Abstract
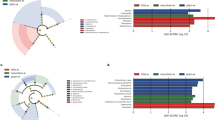
Rhizosphere and root-associated microbial communities are known to be related to soil-borne disease and plant health. In the present study, the microbial communities in rhizosphere soils and roots of both healthy and diseased Panax notoginseng were analyzed by high-throughput sequencing of 16S rRNA for bacteria and 18S rRNA internal transcribed spacer for fungi, to reveal the relationship of microbial community structure with plant health status. In total, 5593 bacterial operational taxonomic units (OTUs) and 963 fungal OTUs were identified in rhizosphere soils, while 1794 bacterial and 314 fungal OTUs were identified from root samples respectively. Principal coordinate analysis separated the microbial communities both in the rhizosphere soils and roots of diseased P. notoginseng from healthy plants. Compared to those of healthy P. notoginseng, microbial communities in rhizosphere soils and roots of diseased plants showed a decrease in alpha diversity. By contrast, bacterial community dissimilarity increased and fungal community dissimilarity decreased in rhizosphere soils of diseased plants, while both bacterial and fungal community dissimilarity in roots showed no significant difference between healthy and diseased plants. Redundancy analysis at the phylum level showed that mycorrhizal colonization and soil texture significantly affected microbial community composition in rhizosphere soils, whereas shoot nutrition status had a significant effect on microbial community composition in root samples. Our study provided strong evidence for the hypothesis that microbial diversity could potentially serve as an indicator for disease outbreak of medicinal plants, and supported the ecological significance of microbial communities in maintaining plant healthy and soil fertility.





Similar content being viewed by others
References
Abarenkov K, Henrik Nilsson R, Larsson KH, Alexander IJ, Eberhardt U, Erland S, Høiland K, Kjøller R, Larsson E, PennanenT Sen R, Taylor AF, Tedersoo L, Ursing BM, Vrälstad T, Liimatainen K, Peintner U, Kõljalg U (2010) The UNITE database for molecular identification of fungi-recent updates and future perspectives. New Phytol 186(2):281–285
Acosta-Martinez V, Dowd S, Sun Y, Allen V (2008) Tag-encoded pyrosequencing analysis of bacterial diversity in a single soil type as affected by management and land use. Soil Biol Biochem 40(11):2762–2770
Al-Karaki GN (2006) Nursery inoculation of tomato with arbuscular mycorrhizal fungi and subsequent performance under irrigation with saline water. Sci Hortic 109(1):1–7
Bailey KL, Lazarovits G (2003) Suppressing soil-borne diseases with residue management and organic amendments. Soil Tillage Res 72(2):169–180
Bakker MG, Glover JD, Mai JG, Kinkel LL (2010) Plant community effects on the diversity and pathogen suppressive activity of soil streptomycetes. Appl Soil Ecol 46(1):35–42
Bergmann GT, Bates ST, Eilers KG, Lauber CL, Caporaso JG, Walters WA, Knight R, Fierer N (2011) The under-recognized dominance of Verrucomicrobia in soil bacterial communities. Soil Biol Biochem 43(7):1450–1455
Beyer L, Wachendorf C, Balzer FM, Balzer-Graf UR (1992) The effect of soil texture and soil management on microbial biomass and soil enzyme activities in arable soils of Northwest Germany. Agribiol Res (Germany) 45:276–283
Brimecombe MJ, De Leij FA, Lynch JM (2001) The effect of root exudates on rhizosphere microbial populations. In: Pinton R, Varanini Z, Nannipieri P (eds) The rhizosphere biochemistry and organic substances at the soil-plant interface. Marcel Dekker, New York, pp 95–140
Brussaard L, De Ruiter PC, Brown GG (2007) Soil biodiversity for agricultural sustainability. Agric Ecosyst Environ 121(3):233–244
Buee M, Reich M, Murat C, Morin E, Nilsson RH, Uroz S, Martin F (2009) 454 Pyrosequencing analyses of forest soils reveal an unexpectedly high fungal diversity. New Phytol 184(2):449–456
Bulluck LR III, Ristaino JB (2002) Effect of synthetic and organic soil fertility amendments on southern blight, soil microbial communities, and yield of processing tomatoes. Phytopathology 92(2):181–189
Cabral A, Groenewald JZ, Rego C, Oliveira H, Crous PW (2012) Cylindrocarpon root rot: multi-gene analysis reveals novel species within the Ilyonectria radicicola species complex. Mycol Prog 11(3):655–688
Clarke KR (1993) Non-parametric multivariate analyses of changes in community structure. Aust J Ecol 18(1):117–143
