Abstract
Cellular RNAs in all three kingdoms of life are modified with diverse chemical modifications. These chemical modifications expand the topological repertoire of RNAs, and fine-tune their functions. Ribosomal RNA in yeast contains more than 100 chemically modified residues in the functionally crucial and evolutionary conserved regions. The chemical modifications in the rRNA are of three types—methylation of the ribose sugars at the C2-positionAbstract (Nm), isomerization of uridines to pseudouridines (Ψ), and base modifications such as (methylation (mN), acetylation (acN), and aminocarboxypropylation (acpN)). The modifications profile of the yeast rRNA has been recently completed, providing an excellent platform to analyze the function of these modifications in RNA metabolism and in cellular physiology. Remarkably, majority of the rRNA modifications and the enzymatic machineries discovered in yeast are highly conserved in eukaryotes including humans. Mutations in factors involved in rRNA modification are linked to several rare severe human diseases (e.g., X-linked Dyskeratosis congenita, the Bowen–Conradi syndrome and the William–Beuren disease). In this chapter, we summarize all rRNA modifications and the corresponding enzymatic machineries of the budding yeast.
You have full access to this open access chapter, Download protocol PDF
Similar content being viewed by others
Key words
- rRNA modification
- Ribose methylation
- Pseudouridylation
- Base methylation
- Aminocarboxypropylation
- Acetylation of cytidines
- Methyltransferase
1 Introduction
RNA modifications are present in all three kingdoms of life and detected in all classes of cellular RNA s. RNA modifications are diverse, with more than 100 types of chemical modifications identified to date [1]. Ribosomes are molecular assemblies of RNA and proteins and are responsible for the synthesis of all proteins in the cells [2]. Structural and functional analyses of ribosomes have revealed that it is the ribosomal RNA (rRNA) that makes the structural framework of ribosomes and catalyzes the joining of amino acids (peptidyl transfer) during translation , hence making the ribosome a ribozyme [3]. Though the chemical composition of RNA seems to be rather insufficient to provide the structural complexity to RNA , the composition analysis of rRNA has shown that rRNA contains different chemical modifications that are added both co- and posttranscriptionally [4, 5]. Ribosomal RNA (rRNA) contains three types of chemical modifications , methylation of the ribose sugars at the C2-position (Nm), isomerization of uridines to pseudouridines (Ψ), and base modifications (methylation (mN) such as acetylation (acN) and aminocarboxypropylation (acpN) ) [6]. Mutations in factors involved in rRNA modifications are associated with several rare severe human diseases (e.g., X-linked dyskeratosis congenita, the Bowen–Conradi syndrome, Hutchison Gilford syndrome , and the William–Beuren disease) [7,8,9,10,11,12,13]. Emerging evidences indicate that some bases are not always completely modified providing heterogeneity with respect to RNA modification [14,15,16]. Heterogeneity in rRNA modification has been correlated with disease etiology (cancer) and shown to play a role in cell differentiation (hematopoiesis) [17]. Remarkably, alterations in rRNA modification patterns profoundly affect the preference of ribosomes for cap- versus IRES-dependent translation having major consequences on cell physiology [18, 19]. Here, we summarize all known rRNA modifications of the budding yeast with an emphasis on base modifications (see Tables 1, 2, 3 and 4).
In Saccharomyces cerevisiae , 18S rRNA of the small subunit contains 14 Ψs, 18 Nms (2′O-methylated Ns), 4 mNs (methylated Ns), and 2 acNs (acylated Ns) (Tables 1 and 2), whereas 25S rRNA of the large subunit contains 30 Ψs, 35 Nms, and 6 base methylations (Tables 3 and 4). Mapping of these modifications has revealed that these modifications cluster in the functionally conserved regions of the ribosomes like the intersubunit and the peptidyl transferase center [6, 20, 21]. Due to technical limitations, the chemical modification profile of rRNA , especially for the base modifications, remained poorly characterized for a long time. Using state-of-the-art RP-HPLC and mass spectrometry together with “reverse genetics,” we and others have recently completed the characterization and mapping of the complete set of yeast rRNA modifications , and have identified the corresponding enzymatic machinery involved in adding these modifications to the RNA [16, 22, 23].
