regulations on both national and international levels by the World Health Organization, (WHO), which proposed the maximum level for OTA in cereals of 5 µg kg−1 (WHO 1991). Moreover, the European Union member states set new limits for dietary intake of OTA ranged between 15 and 60 ng kg−1 bw per week for adult consumers (European Food Safety Authority, EFSA, 2006).
The particularity of OTA is due to its high stability, as it is partially degraded during cooking conditions (Müller 1982), and it resists the food processing temperature. So that, it can be detected in processed food products such as wine, beer and bread (Duarte et al. 2010). It can resist 3 h of steam sterilization at 121 °C (Trivedi et al. 1992), and even at 250 °C, the destruction of the toxin is not complete (Boudra et al. 1995). It is also highly resistant to conventional treatment processes such as thermal sterilization and fermentation (Bullerman and Bianchini, 2007; Kabak 2009).
A considerable attention has been paid for the biodegradation of OTA, either in food products or in aqueous solutions. Cell cultures of plants (wheat, maize, tomato, soybean, sweet potato tubers) completely transformed OTA into a number of other products (Karlovsky 1999). Streptococcus salivarius subsp. thermophilus, Bifidobacterium bifidum, and yogurt bacteria have eliminated OTA levels in milk samples containing 0.05 and 0.1 mg OTA L−1; Lactobacillus delbrueckii subsp. bulgaricus reduced OTA level in milk samples containing 0.5 mg OTA L−1 (Škrinjar et al. 1996). In literature, some filamentous fungi showed potentiality of OTA biodegradation. A. fumigatus, A. japonicus and A. niger completely degraded 2 mg OTA L−1 after 10 days of incubation at 30 °C, it was degraded into OTα and further degradation into an unknown compound was observed (Varga et al. 2000). OTA was partially or completely degraded by A. niger and other filamentous fungi, and OTα was detected, especially in the assays carried out by A. niger and other black fungi (Abrunhosa et al. 2002). A. ochraceus (OTA non-producer), and some strains of A. wentii completely degraded OTA producing unidentified degradation metabolites (Varga et al. 2000). This study presents new insights on the biodegradation of OTA using endophytic strains of T. koningii to less- and non-toxic compounds. This is promising in food and agricultural application for the minimizing the toxicity of OTA.
Materials and methods
Chemicals and OTA standard
All chemicals and solvents used in this study were of high degree of purity. The standard OTA was obtained from Sigma-Aldrich, Taufkirchen, Germany. HPLC-grade solvents such as n-hexane, methylene chloride, chloroform and methanol were used for extraction procedures and thin layer chromatographic (TLC) analysis. Acetonitrile, methanol and citric acid monohydrate used for HPLC analysis, were purchased from Merck, Darmstadt, Germany.
Fungal strains used in the current study
Ochratoxigenic fungus
Aspergillus niger T2, was isolated from a tomato sample (Solanum lycopersicum L.), obtained from a retail market in Sharkia governorate, Egypt. It was isolated by dilute plate method on potato-dextrose agar (PDA) (Nazir et al. 2014). The fungal isolate was molecularly identified based on the sequence of PCR-amplified ITS1-5.8S and ITS4 rRNA-gene analysis performed at The Animal Health Research Institute, Dokki, Giza, Egypt. Sequence was further analyzed using Basic Local Alignment Search Tool (BLAST) program (Altschul et al. 1997) from the National Center of Biotechnology Information (NCBI) website. The sequence size of the fungal strain was successfully deposited in the Genebank with accession number MW513392.1.
Trichoderma koningii strains used in the biodegradation assay
Four endophytic T. koningii strains were used for OTA detoxification activities. These are T. koningii CTX1185, T. koningii CTX1172, T. koningii TD5391 and T. koningii TR2715. The first and second strains were previously isolated from Cupressus macrocarpa twig, while the third and fourth strains were isolated from the bark of Terminalia distichum and T. arjuna, respectively. They were morphologically identified and deposited in the Assiut University Mycological Center (AUMC, http://www.aun. edu.eg/aumc/aumc.htm) with accession numbers: AUMC11519, AUMC11520, AUMC11521, and AUMC11522, respectively (Ismaiel and Ali 2017).
Inoculum preparation and growth conditions
The ochratoxigenic fungal strain, A. niger MW513392.1 was cultivated in 250 mL Erlenmeyer flasks containing 50 mL PD broth with pH 5.6 at 30 °C. For inoculum preparation, fungal conidia from 7-day old cultures of the strain were harvested in sterile distilled water containing 0.01% Tween 80, and gently scrapped off with a sterile glass rod. The spore suspensions were then adjusted to final concentrations of 106 conidia mL−1 using a hemocytometer. One mL of the freshly prepared inoculum of the fungal strain was added to the Erlenmeyer flasks containing medium. Culture flasks were then dark-incubated at static conditions for 10 days at 30 °C.
Extraction and purification of OTA
Extraction of OTA was performed in two steps. Firstly, the culture filtrate of Aspergillus niger T2 (MW513392.1) was defatted with n-hexane, after which OTA was extracted with an equal volume of methylene chloride, then the mixture was shaken for 30 min and allowed to stand for 30 min in a separating funnel. The methylene chloride layer was filtered over anhydrous sodium sulfate and then evaporated under a vacuum to dryness (Daradimos et al. 2000; Téren et al. 1996; Valenta et al. 1993). OTA samples were then purified using column chromatography. The dried extract was dissolved in 0.01 M HCl (5 mL), filtered through glass microfiber filter (55-mm diameter GF/A, Whatman, Maidstone, UK). The solution was subjected to solid-phase extraction column (SPE) (Al-Hadithi et al. 2015). The packing material used in the SPE column was silica gel, C18 bonded to silica gel (Zheng et al. 2006). A rubber syringe plunger was used to push the sample extract through the SPE column which retains the impurities and purified extract was collected in a test tube (Malone et al. 1998).
