Abstract
Main conclusion
The foxtail millet NAC transcription factor NAC1, an ortholog of Arabidopsis NAP, is induced by ABA and senescence and accelerates leaf senescence by promoting ABA biosynthesis.
Leaf senescence, a unique developmental stage involving macromolecule degradation and nutrient remobilization, is finely tuned and tightly controlled by different gene families. NO APICAL MERISTEM, ARABIDOPSIS ATAF1, and CUP-SHAPED COTYLEDON (NAC) transcription factors have been demonstrated to be involved in the modulation of leaf senescence in many land plant species. Foxtail millet (Setaria italica L.), an important food and fodder crop, has been studied for its strong stress tolerance and potential to be a biofuel model plant. However, the functional roles of senescence-associated NACs in foxtail millet are still unknown. In this study, we characterized a nuclear localized NAC transcription factor, SiNAC1, which is induced by senescence and concentrated in senescent leaves in foxtail millet. SiNAC1 also positively responds to abscisic acid (ABA) treatment in foxtail millet. Moreover, SiNAC1 promotes the natural and dark-induced leaf senescence by an ABA-dependent manner in Arabidopsis thaliana. NCED2 and NCED3 are elevated by SiNAC1 overexpression, which subsequently promotes ABA biosynthesis in Arabidopsis. Finally, as a homolog of AtNAP, SiNAC1 can partially rescue the delayed leaf senescence phenotype in atnap mutants. Overall, our results demonstrate that SiNAC1 functions as a positive regulator of leaf senescence and is involved in a positive feedback loop via ABA biosynthesis and leaf senescence.







Similar content being viewed by others
Abbreviations
- AAO3:
-
ABSCISIC ALDEHYDE OXIDASE 3
- NAP:
-
NAC-LIKE, ACTIVATED BY AP3/PI
- NCED:
-
9-Cis-epoxycarotenoid dioxygenase
- SAGs:
-
Senescence-associated genes
- senTFs:
-
Senescence-associated transcriptional factors
- TRR:
-
Transcription regulatory region
References
Austin DF (2006) Fox-tail millets (Setaria: Poaceae): Abandoned food in two hemispheres. Econ Bot 60:143–158
Balazadeh S, Riano-Pachon DM, Mueller-Roeber B (2008) Transcription factors regulating leaf senescence in Arabidopsis thaliana. Plant Biol 10(Suppl 1):63–75. doi:10.1111/j.1438-8677.2008.00088.x
Buchanan-Wollaston V, Earl S, Harrison E, Mathas E, Navabpour S, Page T, Pink D (2003) The molecular analysis of leaf senescence—a genomics approach. Plant Biotechnol J 1:3–22. doi:10.1046/j.1467-7652.2003.00004.x
Chen Q, Wang Q, Xiong L, Lou Z (2011a) A structural view of the conserved domain of rice stress-responsive NAC1. Protein Cell 2:55–63. doi:10.1007/s13238-011-1010-9
Chen Y, Qiu K, Kuai B, Ding Y (2011b) Identification of an NAP-like transcription factor BeNAC1 regulating leaf senescence in bamboo (Bambusa emeiensis ‘Viridiflavus’). Physiol Plant 142:361–371. doi:10.1111/j.1399-3054.2011.01472.x
Dong T, Park Y, Hwang I (2015) Abscisic acid: biosynthesis, inactivation, homoeostasis and signalling. Essays Biochem 58:29–48. doi:10.1042/bse0580029
Ernst HA, Olsen AN, Larsen S, Lo Leggio L (2004) Structure of the conserved domain of ANAC, a member of the NAC family of transcription factors. EMBO Rep 5:297–303. doi:10.1038/sj.embor.7400093
Fan K, Bibi N, Gan S, Li F, Yuan S, Ni M, Wang M, Shen H, Wang X (2015) A novel NAP member GhNAP is involved in leaf senescence in Gossypium hirsutum. J Exp Bot 66:4669–4682. doi:10.1093/jxb/erv240