Clarke KR, Warwick RM (2001) Change in marine communities: an approach to statistical analysis and interpretation, 2nd edn. PRIMER-E, Plymouth
Cornfield AH (1960) Ammonia released on treating soils with N sodium hydroxide as a possible means of predicting the nitrogen-supplying power of soils. Nature 187:260–261
Davies TJ, Pedersen AB (2008) Phylogeny and geography predict pathogen community similarity in wild primates and humans. Proc R Soc B 275(1643):1695–1701
Donegan KK, Schaller DL, Stone JK, Ganio LM, Reed G, Hamm PB, Seidler RJ (1996) Microbial populations, fungal species diversity and plant pathogen levels in field plots of potato plants expressing the Bacillus thuringiensis var. tenebrionis endotoxin. Transgenic Res 5(1):25–35
Droux M (2004) Sulfur assimilation and the role of sulfur in plant metabolism: a survey. Photosynth Res 79(3):331–348
Gamliel A, Austerweil M, Kritzman G (2000) Non-chemical approach to soilborne pest management-organic amendments. Crop Prot 19(8):847–853
Griffiths BS, Ritz K, Wheatley R, Kuan HL, Boag B, Christensen S, Ekelund F, Sorensen SJ, Muller S, Bloem J (2001) An examination of the biodiversity-ecosystem function relationship in arable soil microbial communities. Soil Biol Biochem 33:1713–1722
Guo RY, Chen Q, Li XL (2005) The influence of soil microorganism community on the soil healthy and disease suppressiveness. China Veg 138:78–82
Guo HB, Cui XM, An N, Cai GP (2010) Sanchi ginseng (Panax notoginseng (Burkill) FH Chen) in China: distribution, cultivation and variations. Genet Resour Crop Evol 57(3):453–460
Gyaneshwar P, Kumar GN, Parekh LJ, Poole PS (2002) Role of soil microorganisms in improving P nutrition of plants. Food security in nutrient-stressed environments: exploiting plants’ genetic capabilities. Springer, Netherlands, pp 133–143
Hao X, Jiang R, Chen T (2011) Clustering 16S rRNA for OTU prediction: a method of unsupervised Bayesian clustering. Bioinformatics 27:611–618
Hao Z, Fayolle L, van Tuinen D, Chatagnier O, Li X, Gianinazzi S, Gianinazzi-Pearson V (2012) Local and systemic mycorrhiza-induced protection against the ectoparasitic nematode Xiphinema index involves priming of defence gene responses in grapevine. J Exp Bot 63(10):3657–3672
Henrik Nilsson R, Tedersoo L, Lindahl BD, Kjøller R, Carlsen T, Quince C, Abarenkov K, Pennanen T, Stenlid J, Bruns T, Larsson KH, Kõljalg U, Kauserud H (2011) Towards standardization of the description and publication of next-generation sequencing datasets of fungal communities. New Phytol 191(2):314–318
Jeffries P, Gianinazzi S, Perotto S, Turnau K, Barea JM (2003) The contribution of arbuscular mycorrhizal fungi in sustainable maintenance of plant health and soil fertility. Biol Fertil Soils 37(1):1–16
Lauber CL, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75(15):5111–5120
Li T, Hu YJ, Hao ZP, Li H, Wang YS, Chen BD (2013) First cloning and characterization of two functional aquaporin genes from an arbuscular mycorrhizal fungus Glomus intraradices. New Phytol 197(2):617–630
Lugtenberg B, Kamilova F (2009) Plant-growth promoting rhizobacteria. Annu Rev Microbiol 63:541–556
Ma L, Cao YH, Cheng MH, Huang Y, Mo MH, Wang Y, Yang JZ, Yang FX (2013) Phylogenetic diversity of bacterial endophytes of Panax notoginseng with antagonistic characteristics towards pathogens of root-rot disease complex. Antonie Van Leeuwenhoek 103(2):299–312
Mao ZS, Long YJ, Zhu YY, Zhu SS, He XH, Chen ZJ (2014) First report of Cylindrocarpon destructans var. destructans causing black root rot of sanqi (Panax notoginseng) in China. Plant Dis 98(1):162
McLean EO, Watson ME (1985) Soil measurements of plant-available potassium. In: Munson RD (ed) Potassium in agriculture. CSSA SSSA, Madison, pp 277–308
Miao ZQ, Li SD, Liu XZ, Chen YJ, Li YH, Wang Y, Guo RJ, Xia ZY, Zhang KQ (2006) The causal microorganisms of Panax notoginseng root rot disease. Sci Agric Sin 39(7):1371–1378
Nitta T (1991) Diversity of root fungal floras: its implications for soil-borne diseases and crop growth. Jpn Agric Res Q 25(1):6–11
O'Brien HE, Parrent JL, Jackson JA, Moncalvo JM, and Vilgalys R (2005) Fungal community analysis by large-scale sequencing of environmental samples. Appl Environ Microbiol 71(9):5544–5550
Oksanen J (2011) Multivariate analysis of ecological communities in R: vegan tutorial. R package version 1(7)
Olden JD, Poff NL, Douglas MR, Douglas ME, Fausch KD (2004) Ecological and evolutionary consequences of biotic homogenization. Trends Ecol Evol 19(1):18–24