2 Ribose Methylation
Methylation of 2′-OH of ribose sugar to a 2′-O-methyl-ribose is a characteristic modification in mRNA and many noncoding RNA (ncRNA) including tRNA, rRNA and siRNAs (Fig. 1). Ribose methylation favors a 3′-endo conformation of the ribose and since 3′-endo conformations are known to stabilize A-form helices, methylation of ribose increases the rigidity of the RNA by promoting base stacking. Furthermore, ribose methylation provides RNA stability against base and nuclease hydrolysis by insulating the otherwise active 2′-OH group. Our recent analyses together with other groups have identified several partial rRNA modifications in eukaryotes including humans. These observations suggest the existence of a heterogeneous population of ribosomes supporting the “specialized” translation model, which could possibly play an important role during embryonic development [14,15,16, 22, 24, 25].
2.1 C/D Box snoRNPs
All 2′-OH ribose methylation occur at distinct positions in eukaryotic rRNAs (Tables 1 and 3). The respective positions are targeted via specific C/D box snoRNAs having complementary guide sequences to the respective rRNAs together with distinguishing sequence elements called boxes C and D.
The methylation guide sequence is located upstream of the box D/D′ element and consists of 10–21 nucleotides. The guide sequences direct ribose methylation to the nucleotide base paired to the fifth nucleotide (nt) upstream of the box D or D′ sequence (box D + 5 rule) [26]. The C/D box snoRNPs consist of four common core proteins: Fibrillarin (human)/Nop1 (yeast ), Nop58 , Nop56 , and Snu13 [27]. Interestingly, C/D box snoRNPs involved in functions apart from catalysis like U3, U14 , U8, U22 , snR4 , and snR45 also contain additional proteins [27, 28]. Nop1 is a S-adenosyl methionine (SAM) dependent methyltransferase and catalyzes the 2′-O methylation reaction. Snu13 binds to the kink-turn (loop-stem structure that includes the canonical C/D elements in the loop portion) in the C/D box of snoRNAs . Nop56 and Nop58 are characterized by extensive coiled-coil domains, likely responsible for heterodimerization and providing stability to the snoRNA .
3 Pseudouridylation
Pseudouridylation (ψ) is a C-glycoside rotation isomer of uridine (Fig. 1). Due to rotation, the nitrogen atom at position 1 (N1) forms no longer a glycosidic bond to the ribose and is protonated at physiological pH. Compared to uracil, in pseudouracil both N1 and N3 participate in hydrogen bonding. The N1 proton makes hydrogen bonding with a phosphate group from the same or neighboring nucleotide and provides stability of the structure.
Furthermore, like ribose methylation , pseudouridine also favors a C3-endo sugar pucker (the conformation preferred by an A-form RNA helix) and has been shown to increase the thermal stability of RNA by up to 2 °C [29]. Therefore, the presence of Ψ also plays an important role in RNA stability that may not be essential, but seemingly provides a significant advantage [5].
3.1 H/ACA snoRNPs
The majority of pseudoridines in rRNAs are added via specific H/ACA box snoRNPs having complementary guide sequences to the respective rRNAs and contain small sequence elements referred to as boxes H and ACA [30]. Similar to C/D box snoRNAs , substrate targeting involves base pairing through two short guide sequences in a loop portion of the duplex structures (Tables 1 and 2). The guide sequences are 14–15 nucleotides from the H or ACA box. The binding of the substrate places the target uridine in a pseudouridylation pocket between the flanking paired regions [30].
H/ACA snoRNPs contain four core proteins in yeast : Cbf5p (Dyskerin in humans), Gar1 , Nhp2 , and Nop10 . Cbf5 is the catalytic pseudouridine synthase. Although the crystal structure from the eukaryotic H/ACA snoRNP is still missing, the archaeal H/ACA structure has provided noteworthy details about the organization of various core proteins [31]. Cbf5 has been shown to contact the ACA motif and both P1 and P2 stem. Nhp2 , the yeast homolog of archaeal L7Ae binds to the K loop structure in the upper half of the RNA . Gar1 does not bind to the RNA directly but rather joins the complex through its interaction with Cbf5. Further structural analyses of Cbf5 have revealed that its interaction with Gar1 is essential for substrate binding and release [27].