Feng ZJ, Xu ZS, Sun J, Li LC, Chen M, Yang GX, He GY, Ma YZ (2016) Investigation of the ASR family in foxtail millet and the role of ASR1 in drought/oxidative stress tolerance. Plant Cell Rep 35:115–128. doi:10.1007/s00299-015-1873-y
Gao S, Gao J, Zhu X, Song Y, Li Z, Ren G, Zhou X, Kuai B (2016) ABF2, ABF3, and ABF4 promote ABA-mediated chlorophyll degradation and leaf senescence by transcriptional activation of chlorophyll catabolic genes and senescence-associated genes in Arabidopsis. Mol Plant 9:1272–1285. doi:10.1016/j.molp.2016.06.006
Gepstein S, Thimann KV (1980) Changes in the abscisic acid content of oat leaves during senescence. Proc Natl Acad Sci USA 77:2050–2053
Guinn G (1982) Fruit age and changes in abscisic acid content, ethylene production, and abscission rate of cotton fruits. Plant Physiol 69:349–352
Guo Y, Gan S (2006) AtNAP, a NAC family transcription factor, has an important role in leaf senescence. Plant J 46:601–612. doi:10.1111/j.1365-313X.2006.02723.x
Guo Y, Gan SS (2012) Convergence and divergence in gene expression profiles induced by leaf senescence and 27 senescence-promoting hormonal, pathological and environmental stress treatments. Plant, Cell Environ 35:644–655. doi:10.1111/j.1365-3040.2011.02442.x
Hao YJ, Song QX, Chen HW, Zou HF, Wei W, Kang XS, Ma B, Zhang WK, Zhang JS, Chen SY (2010) Plant NAC-type transcription factor proteins contain a NARD domain for repression of transcriptional activation. Planta 232:1033–1043. doi:10.1007/s00425-010-1238-2
He Y, Gan S (2002) A gene encoding an acyl hydrolase is involved in leaf senescence in Arabidopsis. Plant Cell 14:805–815
Hu R, Qi G, Kong Y, Kong D, Gao Q, Zhou G (2010) Comprehensive analysis of NAC domain transcription factor gene family in Populus trichocarpa. BMC Plant Biol 10:145–167. doi:10.1186/1471-2229-10-145
Jensen MK, Kjaersgaard T, Nielsen MM, Galberg P, Petersen K, O’Shea C, Skriver K (2010) The Arabidopsis thaliana NAC transcription factor family: structure-function relationships and determinants of ANAC019 stress signalling. Biochem J 426:183–196. doi:10.1042/BJ20091234
Jibran R, Hunter DA, Dijkwel PP (2013) Hormonal regulation of leaf senescence through integration of developmental and stress signals. Plant Mol Biol 82:547–561. doi:10.1007/s11103-013-0043-2
Kalivas A, Pasentsis K, Argiriou A, Tsaftaris AS (2010) Isolation, characterization, and expression analysis of an NAP-Like cDNA from crocus (Crocus sativus L.). Plant Mol Biol Rep 28:654–663. doi:10.1007/s11105-010-0197-x
Khan M, Rozhon W, Poppenberger B (2014) The role of hormones in the aging of plants—a mini-review. Gerontology 60:49–55. doi:10.1159/000354334
Khanna-Chopra R (2012) Leaf senescence and abiotic stresses share reactive oxygen species-mediated chloroplast degradation. Protoplasma 249:469–481. doi:10.1007/s00709-011-0308-z
Kim JH, Woo HR, Kim J, Lim PO, Lee IC, Choi SH, Hwang D, Nam HG (2009) Trifurcate feed-forward regulation of age-dependent cell death involving miR164 in Arabidopsis. Science 323:1053–1057. doi:10.1126/science.1166386
Kim J, Chang C, Tucker ML (2015) To grow old: regulatory role of ethylene and jasmonic acid in senescence. Front Plant Sci 6:20. doi:10.3389/fpls.2015.00020
Kim HJ, Nam HG, Lim PO (2016) Regulatory network of NAC transcription factors in leaf senescence. Curr Opin Plant Biol 33:48–56. doi:10.1016/j.pbi.2016.06.002
Koyama T (2014) The roles of ethylene and transcription factors in the regulation of onset of leaf senescence. Front Plant Sci 5:650. doi:10.3389/fpls.2014.00650