Determination of OTA
TLC
In order to detect OTA, the TLC plates were prepared according to Stahl (1969). Ten grams of silica powder GF-254, purchased from Sigma-Aldrich, Taufkirchen, Germany, were mixed with 30 mL warm distilled water with continuous stirring till the formation of a slurry, then poured on a glass plate (20 × 20 cm) and allowed to solidify at room temperature. The plates were kept in an electric oven at 110 °C for 1 h in a vertical position, then used immediately. OTA was then detected qualitatively according to Téren et al. (1996). The dried crude extract was re-dissolved in absolute methanol (250 μL), spotted on TLC plates along with OTA standard solutions using glass capillary tubes approximately 2 cm away from the bottom and 2 cm away from the edges of the plate and in-between the spots. After that, the TLC plates were placed into a solvent tank containing chloroform: methanol (93:7, v/v) as a developing system. The solvent was removed when rises up and reaches approximately 2 cm from the end of the plates. The developed TLC plates were allowed to dry at room temperature. After which, OTA spots of samples and standard were visualized as greenish-blue fluorescence under UV (366 nm). The rate of flow (Rf) of OTA is calculated by using the following formula (Snyder 2008):
The spots of both samples and standard OTA scrapped off, eluted in 3 mL methanol, and centrifuged at 2516 ×g for 10 min (Nesheim 1976). The OTA supernatants were estimated using a UV spectrophotometer (6800UV/ Vis. Spectrophotometer, Jenway) at 365 nm against 3 mL of methanol as control and concentrations were obtained from a standard curve (Nesheim 1976).
HPLC
HPLC analysis was performed at Animal Health Research Institute, Dokki, Giza, Egypt, using Agilent Series 1200 quaternary gradient pump, Series 1200 auto sampler, Series 1200 FLD detector, and HPLC 2D Chemstation software (Hewlett-Packard, Les Ulis, Germany). The chromatographic separation was performed using a reversed-phase column (Extend-C18, Zorbax column, 4.6 mm i.d., 250 mm, 5 μm, Agilent Co.), in which the mobile phase should be sufficiently transparent at the wavelength of detection. The mobile phase used was water: acetonitrile: methanol (60: 20: 20, v:v:v) (Durguti et al. 2014). The fluorescent detector was set to a wavelength of 330 nm for excitation and 460 nm for emission. The column temperature adjusted at 30 °C at a flow rate of 1.0 mL min−1 to achieve the optimum resolution of the OTA. The injection volume was maintained at 20 μL for both sample and standard. Calibration curve was prepared using different concentrations of OTA standard. The linearity of detector response for the standard was determined by means of linear regression.
OTA concentration used in degradation tests
The chromatographically separated OTA from A. niger MW513392.1 was pooled and dissolved in chloroform (AnalaR) at concentration of 10 mg mL−1 and stored in the dark at – 20 °C. In order to test the biodegradation process of OTA, the chloroform was evaporated and the crystalline OTA was dissolved in 5 mL of DMSO and the dilution was made to achieve an initial OTA concentration of 5 µg mL−1 (Wei et al. 1985).
Degradation assay of OTA
T. koningii strains (AUMC11519, AUMC11520, AUMC11521 and AUMC11522) were first grown in PDA for 7 days in the dark at 30 °C for inoculum generation. The fungal strains were then grown in test tubes containing 3 mL of PDB amended with 5 µg mL−1 of OTA. Test tubes were inoculated with a dense conidial suspension of the T. koningii strains, the spore suspension was adjusted to a concentration of 106 spores mL−1 and incubated at 30 °C for 10 days in the dark. After which, the fungal cultures were filtered by Whatman No. 1 filter papers. A negative control, a culture medium-containing OTA, was used to calculate OTA removal percentage. All assays were performed in triplicate (Varga et al. 2000). OTA residues were extracted and quantified spectrophotometrically as mentioned earlier.
where C0 and C represented the initial and residual concentrations of OTA, respectively (Bejaoui et al. 2006).
Effect of EDTA on OTA degrading activity
T. koningii strains were grown separately in PD broth for 7 days at 30 °C. The cells from the 50-mL T. koningii cultures were harvested by centrifugation at 6440 ×g at 4 °C for 20 min. The supernatant was collected and processed by surface sterilization to produce a cell-free supernatant by filtering 0.22 µm filters. Thereafter, the cell-free supernatants were amended with 5 and 10 mmol−1 of EDTA. The EDTA-free broth served as a control (Zhang et al. 2017). In all of these assays, the initial concentration of OTA was 5 µg mL−1.
Fourier-transform infrared (FTIR) analysis
The filtrate of both samples and controls resulted from degradation assays processed above were analyzed by FTIR. In which, the filtrates were extracted with chloroform and the dry films were stored at − 20 °C till the analysis by FTIR which was performed at Microanalytical Center, Faculty of Science, Cairo University, Giza, Egypt. Infrared spectra of treated and non-treated OTA resulted from detoxification experiments were determined over the region 400–4000 cm−1 with Pelkin-Elmer FTIR 1650 spectrophotometer.