4 Base Modifications
The rRNAs contain three different types of base modifications —methylation (m), acetylation (ac), and aminocarboxypropylation (acp). Methylation is the most common base modification in rRNA . All four nitrogenous bases of RNA undergo methylation either at nitrogen (N) or carbon (C) atoms. On the other hand, only cytosine bases are acetylated, and aminocarboxypropyl is added either to a uridine or pseudouridine residues in yeast . The base modifications of RNA are primarily catalyzed by snoRNA-independent enzymes with only exception being 18S rRNA acetylation that requires particular C/D box snoRNAs (Tables 2 and 4).
4.1 Base Methylation
Methylation of nitrogenous base strongly affect their physical and chemical properties. Methylation promotes base stacking by increasing the hydrophobicity and the polarizability. Furthermore, methylation also influences the structure by increasing steric hindrance, blocking canonical (Watson–Crick) hydrogen bonding and fostering noncanonical Hoogsteen base paring . This presumably helps ncRNA like rRNA to attain and maintain specific conformations, essential for their corresponding function—both with respect to their structure and their enzymatic activity.
4.2 Methyl transferases
Methyltransferases are the enzymes that catalyze specific transfer of methyl group form a methyl donor to various substrates. S-adenosyl-methionine (SAM or AdoMet) is the most common methyl donor by virtue of the presence of a charged methylsulfonium center [32]. Methyltransferases that utilize the methyl group of SAM for the methylation reaction are called SAM dependent methyltransferases. Based on their structural analysis, all currently known SAM dependent methyltransferases have been divided into five distinct classes (Class I to Class V) [33].
Class I methyltransferases are characterized by a Rossmann-fold like domain and methylate a wide variety of substrates (DNA, RNA and proteins along with other small molecules). Rossmann folds are nucleotide (especially NAD(P)) binding domains that also contain an alternating α/β strands topology in which two Rossmann fold domains are linked into 6 parallel β stands sandwiched by a pair of α-helices [32]. The Rossmann-like fold comprises of alternating β-stranded and α-helical regions, with all strands forming a central relatively planar β-sheet, and helices stuffing two layers, one on each side of the plane.
Class II methyltransferases are characterized by a methionine synthase activation domain. The Met synthase activation contains an unusual fold with long, central, antiparallel β-sheet flanked by groups of helices at either end, which makes it structurally distinct from Class I methyltransferases [32].
Class III methyltransferases act on ring carbons of the large, planar precorrin substrates during cobalamin biosynthesis (e.g., CbiF) [32].
Class IV methyltransferases belong to the SPOUT family and methylate either RNA or proteins [32, 34]. These enzymes contain a six- stranded parallel β sheet flanked by seven α-helices. Interestingly the first three strands of these methyltransferases form half of a Rossmann fold . The active site of SPOUT methyltransferases is located near the subunit interface of a homodimer.
Class V methyltransferases contain typical SET domains, discovered originally as conserved domain shared by chromatin remodeling proteins Su (var) 3–9, E (Z) (short for Enhancer of Zeste) and Trithorax [32]. Most of the currently known SET methyltransferases methylate lysine residues of various nuclear proteins involved in chromatin remodeling and transcriptional regulation. The SAM binding site and the catalytic center of SET domains contains all-β (eight curved β strands forming three small sheets)and knot-like structures like Class IV methyltransferases but based on a different topology [32].
4.3 N4- Acetylation of Cytidine (ac4C)
Acetylation of cytidine residues is a highly conserved base modification present in 18S rRNA , as well as leucine and serine tRNAs of yeast [35, 36]. Molecular dynamics simulation and in vitro studies using the noninitiator methionine-accepting tRNA of E. coli have reported that acetylation of cytidine residues stabilize the C3′-endo puckering conformation of ribose, and stabilizes the G–C base pairing in the RNA , which has been highlighted as an important function of this modification in counteracting mistakes that may occur during translation due to misreading of the isoleucine AUA codon by tRNAMet [37,38,39]. Interestingly, in the 18S rRNA both ac4C residues [ac4C 1280 (helix 34) and ac4C1773 (helix 45)] are also involved in C-G base pairings that are fundamental for ribosome functionality [6, 40]. The disruption of these base pairing was found to be lethal for yeast cells (unpublished data).