Kusaba M, Tanaka A, Tanaka R (2013) Stay-green plants: what do they tell us about the molecular mechanism of leaf senescence. Photosynth Res 117:221–234. doi:10.1007/s11120-013-9862-x
Lata C, Gupta S, Prasad M (2013) Foxtail millet: a model crop for genetic and genomic studies in bioenergy grasses. Crit Rev Biotechnol 33:328–343. doi:10.3109/07388551.2012.716809
Le DT, Nishiyama R, Watanabe Y, Mochida K, Yamaguchi-Shinozaki K, Shinozaki K, Tran LS (2011) Genome-wide survey and expression analysis of the plant-specific NAC transcription factor family in soybean during development and dehydration stress. DNA Res 18:263–276. doi:10.1093/dnares/dsr015
Lee IC, Hong SW, Whang SS, Lim PO, Nam HG, Koo JC (2011) Age-dependent action of an ABA-inducible receptor kinase, RPK1, as a positive regulator of senescence in Arabidopsis leaves. Plant Cell Physiol 52:651–662. doi:10.1093/pcp/pcr026
Lescot M (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327. doi:10.1093/nar/30.1.325
Li Z, Peng J, Wen X, Guo H (2013) Ethylene-insensitive3 is a senescence-associated gene that accelerates age-dependent leaf senescence by directly repressing miR164 transcription in Arabidopsis. Plant Cell 25:3311–3328. doi:10.1105/tpc.113.113340
Li Z, Zhao Y, Liu X, Peng J, Guo H, Luo J (2014) LSD 2.0: an update of the leaf senescence database. Nucleic Acids Res 42:D1200–D1205. doi:10.1093/nar/gkt1061
Li Z, Zhao Y, Liu X, Jiang Z, Peng J, Jin J, Guo H, Luo J (2017) Construction of the leaf senescence database and functional assessment of senescence-associated genes. Methods Mol Biol 1533:315–333. doi:10.1007/978-1-4939-6658-5_19
Liang C, Wang Y, Zhu Y, Tang J, Hu B, Liu L, Ou S, Wu H, Sun X, Chu J, Chu C (2014) OsNAP connects abscisic acid and leaf senescence by fine-tuning abscisic acid biosynthesis and directly targeting senescence-associated genes in rice. Proc Natl Acad Sci USA 111:10013–10018. doi:10.1073/pnas.1321568111
Lim PO, Kim HJ, Nam HG (2007) Leaf senescence. Annu Rev Plant Biol 58:115–136. doi:10.1146/annurev.arplant.57.032905.105316
Liu N, Xie K, Jia Q, Zhao J, Chen T, Li H, Wei X, Diao X, Hong Y, Liu Y (2016) Foxtail Mosaic Virus-induced gene silencing in monocot plants. Plant Physiol 171:1801–1807. doi:10.1104/pp.16.00010
Lu H, Zhang J, Liu KB, Wu N, Li Y, Zhou K, Ye M, Zhang T, Zhang H, Yang X, Shen L, Xu D, Li Q (2009) Earliest domestication of common millet (Panicum miliaceum) in east Asia extended to 10,000 years ago. Proc Natl Acad Sci USA 106:7367–7372. doi:10.1073/pnas.0900158106
Mao C, Lu S, Lv B, Zhang B, Shen J, He J, Luo L, Xi D, Chen X, Ming F (2017) A rice NAC transcription factor promotes leaf senescence via ABA biosynthesis. Plant Physiol 174:1747–1763. doi:10.1104/pp.17.00542
Nambara E, Marion-Poll A (2005) Abscisic acid biosynthesis and catabolism. Annu Rev Plant Biol 56:165–185. doi:10.1146/annurev.arplant.56.032604.144046
Noodén LD (1988) The phenomena of senescence and aging. In: Nooden LD (ed) Senescence and aging in plants. Elsevier Academic Press, San Diego, pp 1–50
Nuruzzaman M, Manimekalai R, Sharoni AM, Satoh K, Kondoh H, Ooka H, Kikuchi S (2010) Genome-wide analysis of NAC transcription factor family in rice. Gene 465:30–44. doi:10.1016/j.gene.2010.06.008
Oka M, Shimoda Y, Sato N, Inoue J, Yamazaki T, Shimomura N, Fujiyama H (2012) Abscisic acid substantially inhibits senescence of cucumber plants (Cucumis sativus) grown under low nitrogen conditions. J Plant Physiol 169:789–796. doi:10.1016/j.jplph.2012.02.001