Liquid chromatography-mass spectroscopy/ mass spectroscopy (LC–MS/MS) analysis
To distinguish the products of OTA biodegradation by the most efficient T. koningii AUMC11521, LC–MS/MS was performed with compound-specific modification to compare the fluorescent and mass transitional chromatograms. The fluorescent detector was set to a wavelength of 333 nm for excitation and 460 nm for emission. LC–MS/MS was performed in the Regional Center for Food and Feed (RCFF), Agricultural Research Center, Giza, Egypt. The analysis was performed using Agilent 1200 series liquid chromatography system equipped with Applied Biosystems (API 4000 Qtrape) tandem mass spectrometers with electrospray ionization (ESI) interface. Agilent 1200 series liquid chromatography system was equipped with Applied Biosystems (API 4000 Qtrape) tandem mass spectrometers with electrospray ionisation (ESI) interface. Separation was performed on a C18 column ZORBAX Eclipse XDBC18 4.6 mm × 150 mm, 5 μm particle sizes. The injection volume was 25 μL. A mobile phase was at 0.3 mL/min flow rate, in which one reservoir contained 10 mM ammonium format solution in methanol: water (1:9, v/v). The ESI source was used in the positive mode, and nitrogen was used as nebulizer gas, curtain gas, heater gas and collision gas according to manufacturer’s settings; source temperature was 300 °C, ion spray potential 5500 V. Declustering potential and collision energy were optimized by using the Harvard apparatus syringe pump. The Multiple Reaction Monitoring Mode (MRM) was used in which one MRM was used for quantification and other was used for confirmation (qualifier peaks).
Statistical analysis
Data obtained were subjected to the statistical analysis of variance according to Snedecor and Cochran (1980), and means separation were done according to Duncan (1955).
Results
Production and purification of OTA from A. niger T2 (MW513392.1)
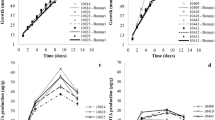
The ochratoxigenic strain, A. niger T2, was isolated from a tomato seeds sample collected from a retail shop during a survey of fungal contamination. Based on the colony appearance, morphological criteria, and conidial arrangement, the isolate was identified. It was further molecularly identified and its sequence (480 pb) has been deposited in GenBank with accession number MW513392.1. The strain was tested for the production of OTA in PDB after incubation for 10 days. The chloroform extract of the fungal filtrate showed the presence of OTA spot on TLC at Rf value = 0.89. The fungal strain was found to produce OTA in amount up to 86 µg L−1 using PDB after 10 days of incubation. After purification with column chromatography, the purified OTA using HPLC analysis gave a single peak at a retention time of 5.555 min that was identical with the standard peak (Fig. 1a, b). The recovered OTA was used later in the biodegradation studies by T. koningi strains.
OTA degrading activity of T. koningii strains
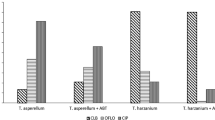
Four endophytic strains of T. koningii (AUMC11519, AUMC11520, AUMC11521 and AUMC11522) were inoculated individually in PDB spiked with OTA (5 μg mL−1) and incubated for 10 days at 30 °C in order to demonstrate the biodegradation activity. Data presented in Fig. 2 showed that three strains, AUMC11519, AUMC11520 and AUMC11521 completely eliminated OTA from the culture medium in a percent of 100%, while AUMC11522 strain eliminated only 41.82% of OTA recording significant differences (P ≤ 0.05) with the other test strains.
Biodegradation of OTA by T. koningii strains after incubation for 10 days at 30 °C under static conditions. Data was expressed in percentage of OTA removal. Values are represented as means ± SD of three replicate analyses from two independent experiments. Different letters on the bars for OTA removal (%) indicate significant differences
Effect of EDTA on OTA degrading activity
T. koningii strains were grown separately in PD broth for 7 days at 30 °C. After centrifugation, the cell-free supernatants were supplemented separately with 5 and 10 mmol−1 EDTA. The EDTA-free supernatants of the three test strains showed a total 100% effective removal of OTA. EDTA-treated supernatants showed a dramatic significant reduction in the percent of OTA removal ability (P ≤ 0.05) (Fig. 3). The 5 mmol−1 EDTA-amended supernatants of the three strains inhibited OTA degradation ability by 58.59% (AUMC11520), 34.3% (AUMC 11,521), and 32.68% (AUMC11519). Upon treatment with 10 mmol−1 EDTA, the inhibition of degradation efficiency (%) was significantly reduced by 77.57% (AUMC11520), 76.34% (AUMC11521) and 73.15% (AUMC11519).
Effect of cell-free supernatants of T. koningii strains (AUMC11519, AUMC11520, and AUMC11521) treated with 5 and 10 mmol−1 EDTA on OTA removal%. Samples were incubated at 30 °C for 7 days with 5 μg OTA mL−1. Values were represented as means ± SD of three replicate analyses from two independent experiments. Different letters on the bars for either treatment with 5 mmol−1 EDTA or 10 mmol−1 indicate significant differences
Characterization of OTA biodegradation by the most potent T. koningii strains.
FTIR
To estimate the structural changes of the biodegraded OTA, the FTIR spectra of OTA before and after incubation for 7 days at 30 °C with T. koningii AUMC11519, AUMC11520, and AUMC11521 were analyzed (Fig. 4a–d, respectively). According to FTIR spectrum of OTA treated with AUMC11521 strain (Fig. 4d), shifts at 3426.9 cm−1, 1640.2 cm−1 (∆ 6 cm−1), and 699.1 cm−1 (∆ 10 cm−1) coupled with the disappearance of peak at 1722.1 cm−1 and new peak at 612.3 cm−1 confirmed the cleavage of C=O of carboxylic acid and C=O of amide bond NH–CO and=C–H with C–Cl. FTIR spectra of OTA treated with AUMC11519 strain (Fig. 4b) and AUMC11520 strain (Fig. 4c) showed a shift at 1722.1 cm−1 (∆ 6 cm−1). New peaks at 1278.6 cm−1 (AUMC11519 strain) and 1386.6 cm−1 (AUMC11520 strain) are assigned to stretching mode of C–O (ether and carboxylic acid), C–F, NO2 (aliphatic nitro) and binding mode of C–O–H, respectively. A remarkable shift at 799.4 cm−1 (∆80 cm−1) in the case of AUMC11519 strain attributed to the bending mode of=C–H (aromatic ring), and N–H, and C–Cl stretching mode in OTA structure. From the results of FTIR spectra of T. koningii strains incubated with OTA, the AUMC11521 strain had incurred more changes in OTA degradation, compared with the other strains. Therefore, further characterization of products resulted from OTA degradation by this strain was proceeded by LC–MS/MS.