Structural and functional analysis of bacterial RNA acetyltransferase TmcA has revealed that RNA acetyltransferases utilize acetyl-CoA as an acetyl group donor, which is transferred in an ATP-dependent manner to the cytosine residues [41]. RNA acetyltransferases contain an N- terminal RNA helicase domain similar to that of DEAD-box RNA helicases paired with a C- terminal Gcn5-related N-acetyltransferase (GNAT) fold. TmcA is a stand-alone enzyme and does not necessitate auxiliary factors for substrate specificity, which is likely due to similar substrates that it acetylates [42].
In contrast to TmcA, the yeast and higher eukaryotes TmcA homologs Kre33 /NAT10 utilize protein factors Tan1 and THUMPD1, respectively, for tRNA acetylation and snoRNAs snR4 and snR45 for 18S rRNA acetylation [28, 43, 44].
4.4 3-Amino carboxy propylation acp
The addition of 3-aminocarboxypropyl (acp) to the RNA is another highly conserved modification in eukaryotic cellular RNAs derived from S-adenosyl-L-methionine (SAM ) [45]. This modification is mostly added to the uridine and pseudouridine residues and impacts the ability of these bases to establish hydrogen bonding with otherwise complementary adenosine residue [46, 47].
RNA aminocarboxypropyl transferase is a novel class of SAM dependent acp transferase with a significant homology to the SPOUT-class RNA-methyltransferases [46, 47]. Structure analysis of its archaeal Tsr3 homologs has revealed that these enzymes contains the same fold as in SPOUT- class-RNA methyltransferase [34], but due to a discrete SAM binding arrangement acp transferases transfer the acp instead of the methyl group of SAM to its substrate [47].
5 Base Modifications of 18S rRNA
The 18S rRNA of ribosome small subunit (SSU) contains four base methylation : one m1acp3Ψ1191, two m6 2A1781, m6 2A1782, and a single m7G1575, and two base acetylation —ac4C 1280 and 1773 (Fig. 1 and Table 3).
5.1 Dim1 Catalyzes m6 2A1781 and m6 2A1782 dimethylation
The first base methylation analyzed in rRNAs of S. cerevisiae were the m6 2A1781 and m6 2A1782 dimethylations [48]. Both dimethylations of adenine residues at carbon atom 6 (C-6) in the 3′of SSU rRNA are highly conserved and are catalyzed by universally conserved orthologous methyltransferases. In S. cerevisiae , Dim1 catalyzes the methylation of both adenine residues [48, 49]. Dim1 is an essential protein and plays a key role in 40S biogenesis. Interestingly, both these dimethylations of 18S are performed in the cytoplasm and it has been shown that 40S subunits lacking dimethylated 18S rRNA are not competent for translation in vitro [48, 49].
Nevertheless, further analysis of this methylation and its corresponding enzymes revealed that the methylation deficient mutant of dim1 is viable albeit influences the 35S rRNA processing, suggesting that methylations are dispensable for ribosome function in vivo [48, 49].
5.2 Bud23 Catalyzes m7G1575 Methylation
Bud23 catalyzes the methylation of nitrogen atom 7 in the guanosine residue at position 1575 (m7G1575) of 18S rRNA [50]. Residue G1575 is highly conserved and interacts with the anticodon loop of the P-site tRNA [40]. Bud23 is a eukaryote specific enzyme and no orthologs have been detected in prokaryotes. It is a nucleolar protein and has been shown to be critical for the efficient export of the small subunit and its cytoplasmic maturation [50]. Intriguingly, as observed for Dim1 , the m7G1575 methylation also turned out to be not the essential function of Bud23 as the loss of methylation does not exhibit any 40S processing and export defects. Conspicuously, Bud23 needs the assistance of methyltransferase adaptor protein, Trm112 for the methylation of G1575 [50,51,52].