Olsen AN, Ernst HA, Leggio LL, Skriver K (2005) NAC transcription factors: structurally distinct, functionally diverse. Trends Plant Sci 10:79–87. doi:10.1016/j.tplants.2004.12.010
O’Shea C, Kryger M, Stender EG, Kragelund BB, Willemoes M, Skriver K (2015) Protein intrinsic disorder in Arabidopsis NAC transcription factors: transcriptional activation by ANAC013 and ANAC046 and their interactions with RCD1. Biochem J 465:281–294. doi:10.1042/BJ20141045
Pourtau N, Mares M, Purdy S, Quentin N, Ruel A, Wingler A (2004) Interactions of abscisic acid and sugar signalling in the regulation of leaf senescence. Planta 219:765–772. doi:10.1007/s00425-004-1279-5
Puranik S, Bahadur RP, Srivastava PS, Prasad M (2011) Molecular cloning and characterization of a membrane associated NAC family gene, SiNAC from foxtail millet [Setaria italica (L.) P. Beauv]. Mol Biotechnol 49:138–150. doi:10.1007/s12033-011-9385-7
Puranik S, Sahu PP, Srivastava PS, Prasad M (2012) NAC proteins: regulation and role in stress tolerance. Trends Plant Sci 17:369–381. doi:10.1016/j.tplants.2012.02.004
Puranik S, Sahu PP, Mandal SN, Parida SK, Prasad M (2013) Comprehensive genome-wide survey, genomic constitution and expression profiling of the NAC transcription factor family in foxtail millet (Setaria italica L.). PLoS One 8:e64594. doi:10.1371/journal.pone.0064594
Qiu K, Li Z, Yang Z, Chen J, Wu S, Zhu X, Gao S, Gao J, Ren G, Kuai B, Zhou X (2015) EIN3 and ORE1 accelerate degreening during ethylene-mediated leaf senescence by directly activating chlorophyll catabolic genes in Arabidopsis. PLoS Genet 11:e1005399. doi:10.1371/journal.pgen.1005399
Riov J, Dagan E, Goren R, Yang SF (1990) Characterization of abscisic acid-induced ethylene production in citrus leaf and tomato fruit tissues. Plant Physiol 92:48–53
Ruberti C, Barizza E, Bodner M, La Rocca N, De Michele R, Carimi F, Lo Schiavo F, Zottini M (2014) Mitochondria change dynamics and morphology during grapevine leaf senescence. PLoS One 9:e102012. doi:10.1371/journal.pone.0102012
Rushton PJ, Bokowiec MT, Han S, Zhang H, Brannock JF, Chen X, Laudeman TW, Timko MP (2008) Tobacco transcription factors: novel insights into transcriptional regulation in the Solanaceae. Plant Physiol 147:280–295. doi:10.1104/pp.107.114041
Sakuraba Y, Jeong J, Kang MY, Kim J, Paek NC, Choi G (2014) Phytochrome-interacting transcription factors PIF4 and PIF5 induce leaf senescence in Arabidopsis. Nat Commun 5:4636. doi:10.1038/ncomms5636
Schippers JH (2015) Transcriptional networks in leaf senescence. Curr Opin Plant Biol 27:77–83. doi:10.1016/j.pbi.2015.06.018
Shen H, Yin YB, Chen F, Xu Y, Dixon RA (2009) A bioinformatic analysis of NAC genes for plant cell wall development in relation to lignocellulosic bioenergy production. Bioenergy Res 2:217–232. doi:10.1007/s12155-009-9047-9
Sun L, Sun Y, Zhang M, Wang L, Ren J, Cui M, Wang Y, Ji K, Li P, Li Q, Chen P, Dai S, Duan C, Wu Y, Leng P (2012) Suppression of 9-cis-epoxycarotenoid dioxygenase, which encodes a key enzyme in abscisic acid biosynthesis, alters fruit texture in transgenic tomato. Plant Physiol 158:283–298. doi:10.1104/pp.111.186866
Tang Y, Liu M, Gao S, Zhang Z, Zhao X, Zhao C, Zhang F, Chen X (2012) Molecular characterization of novel TaNAC genes in wheat and overexpression of TaNAC2a confers drought tolerance in tobacco. Physiol Plant 144:210–224. doi:10.1111/j.1399-3054.2011.01539.x