Confirmation of OTA biodegradation products by LC–MS/MS
In order to identify the biodegradation products resulted from incubation of OTA with the most efficient OTA degrading strain (T. koningii AUMC11521) compared with control OTA (before incubation), LC–MS/MS analyses were employed (Fig. 5). Based on the LC–MS/MS spectra, the detected m/z 402.9 (Fig. 5a), m/z 258.2, m/z 257.0, m/z 165.0, and m/z 150.2 (Fig. 5b) were assigned to the separated OTA, OTα-amide, OTα, L-β-phenylalanine, and 3-phenylpropanoic acid, respectively. Their chemical formula were C20H18ClNO6 (OTA), C11H12ClNO4 (OTα-amide), C11H9ClO5 (OTα), phenylalanine (C9H11NO2), and C9H10O2 (3-phenyl propanoic acid) (Table 1). These changes confirmed the two hypothesized biodegradation pathways of OTA (Fig. 6). This is in accordance with the finding of FTIR studies which confirmed the cleavage of C=O of carboxylic acid and C=O of NH–CO and=C–H with C–Cl. The degradation products with their molecular weight (MW), chemical formula, and structural formula were presented in Table 1.
Biodegradation of OTA by T. koningii AUMC11521 grown in PDB for 7 days at 30ºC. i Ochratoxinases with amido-hydrolase activity are produced to break the amide bond in OTA, releasing two non-toxic products: OTα and L-β-phenylalanine; ii Hydrolases also produce less toxic metabolites: OTα-amide and 3-phenylpropanoic acid. Arrows refer to the positions of the cleavage of the bond responsible for the breakdown of OTA molecule
Discussion
The toxic metabolites of storage molds are known as mycotoxins and the health hazards caused by them are known as mycotoxicosis. The entire problem of mycotoxins production and health hazards in humans were well documented (Klich et al. 2003; Trucksess et al. 2003). Due to OTA toxicity on human and animal health, this toxin is not allowed to be present above maximum permitted levels in agricultural products intended to be used as foods or animal feed. Substantial efforts have been exerted to study the critical points of OTA presence in food and feed commodities, and its detoxification methods have also been investigated (Abrunhosa et al. 2010). Although it is almost impossible to entirely prevent the formation of OTA in all cases, OTA accumulation can be minimized. A remarkable attention has been paid regarding demonstrating the effective methods for detoxification of mycotoxins contaminated commodities. Decontamination or detoxification is a pressing issue; this is useful in order to recondition mycotoxin contaminated agricultural products for use as animal feeds. Although certain treatments have been found to reduce levels of specific mycotoxins, however no single method has been developed that is equally effective against the wide variety of mycotoxins which may co-occur in different commodities (Abrunhosa et al. 2010). Regarding OTA, the most promising approaches included the use of microbes or their enzymes for decontamination purposes (Abrunhosa et al. 2010).
Biodegradation or biodetoxification of OTA by fungi include the application of microbes or their enzymes for decontamination purposes (Abrunhosa et al. 2010). There are two different biodegradation pathways of OTA by enzymes. First biodegradation pathway occurred through the hydrolysis of the amide bond resulting in the production of L-β-phenylalanine molecule and OTα (Fig. 7i), which are non-toxic, so that this process is considered detoxification of OTA (Abrunhosa et al. 2010; Karlovsky 1999). The second pathway, biodegradation occurred through the hydrolysis of the lactone ring, resulting in the production of open ring OTA (Fig. 7ii), which have the same toxicity as OTA, so that this process is not considered as detoxification but it is just break down of OTA molecules (Abrunhosa et al. 2010; Karlovsky 1999). There is a third hypothetical pathway that was mostly observed during thermal degradation of OTA, in which the degradation occurred through the hydrolysis of the carbonyl of carboxylic acid resulting in the formation of the non-toxic OTα-amide (Fig. 7iii), this process is also considered detoxification (Bittner et al. 2015).
The current study showed a degradation percent ranged from 41.82 ± 1.57% to 100% of OTA with T. koningii strains. Similarly, Trichoderma spp. were able to degrade more than 80% of aflatoxin B1 (Shantha 1999). These differences in the biodegradation values occurred due to the strain-specificity (Petruzzi et al. 2015). Trichoderma spp. were previously used to inhibit OTA production by different ochratoxigenic fungi and suppress the growth of these fungi by production of certain enzymes as a result of antagonism (Vankudoth et al. 2016). In literature, no reports regarding using T. koningii strains in biodegradation of OTA are found, but their OTA-degrading activities may be attributed to the ability of Trichoderma spp.to produce extracellular enzymes such as hydrolases (Patil et al. 2016), proteolytic enzymes (proteases) (Aziz et al. 1993; Schuster and Schmoll 2010), including carboxypeptidase A (CPA) (Kupski et al. 2018). Therefore, Trichoderma spp. have a major impact on the biological control potential (Viterbo et al. 2002). CPA belongs to extracellular proteases and produced by Trichoderma spp. was found to be responsible for OTA degradation (Abrunhosa et al. 2002, 2006; Abrunhosa and Venâncio 2007; Schuster and Schmoll, 2010; Kupski et al. 2018).