5.3 Nep1 (Emg1 ) and Tsr3 Catalyze Ψ1191 Modification (m1 acp3Ψ1191)
The most interesting modification in the rRNA result in 1-methyl-3-(3-amino-3-carboxypropyl )-pseudouridine (m1acp3Ψ1191) [4]. S. cerevisiae 18S rRNA contains a single m1acp3Ψ hypermodification at residue 1191, located in the decoding center of the 40S subunit, proximal to the P site-tRNA [40]. The m1acp3Ψ1191 modification comprises of three reactions: (1) pseudouridylation , (2) methylation , and finally (3) the addition of 3-amino-3-carboxypropyl [8, 47, 53]. During maturation of 18S rRNA , these modifications are added sequentially. First, snR35 snoRNP catalyzes the pseudouridylation of U1191, thus unlinking the N1 of the U1191 from the glycosidic bond with the ribose sugar [30]. Next, Nep1 (Emg1 ) methylates this N1 of Ψ1191. Nep1 , also described as Emg1 is a highly conserved and an essential pseudouridine-N1-specific methyltransferase [8] with an extended SPOUT-class methyltransferase fold and a pre-organized SAM-binding site [9, 54, 55]. Remarkably, a point mutation in the human Nep1 (Emg1) protein (D86G) has been shown to be responsible for Bowen–Conradi syndrome (BSC mutation) [7]. As already observed in case of Bud23 and Dim1 , the methylation appeared not to be the essential function of the Nep1 . Instead, the BSC-mutated yeast Nep1 protein showed enhanced dimerization propensity and increased affinity for its RNA-target in vitro. Furthermore, the BCS mutation prevented nucleolar accumulation of Nep1, which could be the reason for reduced growth in yeast and the Bowen-Conradi syndrome [8]. Furthermore, loss of methylation leads to defects in 40S subunits and affects antibiotic sensitivity [8, 56, 57]. Genetic analysis of Nep1 has provided understanding of its possible essential function [56, 58]. Overexpression of Rps19 protein has been demonstrated to suppress the deletion of otherwise essential Nep1 [58]. This has led to the hypothesis that perhaps the essential function of Nep1 is to assist the incorporation of Rps19 protein during 40S biogenesis. Nevertheless, this model awaits the biochemical experiments in its support [59].
Whereas Ψ (snR35) and N1-CH3 (Nep1 ) are introduced in the nucleus, it was shown that the addition of 3-amino-3-carboxypropyl to the Ψ1191 takes place in the cytoplasm [53]. The enzyme responsible for the transfer of 3-amino-3-carboxypropyl (acp) to the N3 atom of Ψ1191 was recently identified as Tsr3 [47]. A homologous enzyme was also identified in E. coli that catalyzes acp3U modification at position 47 in the variable loop of eight E. coli tRNAs [46].
5.4 Kre33/NAT10 Catalyzes ac4C1280 and ac4C1773 Acetylation of the 18S rRNA in a snoRNA Dependent Manner
Eukaryotes contain only one cytidine acetyltransferase Kre33 (yeast ) and NAT10 (human) that catalyzes both rRNA and tRNA cytidine acetylation [28, 43, 44]. Kre33 contains a N terminal helicase domain fused to the C terminal acetyltransferase domain related to GCN5 [42, 44]. In S. cerevisiae Tan1 was initially identified as a protein that is required for acetylation of ac4C formation at position 12 of tRNA-Ser (CGA) [36]. Nevertheless, Tan1 does not contain any catalytic (acetyltransferase) domain for ac4C formation but rather contains the THUMP domain, responsible for tRNA binding.
Interestingly, the acetylation of two cytosine residues in 18S rRNA catalyzed by Kre33 are guided by the two box C/D snoRNAs snR4 and snR45 which are not known to be involved in methylation [28]. This is in contrast to tRNA acetylation where protein Tan1 likely guides Kre33 to the acetylated cytidine [28, 43, 44]. These results highlighted yet another example of an incredible cellular modularity, where a single enzyme is targeted to diverse substrates by means of either protein or RNA adaptors. Both snR4 and snR45 establish extended bipartite complementarity around the cytosines targeted for acetylation [28].
The viability of catalytically deficient Kre33 mutants demonstrated that acetylation of 18S rRNA is dispensable for cell viability. Although both catalytic deficient mutants, kre33-H545A and kre33-R637A exhibited severe growth defects at 37 °C, similar to what was observed previously for the Tan1 deletion mutant [36, 44]. Since snoRNA deletions did not exhibit this phenotype at 37 °C, we concluded that this is probably due to the loss of tRNA acetylation .