Uauy C, Distelfeld A, Fahima T, Blechl A, Dubcovsky J (2006) A NAC gene regulating senescence improves grain protein, zinc, and iron content in wheat. Science 314:1298–1301. doi:10.1126/science.1133649
Wang D, Gao Z, Du P, Xiao W, Tan Q, Chen X, Li L, Gao D (2015) Expression of ABA metabolism-related genes suggests similarities and differences between seed dormancy and bud dormancy of peach (Prunus persica). Front Plant Sci 6:1248. doi:10.3389/fpls.2015.01248
Weaver LM, Gan S, Quirino B, Amasino RM (1998) A comparison of the expression patterns of several senescence-associated genes in response to stress and hormone treatment. Plant Mol Biol 37:455–469
Wu A, Allu AD, Garapati P, Siddiqui H, Dortay H, Zanor MI, Asensi-Fabado MA, Munne-Bosch S, Antonio C, Tohge T, Fernie AR, Kaufmann K, Xue GP, Mueller-Roeber B, Balazadeh S (2012) JUNGBRUNNEN1, a reactive oxygen species-responsive NAC transcription factor, regulates longevity in Arabidopsis. Plant Cell 24:482–506. doi:10.1105/tpc.111.090894
Yang SD, Seo PJ, Yoon HK, Park CM (2011) The Arabidopsis NAC transcription factor VNI2 integrates abscisic acid signals into leaf senescence via the COR/RD genes. Plant Cell 23:2155–2168. doi:10.1105/tpc.111.084913
Yang J, Worley E, Udvardi M (2014) A NAP-AAO3 regulatory module promotes chlorophyll degradation via ABA biosynthesis in Arabidopsis leaves. Plant Cell 26:4862–4874. doi:10.1105/tpc.114.133769
Zhang K, Gan SS (2012) An abscisic acid-AtNAP transcription factor-SAG113 protein phosphatase 2C regulatory chain for controlling dehydration in senescing Arabidopsis leaves. Plant Physiol 158:961–969. doi:10.1104/pp.111.190876
Zhang H, Zhou C (2013) Signal transduction in leaf senescence. Plant Mol Biol 82:539–545. doi:10.1007/s11103-012-9980-4
Zhang G, Liu X, Quan Z, Cheng S, Xu X, Pan S, Xie M, Zeng P, Yue Z, Wang W, Tao Y, Bian C, Han C, Xia Q, Peng X, Cao R, Yang X, Zhan D, Hu J, Zhang Y, Li H, Li H, Li N, Wang J, Wang C, Wang R, Guo T, Cai Y, Liu C, Xiang H, Shi Q, Huang P, Chen Q, Li Y, Wang J, Zhao Z, Wang J (2012) Genome sequence of foxtail millet (Setaria italica) provides insights into grass evolution and biofuel potential. Nat Biotechnol 30:549–554. doi:10.1038/nbt.2195
Zhao Y, Chan Z, Gao J, Xing L, Cao M, Yu C, Hu Y, You J, Shi H, Zhu Y, Gong Y, Mu Z, Wang H, Deng X, Wang P, Bressan RA, Zhu JK (2016) ABA receptor PYL9 promotes drought resistance and leaf senescence. Proc Natl Acad Sci USA 113:1949–1954. doi:10.1073/pnas.1522840113
Acknowledgements
Arabidopsis T-DNA insertion mutants were provided by ABRC. Seeds of foxtail millet were obtained from Dr. Xianmin Diao (Institute of Crop Sciences, Chinese Academy of Agricultural Sciences, Beijing, China). This work was supported by the grants from the National Science Foundation of China (Grant no. 31671694 to CZ, and Grant no. 31600227 to GW), the Natural Science Fund for Distinguished Young Scholars of Hebei Province (Grant no. 2014C2014205165), the Support Program for Hundreds of Outstanding Innovative Talents in Higher Education Institutions of Hebei Province (III) (Grant no. SLRC2017043) to CZ, and the Youth Foundation of Hebei Education Department (Grant no. QN2015215) to GW.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2017_2770_MOESM1_ESM.pptx