In order to prove that extracellular CPA is responsible for OTA degradation, the cell-free supernatants of T. koningii strains (AUMC11519, AUMC11520, and AUMC11521) were amended with 5 and 10 mmol−1 of EDTA. Results showed a dramatic reduction in the OTA elimination ability of T. koningii strains. Based on previous studies concerning enzyme inhibition data (Abrunhosa and Venâncio 2007; Zhang et al. 2017), the OTA degrading enzyme from T. koningii strains was strongly inhibited by EDTA, demonstrating that the enzyme produced by T. koningii strains is a metalloenzyme. These results strongly suggest that T. koningii strains produce an enzyme that is responsible for biodegradation of OTA through the hydrolysis of carbonyl C-14 and the nitrogen atom of amine group, which resulted in the production of OTα-amide. In agreement with our results, studies carried out by Abrunhosa et al. (2002) on other filamentous fungi including A. niger showed completely or partially degradation of OTA after growth in 1 mg L−1 OTA for 6 days at 25 °C, OTα was also detected as a degradation product in their study. The authors concluded that a carboxypeptidase may be involved in the degradation of OTA by their strains. In a previous study (Zhang et al. 2017), Alcaligenes faecalis isolated from soil samples was found to degrade OTA efficiently and OTα was confirmed as a degradation product in the intracellular extract of A. faecalis using UPLC-MS/MS. It was suggested that the biodegradation of OTA occurs through the hydrolysis of the OTA amide bond by a putative peptidase (Zhang et al. 2017). In the current study, the OTA degradation by T. koningii strains released OTα-amide, 3-phenylpropionic acid, OTα and phenylalanine, indicating breakage of the amide bond and suggesting that the possible mechanism involved in OTA degradation by T. koningii strains is enzymatic.
Enzymatic degradation of OTA has specific advantages (Zhao et al. 2020). First, the degradation products have low toxicity. Second, OTα is a non-toxic compound with a tenfold shorter half-life time in humans (Bui-Klimke and Wu 2015; Zhao et al. 2020) and is 1000 times less toxic than OTA (Rodriguez et al. 2011). Compared with adsorption, degradation has obvious advantages. Hence, the conversion processes of OTA into OTα contribute substantially to reducing the toxic effects of OTA and are considered promising routes for OTA detoxification. Additionally, the enzymatic degradation is a non-reversible process that does not cause secondary pollution. Cell wall adsorption by microorganisms such as yeast and lactic acid bacteria does not completely eliminate OTA, and there are still hidden dangers of desorption (Zhao et al. 2020).
The FTIR spectra were carried out to identify the changes occurred in OTA structure in the range of 4000–400 cm−1. The position or the intensities of peaks of OTA are expected to be changed upon this interaction. On the basis of FTIR spectra of OTA (before and after biodegradation) T. koningii AUMC11521 was found to be the most effective strain in OTA degradation. FTIR spectrum of OTA treated with this strain showed shifts at 3426.9 cm−1, 1640.2 cm−1 (∆ 6 cm−1), and 699.1 cm−1 (∆ 10 cm−1) coupled with the disappearance of peak at 1722.1 cm−1 and new peak detected at 612.3 cm−1 confirmed the cleavage of C=O of carboxylic acid and C=O of amide bond NH–CO and=C–H with C–Cl. These changes in OTA molecules are assigned to the -OH and -NH groups and hydrogen bonding in fingerprint region between 2500 cm−1 and 3800 cm−1 which corresponds to protein component (Bhat, 2013). Therefore, OTA biodegradation products by the ultraefficient strain AUMC11521 were subjected to further characterization by LC–MS/MS.
LC–MS/MS analyses of fungal OTA before and after incubation with the T. koningii AUMC11521 for 7 days at 30 °C confirmed the two enzymatic hypothesized biodegradation pathway of OTA. According to the LC–MS/MS spectra, the detected m/z m/z 402.9, m/z 258.2, m/z 257.0, m/z 165.0, and m/z 150.2 were assigned to the separated OTA, OTα-amide, OTα, L-β-phenylalanine, and 3-phenylpropanoic acid, respectively. We suggest that OTA degradation occurred by two pathways (Fig. 6i, ii). First one occurred through the hydrolysis of the amide bond (Fig. 6i) via hydrolytic enzymes, such as CPA, carboxypeptidase PJ-1540, protease A, lipase A, ochratoxinases with amido-hydrolase activity, etc. which resulted in the production of L-β-phenylalanine molecule and a non-toxic OTα (Dobritzsch et al. 2014; Liuzzi et al. 2017). The second pathway involved the breakage between C-14 and the nitrogen atom of amine group as indicated by arrow in Fig. 6ii. This pathway resulted in the formation of OTα-amide and 3-phenylpropanoic acid. This is in accordance with Bittner et al. (2015). The cleavage of the amide bond (CO–NH) and NH–CH was confirmed as mentioned from FTIR spectra results. FTIR bands found at 1631 and 1600 cm−1 are assigned to C–N stretching vibrations. The four recorded products are less and non-toxic, so that this process is considered as detoxification. There is an additional pathway, occurred through the hydrolysis of the lactone ring by an ochratoxin-lactonase, resulted in the production of open ring OTA, which has the same toxicity as OTA, so that this process is not considered as detoxification but it is just break down of OTA molecules (Leitão and Enguita 2021). It is well known that the kidney is the first target for OTA. Bittner et al. (2015) studied the toxicity of OTα-amide using immortalized human kidney epithelial cells, a cell line known to be sensitive against OTA. The authors found that OTα-amide had no toxicity up to concentrations of 50 μM. Interestingly, 3-phenyl propionic acid is extensively used in cosmetics and pharmaceuticals as well as a preservative and flavoring agent in the food industry and it has a role as an antifungal agent (Korneev 2013).