6 Base Modifications of 25S rRNA
The 25S rRNA of the ribosome large subunit (LSU) in yeast contains 6 base methylation : two m1A (1-methyl adenosine), two m5C (5-methyl cytosine ), and two m3U (3-methyl uridine ) (Fig. 1 and Table 4). Two m5U (5-methyl uridine) methylations were also reported previously but the presence of these in the 25S rRNA have been disproven.
6.1 Rrp8 (Bmt1) Catalyzes m1 A 645 Methylation
Rrp8 , a protein previously shown to be involved in the processing of 35S rRNA at site A2 [60]. Rrp8 catalyzes m1A645 base methylation of 25S rRNA and is a Class I SAM dependent rRNA methyltransferase [61]. The mapping of m1A645 on the ribosomal RNA revealed that this residue is present in helix 25.1 of domain II of the 25S rRNA . Helix 25.1 is highly conserved in eukaryotes including humans [62]. The in vivo structural probing of 25S rRNA , using both DMS and SHAPE, revealed that the loss of the Rrp8-catalyzed m1A modification alters the conformation of domain I of yeast 25S rRNA causing translation initiation defects detectable as halfmer formation, likely because of incompetent loading of 60S on the 43S-preinitiation complex [62]. Surprisingly, quantitative proteomic analysis of the yeast Δrrp8 mutant strain using 2D-DIGE have exhibited that loss of m1A645 impacts production of a specific set of proteins involved in carbohydrate metabolism, translation , and ribosome synthesis [62]. These findings in yeast point to a role of Rrp8 in primary metabolism. In conclusion, the m1A modification is crucial for maintaining an optimal 60S conformation, which in turn is important for regulating the synthesis of key metabolic enzymes.
6.2 Bmt2 Catalyzes A2142 Methylation (m1A2142)
Bmt2 catalyzes the m1A2142 methylation located in helix 65 (H65) of the 25S rRNA [63]. H65 belongs to domain IV of 25S rRNA , which makes most of the intersubunit surface of the large subunit . Interestingly, L2, a highly conserved protein makes physical contact with H65, especially with its SH3-β barrel globular domain. Like Rrp8 , Bmt2 also belongs to Class I Rossmann-like fold methyltransferases family [63].
A deletion mutant of BMT2 was previously identified to extend hibernating life span and to exhibit peroxide sensitivity [64]. A methyltransferase dead mutant (bmt2-G180R) and rDNA point mutants (A2142G, A2142C, and A2142T) also exhibited hydrogen peroxide sensitivity. Taken together this indicates that the biosynthetic pathways of this modifications interact with the cellular response toward oxidative stress stimulated by hydrogen peroxide.
6.3 Rcm1 (Bmt3 ) and Nop2 (Bmt4 ) Catalyze m5C2278 and m5C2870 Methylation
Yeast 25S rRNA contains two m5C residues at positions 2278 and 2870 [65]. Rcm1 and Nop2 were shown to be responsible for two distinct m5C methylations of 25S rRNA at position C2278 and C2870, respectively [66]. Both Rcm1 and Nop2 belong to Class I methyltransferases characterized by Rossmann-like folds. Interestingly, biochemical and structural analyses of other RNA cytosine methyltransferases have provided an important insight into their catalytic mechanism [67]. All known RNA cytosine methyltransferases utilize two highly conserved cysteine residues in the motif IV and motif VI for the addition of methyl group at C5 of cytosine [65, 67]. The cysteine in motif VI makes a nucleophilic attack on to the C-6 of cytosine and form a covalent adduct with the RNA , whereas the cysteine of motif IV is important for the release of the substrate [67]. For both Rcm1 and Nop2 exchange of cysteine in motif VI with the alanine led to methyltransferase-dead mutants, whereas replacement of cysteine in motif IV with alanine turned out to be lethal [66].
The lethality of both rcm1 and nop2 motif IV cysteine mutant proteins suggested that due to substitution of cysteines in motif IV, the mutant proteins fail to separate themselves from their corresponding targets and forms a stable protein-rRNA complex. This fixing of the mutant protein then blocks the 25S rRNA processing and probably causes cell death. This model was further supported by suppression of this lethality upon exchange of both motif IV and motif VI cysteine residues together in case of Rcm1 and also Nop2 , as this precluded the formation of any covalent complex [66].