Page AL, Miller RH, Keeney DR (1982) Total carbon, organic carbon and organic matter. In: American Society of Agronomy (ed) Methods of soil analysis. part 2, agronomy. American Society of Agronomy, Madison, pp 539–579
Phillips JM, Hayman DS (1970) Improved procedures for clearing roots and staining parasitic and vesicular-arbuscular mycorrhizal fungi for rapid assessment of infection. Trans Br Mycol Soc 55(1):158
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J, Glöckner FO (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35(21):7188–7196
Qiu M, Zhang R, Xue C, Zhang S, Li S, Zhang N, Shen Q (2012) Application of bio-organic fertilizer can control Fusarium wilt of cucumber plants by regulating microbial community of rhizosphere soil. Biol Fertil Soils 48(7):807–816
Rajala T, Peltoniemi M, Pennanen T, Mäkipää R (2012) Fungal community dynamics in relation to substrate quality of decaying Norway spruce (Picea abies [L.] Karst.) logs in boreal forests. FEMS Microbiol Ecol 81(2):494–505
Rescia AJ, Schmitz MF, Martin de Agar P, Pablo CL, Atauri JA, Pineda FD (1994) Influence of landscape complexity and land management on woody plant diversity in northern Spain. J Veg Sci 5(4):505–516
Rodrigues JL, Pellizari VH, Mueller R, Baek K, Jesus EDC, Paula FS, Mirza B, Hamaoli GS, Tsai SM, Feigl B, Tiedje JM, Bohannan BJM, Nüsslein K (2013) Conversion of the Amazon rainforest to agriculture results in biotic homogenization of soil bacterial communities. Proc Natl Acad Sci 110(3):988–993
Roesch LF, Fulthorpe RR, Riva A, Casella G, Hadwin AK, Kent AD, Daroub SH, Camargo FA, Farmerie WG, Triplett EW (2007) Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1(4):283–290
Sanguin H, Sarniguet A, Gazengel K, Moënne-Loccoz Y, Grundmann GL (2009) Rhizosphere bacterial communities associated with disease suppressiveness stages of take-all decline in wheat monoculture. New Phytol 184(3):694–707
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75(23):7537–7541
Stewart CN, Via LE (1993) A rapid CTAB DNA isolation technique useful for RAPD fingerprinting and other PCR applications. Biotechniques 14(5):748–750
Sun HX, Qin F, Ye YP (2005) Relationship between haemolytic and adjuvant activity and structure of protopanaxadiol-type saponins from the roots of Panax notoginseng. Vaccine 23(48):5533–5542
Teixeira LC, Peixoto RS, Cury JC, Sul WJ, Pellizari VH, Tiedje J, Rosado AS (2010) Bacterial diversity in rhizosphere soil from Antarctic vascular plants of Admiralty Bay, maritime Antarctica. ISME J 4(8):989–1001
Van Bruggen AHC, Semenov AM (1999) A new approach to the search for indicators of root disease suppression. Australas Plant Pathol 28:4–10
Van Elsas JD, Garbeva P, Salles J (2002) Effects of agronomical measures on the microbial diversity of soils as related to the suppression of soil-borne plant pathogens. Biodegradation 13(1):29–40
van Elsas JD, Jansson JK, Trevors JT (2007) Modern soil microbiology. CRC Press, Boca Raton
Vigo C, Norman JR, Hooker JE (2000) Biocontrol of the pathogen Phytophthora parasitica by arbuscular mycorrhizal fungi is a consequence of effects on infection loci. Plant Pathol 49(4):509–514
Vitale A, Aiello D, Guarnaccia V, Perrone G, Stea G, Polizzi G (2012) First report of root rot caused by Ilyonectria (=Neonectria) macrodidyma on avocado (Persea americana) in Italy. J Phytopathol 160(3):156–159
Woolson RF, Clarke WR (2011) Statistical methods for the analysis of biomedical data. Wiley, New York
Zeller V, Bardgett RD, Tappeiner U (2001) Site and management effects on soil microbial properties of subalpine meadows: a study of land abandonment along a north-south gradient in the European Alps. Soil Biol Biochem 33(4):639–649
Acknowledgments
The study was financially supported by National Natural Science Foundation of China (41101245) and National Key Technology R&D Program of the Ministry of Science and Technology (2012BAI29B02). We are grateful to the Prof. Yong Wang in Wenshan Sanqi Institute of Wenshan University for identifying the root-rot disease of P. notoginseng.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wu, Z., Hao, Z., Zeng, Y. et al. Molecular characterization of microbial communities in the rhizosphere soils and roots of diseased and healthy Panax notoginseng . Antonie van Leeuwenhoek 108, 1059–1074 (2015). https://doi.org/10.1007/s10482-015-0560-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-015-0560-x