Conclusion
Overall, our endophytic strains of T. koningii (AUMC11519, AUMC11520 and AUMC11521) proved their efficient capability for degradation of OTA in vitro. The degradation products include OTα-amide, 3-phenylpropionic acid, OTα and phenylalanine which are much less toxic than OTA. The OTA degradation process by T. koningii strains is suggested to be enzymatic and CPA has a crucial role that could be used for OTA detoxification contamination in foods and food products. Furthermore, characterization of the genes that are responsible for the degradation of OTA would be a necessity.
Data availability
All relevant data are within the paper and its Supporting Information file.
References
Abrunhosa L, Venâncio A (2007) Isolation and purification of an enzyme hydrolyzing ochratoxin A from Aspergillus niger. Biotechnol Lett 29(12):1909–1914
Abrunhosa L, Serra R, Venâncio A (2002) Biodegradation of ochratoxin A by fungi isolated from grapes. J Agric Food Chem 50(25):7493–7496. https://doi.org/10.1021/jf025747i
Abrunhosa L, Santos L, Venâncio A (2006) Degradation of ochratoxin A by proteases and by a crude enzyme of Aspergillus niger. Food Biotechnol 20(3):231–242. https://doi.org/10.1080/08905430600904369
Abrunhosa L, Paterson RR, Venâncio A (2010) Biodegradation of ochratoxin A for food and feed decontamination. Toxins 2(5):1078–1099. https://doi.org/10.3390/toxins2051078
Al-Hadithi N, Kössler P, Karlovsky P (2015) Determination of ochratoxin A in wheat and maize by solid bar microextraction with liquid chromatography and fluorescence detection. Toxins 7(8):3000–3011. https://doi.org/10.3390/toxins7083000
Ali R, Ismail M, Bhalli JA, Mobeen A, Khan QM (2013) Effect of temperature on ochratoxin A production in common cereals by Aspergillus species. J Anim Plant Sci 23(5):1316–1320
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402. https://doi.org/10.1093/nar/25.17.3389
Aziz AY, Foster HA, Fairhurst CP (1993) Extracellular enzymes of Trichoderma harzianum, T. polysporum and Scytalidium lignicola in relation to biological control of Dutch Elm disease. Arboric J 17(2):159–170. https://doi.org/10.1080/03071375.1993.9746959
Bejaoui H, Mathieu F, Taillandier P, Lebrihi A (2006) Black aspergilli and ochratoxin A production in French vineyards. Int J Food Microbiol 111:S46–S52. https://doi.org/10.1016/j.ijfoodmicro.2006.03.004
Bhat R (2013) Potential use of fourier transform infrared spectroscopy for identification of molds capable of producing mycotoxins. Inter J Food Prop 16(8):1819–1829. https://doi.org/10.1080/10942912.2011.609629
Bittner A, Cramer B, Harrer H, Humpf HU (2015) Structure elucidation and in vitro cytotoxicity of ochratoxin α amide, a new degradation product of ochratoxin A. Mycotoxin Res 31(2):83–90. https://doi.org/10.1007/s12550-014-0218-y
Boudra H, Le Bars P, Le Bars J (1995) Thermostability of ochratoxin A in wheat under two moisture conditions. Appl Environ Microbiol 61(3):1156–1158. https://doi.org/10.1128/aem.61.3.1156-1158.1995
Bui-Klimke TR, Wu F (2015) Ochratoxin A and human health risk: a review of the evidence. Crit Rev Food Sci Nutr 55(13):1860–1869. https://doi.org/10.1080/10408398.2012.724480
Bullerman LB, Bianchini A (2007) Stability of mycotoxins during food processing. Inter J Food Microbiol 119(1–2):140–146. https://doi.org/10.1016/j.ijfoodmicro.2007.07.035
Daradimos E, Marcaki P, Koupparis M (2000) Evaluation and validation of two fluorometric HPLC methods for the determination of aflatoxin B1 in olive oil. Food Addit Contam 17(1):65–73. https://doi.org/10.1080/026520300283603
Dobritzsch D, Wang H, Schneider G, Yu S (2014) Structural and functional characterization of ochratoxinase, a novel mycotoxin-degrading enzyme. Biochem J 462(3):441–452. https://doi.org/10.1042/BJ20140382
Duarte SC, Pena A, Lino CM (2010) A review on ochratoxin A occurrence and effects of processing of cereal and cereal derived food products. Food Microbiol 27(2):187–198. https://doi.org/10.1016/j.fm.2009.11.016
Duarte SC, Lino CM, Pena A (2012) Food safety implications of ochratoxin A in animal-derived food products. Vet J 192(3):286–292. https://doi.org/10.1016/j.tvjl.2011.11.002
Duncan DB (1955) Multiple range and multiple F tests. Biometrics 11(1):1–42
Durguti V, Georgieva A, Angelov A, Bajrami Z (2014) Quantitative determination of ochratoxin A in wine after the clarification and filtration. Croat J Food Sci Technol 6(2):79–83
European Food Safety Authority (EFSA) (2006) Opinion of the scientific panel on contaminants in the food chain [CONTAM] related to ochratoxin A in food. EFSA J 4(6):365. https://doi.org/10.2903/j.efsa.2006.365
Faucet V, Pfohl-Leszkowicz A, Dai J, Castegnaro M, Manderville RA (2004) Evidence for covalent DNA adduction by ochratoxin A following chronic exposure to rat and subacute exposure to pig. Chem Res Toxicol 17(9):1289–1296. https://doi.org/10.1021/tx049877s
García-Cela E, Crespo-Sempere A, Ramos AJ, Sanchis V, Marin S (2014) Ecophysiological characterization of Aspergillus carbonarius, Aspergillus tubingensis and Aspergillus niger isolated from grapes in Spanish vineyards. Int J Food Microbiol 173:89–98. https://doi.org/10.1016/j.ijfoodmicro.2013.12.012
Hao J, Wu W, Wang Y, Yang Z, Liu Y, Lv Y, Xu W (2015) Arabidopsis thaliana defense response to the ochratoxin A-producing strain (Aspergillus ochraceus 3.4412). Plant Cell Rep 34(5):705–719. https://doi.org/10.1007/s00299-014-1731-30
IARC Working Group on the Evaluation of Carcinogenic Risks to Humans (1993) Ochratoxin A. In: Some Naturally Occurring Substances: Food Items and Constituents, Heterocyclic Aromatic Amines and Mycotoxins. International Agency for Research on Cancer (IARC).