Nop2 is an essential protein in S. cerevisiae [68]. Previous biochemical analyses using the hypomorphic expression system have shown that Nop2 depletion causes severe defects in the rRNA processing and 60S biogenesis. The viability of the nop2-C478A methyltransferase-deadmutant demonstrated, that methylation at C2870 is not the essential function of Nop2 . In other words m5C2870 is not essential for viability. Nevertheless, the polysome profiles and Northern-blotting analysis of the methyltransferase-dead mutant of Nop2 demonstrated that loss of m5C2870 strongly impairs the 35S rRNA processing and 60S biogenesis; the phenotypes observed previously upon depletion of Nop2 [66]. Therefore, this suggested that the Nop2 depletion phenotypes observed previously are apparently the result of reduced m5C modification.
Absence of m5C2278 causes anisomycin hypersensitivity [66]. In contrast to m5C2870, loss of m5C 2278 did not exhibit any defects in 60S biogenesis and 35S rRNA processing. As far as the biological role of m5C2278 is concerned, loss of m5C2278 in helix 70 (H70) causes anisomycin hypersensitivity , indicating that the loss of this methylation disrupts its optimal conformation. Together with the data from the m1A2142 base modification in helix 65, the anisomycin hypersensitivity after the loss of m5C 2278 also suggests that the conformation of domain IV of 25S rRNA is crucial for anisomycin sensitivity as both m1A2142 and m5C 2278 reside in the domain IV of the 25S rRNA [66].
To understand how 25S/28S rRNA cytosine methylation (m5C) and its corresponding enzyme (Rcm1 /NSUN5 ) impact biological functions, in a collaboration with Johannes Grillari’s lab, we found, that reduced levels of the enzyme increase the life span and stress resistance in yeast , worms, and flies [69]. Loss of m5C alters the structural conformation of the ribosome and promotes translational reprogramming by favoring translation of a distinct subset of oxidative stress-responsive mRNAs [69]. Thus, rather than merely being a static molecular machine executing translation , the ribosome exhibits functional diversity by modification of just a single rRNA nucleotide, resulting in an alteration of organismal physiological behavior, and linking rRNA-mediated translational regulation to the modulation of life span, and differential stress response.
6.4 Bmt5 and Bmt6 Catalyze m3U2634 and m3U2843 Methylation
Bmt5 and Bmt6 catalyze the m3U methylations at positions 2634 and 2843 of the 25S rRNA [70]. Both enzymes belong to the Rossmann-like fold protein family. The substitution of highly conserved glycine residues in the SAM binding motif of both Bmt5 and Bmt6 with arginine abolished their catalytic activity. Surprisingly, m3U methylation in the 16S rRNA of E. coli is catalyzed by proteins of the SPOUT methyltransferase family which had led to the assumption that m3U methylation could only be performed by SPOUT methyltransferases [71]. With the identification of Bmt5 and Bmt6 this convention is no more valid.
Both Bmt5 and Bmt6 are highly conserved in lower eukaryotes including yeasts like Schizosaccharomyces pombe , Neurospora crassa, and Kluyveromyces lactis. The Bmt5 and Bmt6 homologs are likely performing the same function in these organisms.
6.5 25S rRNA of Yeast Does Not Contain any m5U Residues
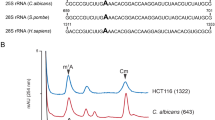
Yeast 25S rRNA was predicted to contain two m5U residues at position 956 and 2924 [20, 72]. Using both RP-HPLC and mass spectrometry this was disproved [70]. Similarly, the 25S rRNA of C. albicans and S. pombe also did not contain any m5U residues [73]. This has been also confirmed for higher eukaryotes , whereas m5U residues have been only observed in tRNAs , which is in consent with the presence of only one m5U methyltransferase, Trm2 (in yeast and its homologs), responsible for m5U modification of tRNAs at position 54. Up till now, Trm2 has not been observed to display any interaction with the rRNA in yeast . Taken together, this would apparently suggest that during the course of evolution m5U methylation have disappeared in rRNA of eukaryotes .