Supplementary material 1 (PPTX 285 kb) Fig. S1 Amino acid analysis between AtNAP and five SiNACs. a Protein sequence alignment of AtNAP and five SiNACs using DNAMAN. b Model of five NAC sub-domain, nuclear localization sequence (NLS) and cis-elements in SiNAC1 coding sequence region and promoter region
425_2017_2770_MOESM2_ESM.pptx
Supplementary material 2 (PPTX 838 kb) Fig. S2 Phylogenetic analysis of SiNACs with other genes related to leaf senescence. The phylogenetic tree was constructed using MEGA based on the multiple alignments of 35 NAP protein sequences. Si, Setaria italic; Os, Oryza sativa; Ta, Triticum aestivum; Hv, Hordeum vulgare; Bd, Brachypodium distachyon; Zm, Zea mays; At, Arabidopsis thaliana
425_2017_2770_MOESM3_ESM.pptx
Supplementary material 3 (PPTX 604 kb) Fig. S3 Expression of SiNAC2, SiNAC3, SiNAC4, SiNAC5 in four developmental stages, different tissues and different senescing parts in one leaf in foxtail millet. Error bars indicate the ± SE. All experiments were repeated at least three times, with similar results
425_2017_2770_MOESM4_ESM.pptx
Supplementary material 4 (PPTX 16069 kb) Fig. S4 Subcellular localization and Transcriptional activation analysis of SiNAC2-SiNAC5. a Subcellular localization of SiNAC2-SiNAC5. Column 1 is the signal of GFP; column 2 is bright field of a same cell; columns 3 are overlaps in the same cell. Scale bars=20 μm. b Transcriptional activation analysis of SiNAC2-SiNAC5.All experiments were repeated at least three times, with similar results
425_2017_2770_MOESM5_ESM.pptx
Supplementary material 5 (PPTX 24760 kb) Fig. S5 Altered sensitivity of SiNAC1-OE plants in response to ABA at the germination and cotyledon greening stages compared with Col-0. Seeds of Col and SiNAC1 overexpression plants were sown on petri dishes with or without ABA. The concentration of ABA were 0.25 μM, 0.5 μM and 0.75 μM, respectively. a The different seed germination and cotyledon greening phenotype of Col and transgenic seedlings when incubation in growth chamber after 5 days. b Seed germination rate under different ABA concentration was calculated after vernalization. c Cotyledon greening rate was calculated from the 2nd day to the 7th day after vernalization. Error bars indicate the ± SE. All experiments were repeated at least three times, with similar results
425_2017_2770_MOESM6_ESM.pptx
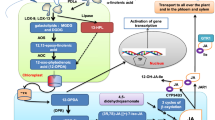
Supplementary material 6 (PPTX 274 kb) Fig. S6 A model for the role of SiNAC1 in participating a positive feedback loop during leaf senescence. Under natural and dark condition, the onset of leaf senescence was induced by various hormones including ABA. When leaf senescence initiation, ABA biosynthesis and ABA signaling pathway were also activated, which further magnified the senescence signals. During this process, SiNAC1 was a senescence induced transcription factor which could also up-regulated by ABA. Moreover, SiNAC1 promoted expression of two key enzymes in ABA biosynthesis NCED2 and NCED3, which subsequently lead ABA accumulation. Therefore, SiNAC1 involved in a ternary positive feedback module and was response for mediating the leaf senescence signaling to ABA biosynthesis
Rights and permissions
About this article
Cite this article
Ren, T., Wang, J., Zhao, M. et al. Involvement of NAC transcription factor SiNAC1 in a positive feedback loop via ABA biosynthesis and leaf senescence in foxtail millet. Planta 247, 53–68 (2018). https://doi.org/10.1007/s00425-017-2770-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-017-2770-0