Ismaiel AA, Ali DM (2017) Antimicrobial properties of 6-pentyl-α-pyrone produced by endophytic strains of Trichoderma koningii and its effect on aflatoxin B1 production. Biologia 72(12):1403–1415. https://doi.org/10.1515/biolog-2017-0173
Ismaiel AA, Papenbrock J (2015) Mycotoxins: producing fungi and mechanisms of phytotoxicity. Agriculture 5(3):492–537. https://doi.org/10.3390/agriculture5030492
Kabak B (2009) The fate of mycotoxins during thermal food processing. J Sci Food Agric 89(4):549–554. https://doi.org/10.1002/jsfa.3491
Karlovsky P (1999) Biological detoxification of fungal toxins and its use in plant breeding, feed and food production. Nat Toxins 7(1):1–23. https://doi.org/10.1002/(SICI)1522-7189(199902)7:1%3C1::AID-NT37%3E3.0.CO;2-9
Klich MA, Cary JW, Beltz SB, Bennett CA (2003) Phylogenetic and morphological analysis of Aspergillus ochraceoroseus. Mycologia 95(6):1252–1260. https://doi.org/10.1080/15572536.2004.11833033
Korneev SM (2013) Hydrocinnamic acids: application and strategy of synthesis. Synthesis 45(08):1000–1015. https://doi.org/10.1055/s-0032-1318475
Kupski L, Queiroz MI, Badiale-Furlong E (2018) Application of carboxypeptidase A to a baking process to mitigate contamination of wheat flour by ochratoxin A. Process Biochem 64:248–254. https://doi.org/10.1016/j.procbio.2017.09.006
Leitão AL, Enguita FJ (2021) Structural insights into carboxylic polyester-degrading enzymes and their functional depolymerizing neighbors. Inter j Mol Sci 22(5):2332. https://doi.org/10.3390/ijms22052332
Liuzzi VC, Fanelli F, Tristezza M, Haidukowski M, Picardi E, Manzari C, Mulè G (2017) Transcriptional analysis of Acinetobacter sp. neg1 capable of degrading ochratoxin A. Front Microbiol 7:2162. https://doi.org/10.3389/fmicb.2016.02162
Malone BR, Humphrey CW, Romer TR, Richard JL (1998) One-step solid-phase extraction cleanup and fluorometric analysis of deoxynivalenol in grains. J AOAC Int 81(2):448–452. https://doi.org/10.1093/jaoac/81.2.448
Müller HM (1982) Decontamination of mycotoxins I. Phys Process Ubersicht Tierernähr 10:95–122
Nazir KHMNH, Hassan J, Durairaj P, Yun H (2014) Isolation and identification of Aspergillus flavus from poultry feed samples using combined traditional-molecular approach and expression of CYP64A1 at mRNA level. Pak J Agri Sci 51(2):287–291
Nesheim S (1976) The ochratoxins and other related compounds. Adv Chem Seri. https://doi.org/10.1021/C1976-0149.ch012
Patil AS, Patil SR, Paikrao HM (2016) Trichoderma secondary metabolites: their biochemistry and possible role in disease management. In: Choudhary DK, Varma A (eds) Microbial-mediated induced systemic resistance in plants. Springer, Singapore, pp 69–102. https://doi.org/10.1007/978-981-10-0388-2_6
Petruzzi L, Baiano A, De Gianni A, Sinigaglia M, Corbo MR, Bevilacqua A (2015) Differential adsorption of ochratoxin A and anthocyanins by inactivated yeasts and yeast cell walls during simulation of wine aging. Toxins 7(10):4350–4365. https://doi.org/10.3390/toxins7104350
Preuss ML, Cameron JC, Berg RH, Jez JM (2014) Immunolocalization of glutathione biosynthesis enzymes in Arabidopsis thaliana. Plant Physiol Biochem 75:9–13. https://doi.org/10.1016/j.plaphy.2013.11.027
Rodriguez H, Reveron I, Doria F, Costantini A, De Las RB, Munoz R, Garcia-Moruno E (2011) Degradation of ochratoxin A by Brevibacterium species. J Agric Food Chem 59(19):10755–10760. https://doi.org/10.1021/jf203061p
Schuster A, Schmoll M (2010) Biology and biotechnology of Trichoderma. Appl Microbiol Biotechnol 87(3):787–799. https://doi.org/10.1007/s00253-010-2632-1
Shantha T (1999) Fungal degradation of aflatoxin B1. Nat Toxins 7(5):175–178. https://doi.org/10.1002/1522-7189(200009/10)7:5%3C175::AID-NT63%3E3.0.CO;2-M
Škrinjar M, Rašić JL, Stojičić V (1996) Lowering of ochratoxin A level in milk by yoghurt bacteria and bifidobacteria. Folia Microbiol 41(1):26–28. https://doi.org/10.1007/BF02816335
Snedecor GW, Cochran WG (1980) Statistical Methods Iowa State University Press, Ames. Statistical methods, 7th edn. The Iowa State University Press, Ames
Snyder L (2008) Solvent selectivity in normal-phase TLC. JPC-J Planar Chromat 21(5):315–323. https://doi.org/10.1556/jpc.21.2008.5.1
Stahl E (1969) Apparatus and general techniques in TLC. In: Stahl E (ed) Thin-layer chromatography. Springer, Berlin. https://doi.org/10.1007/978-3-642-88488-7_3
Téren J, Varga J, Hamari Z, Rinyu E, Kevei F (1996) Immunochemical detection of ochratoxin A in black Aspergillus strains. Mycopathologia 134(3):171–176. https://doi.org/10.1007/BF00436726
Trivedi AB, Doi E, Kitabatake N (1992) Detoxification of ochratoxin A on heating under acidic and alkaline conditions. Biosci Biotechnol Biochem 56(5):741–745. https://doi.org/10.1271/bbb.56.741
Trucksess MW, Avramson D, Dorner J, Eppley RM, Hagler WM, Hald B, Maragos C, Sabino M, Solfrizzo M, van Egmond HP, Ware GM, Whitaker TB, Wilson DM (2003) Committee on natural toxins and food allergens: Mycotoxins. J AOAC Int 86(1):129–138
Valenta H, Kühn I, Rohr K (1993) Determination of ochratoxin A in urine and faeces of swine by high-performance liquid chromatography. J Chromatogr B: Biomed Sci Appl 613(2):295–302. https://doi.org/10.1016/0378-4347(93)80145-T
Van der Merwe KJ, Steyn PS, Fourie L (1965a) 1304. Mycotoxins. Part II. The constitution of ochratoxins A, B, and C, metabolites of Aspergillus ochraceus wilh. J Chem Soc (resumed). https://doi.org/10.1039/JR9650007083
Van der Merwe KJ, Steyn PS, Fourie L, Scott DB, Theron JJ (1965b) Ochratoxin A, a toxic metabolite produced by Aspergillus ochraceus Wilh. Nature 205(4976):1112–1113. https://doi.org/10.1038/2051112a0
Vankudoth KR, Boda A, Sivadevuni G, Solipuram MR (2016) Effect of indigenous fungi on ochratoxin A produced by two species of Penicillium. Anim Nutr 2(3):225–228. https://doi.org/10.1016/j.aninu.2016.04.004
Varga J, Rigó K, Téren J (2000) Degradation of ochratoxin A by Aspergillus species. Int J Food Microbiol 59(1–2):1–7. https://doi.org/10.1016/S0168-1605(00)00230-0
Viterbo A, Ramot O, Chernin L, Chet I (2002) Significance of lytic enzymes from Trichoderma spp. in the biocontrol of fungal plant pathogens. Antonie Van Leeuwenhoek 81(1):549–556. https://doi.org/10.1023/A:1020553421740
Wang Y, Zhao W, Hao J, Xu W, Luo Y, Wu W, Huang K (2014) Changes in biosynthesis and metabolism of glutathione upon ochratoxin A stress in Arabidopsis thaliana. Plant Physiol Biochem 79:10–18. https://doi.org/10.1016/j.plaphy.2014.03.001
Wei YH, Lu CY, Lin TN, Wei RD (1985) Effect of ochratoxin A on rat liver mitochondrial respiration and oxidative phosphorylation. Toxicology 36(2–3):119–130. https://doi.org/10.1016/0300-483X(85)90046-0
WHO (World Health Organization) (1991) Evaluation of certain food additives and contaminants. Thirty-seventh report of the joint FAO/WHO Expert Committee on Food Additives (WHO technical report series 806), Geneva: World Health Organization. pp 29–31. https://doi.org/10.1002/food.19910350833.
Zhang HH, Wang Y, Zhao C, Wang J, Zhang XL (2017) Biodegradation of ochratoxin A by Alcaligenes faecalis isolated from soil. J Appl Microbiol 123(3):661–668. https://doi.org/10.1111/jam.13537
Zhao M, Wang XY, Xu SH, Yuan GQ, Shi XJ, Liang ZH (2020) Degradation of ochratoxin A by supernatant and ochratoxinase of Aspergillus niger W-35 isolated from cereals. World Mycotoxin J 13(2):287–298. https://doi.org/10.3920/WMJ2019.2446
Zheng MZ, Richard JL, Binder J (2006) A review of rapid methods for the analysis of mycotoxins. Mycopathologia 161(5):261–273. https://doi.org/10.1007/s11046-006-0215-6
Acknowledgements
The authors acknowledge Prof. Dr. Mohammed Gomaa Assy, Professor of Organic Chemistry, Department of Chemistry, Faculty of Science, Zagazig University, Zagazig 44519, Egypt, for critical comments concerning with illustration of FTIR and LC-MS/ MS analyses and declaration of OTA degradation products.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). This work was partially supported by Zagazig University, Egypt.
Author information
Authors and Affiliations
Contributions
Study concept and design, AAI and MTE; acquisition of data, HHM; analysis and interpretation of data, MTE and AAI; drafting of the manuscript, MTE, AAI and HHM; critical revision of the manuscript for important intellectual content, AAI and MTE; statistical analysis, HHM; administrative, technical, and material support, AAI, MTE and HHM; study supervision: AAI and MTE
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ismaiel, A.A., Mohamed, H.H. & El-Sayed, M.T. Biodegradation of ochratoxin A by endophytic Trichoderma koningii strains. World J Microbiol Biotechnol 39, 53 (2023). https://doi.org/10.1007/s11274-022-03491-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-022-03491-2