References
Boccaletto P, Machnicka MA, Purta E, Piatkowski P, Baginski B, Wirecki TK, de Crecy-Lagard V, Ross R, Limbach PA, Kotter A et al (2018) MODOMICS: a database of RNA modification pathways. 2017 update. Nucleic Acids Res 46:D303–D307
Woolford JL Jr, Baserga SJ (2013) Ribosome biogenesis in the yeast Saccharomyces cerevisiae. Genetics 195:643–681
Cech TR (2000) Structural biology. The ribosome is a ribozyme. Science 289:878–879
Klootwijk J, Planta RJ (1973) Analysis of the methylation sites in yeast ribosomal RNA. Eur J Biochem 39:325–333
Motorin Y, Helm M (2011) RNA nucleotide methylation. Wiley Interdiscip Rev RNA 2:611–631
Sharma S, Lafontaine DLJ (2015) ‘View from a Bridge’: a new perspective on eukaryotic rRNA base modification. Trends Biochem Sci 40:560–575
Armistead J, Khatkar S, Meyer B, Mark BL, Patel N, Coghlan G, Lamont RE, Liu S, Wiechert J, Cattini PA et al (2009) Mutation of a gene essential for ribosome biogenesis, EMG1, causes Bowen-Conradi syndrome. Am J Hum Genet 84:728–739
Meyer B, Wurm JP, Kotter P, Leisegang MS, Schilling V, Buchhaupt M, Held M, Bahr U, Karas M, Heckel A et al (2011) The Bowen-Conradi syndrome protein Nep1 (Emg1) has a dual role in eukaryotic ribosome biogenesis, as an essential assembly factor and in the methylation of Psi1191 in yeast 18S rRNA. Nucleic Acids Res 39:1526–1537
Wurm JP, Meyer B, Bahr U, Held M, Frolow O, Kotter P, Engels JW, Heckel A, Karas M, Entian KD et al (2010) The ribosome assembly factor Nep1 responsible for Bowen-Conradi syndrome is a pseudouridine-N1-specific methyltransferase. Nucleic Acids Res 38:2387–2398
Doll A, Grzeschik KH (2001) Characterization of two novel genes, WBSCR20 and WBSCR22, deleted in Williams-Beuren syndrome. Cytogenet Cell Genet 95:20–27
Knight SW, Heiss NS, Vulliamy TJ, Greschner S, Stavrides G, Pai GS, Lestringant G, Varma N, Mason PJ, Dokal I et al (1999) X-linked dyskeratosis congenita is predominantly caused by missense mutations in the DKC1 gene. Am J Hum Genet 65:50–58
Nachmani D, Bothmer AH, Grisendi S, Mele A, Bothmer D, Lee JD, Monteleone E, Cheng K, Zhang Y, Bester AC et al (2019) Germline NPM1 mutations lead to altered rRNA 2′-O-methylation and cause dyskeratosis congenita. Nat Genet 51:1518–1529
Venturi G, Montanaro L (2020) How altered ribosome production can cause or contribute to human disease: the spectrum of ribosomopathies. Cell 9:2300
Buchhaupt M, Sharma S, Kellner S, Oswald S, Paetzold M, Peifer C, Watzinger P, Schrader J, Helm M, Entian KD (2014) Partial methylation at Am100 in 18S rRNA of baker's yeast reveals ribosome heterogeneity on the level of eukaryotic rRNA modification. PLoS One 9:e89640
Sloan KE, Warda AS, Sharma S, Entian KD, Lafontaine DLJ, Bohnsack MT (2017) Tuning the ribosome: the influence of rRNA modification on eukaryotic ribosome biogenesis and function. RNA Biol 14:1138–1152
Taoka M, Nobe Y, Yamaki Y, Yamauchi Y, Ishikawa H, Takahashi N, Nakayama H, Isobe T (2016) The complete chemical structure of Saccharomyces cerevisiae rRNA: partial pseudouridylation of U2345 in 25S rRNA by snoRNA snR9. Nucleic Acids Res 44:8951–8961
Vlachos A (2017) Acquired ribosomopathies in leukemia and solid tumors. Hematology Am Soc Hematol Educ Program 2017:716–719
Aspesi A, Ellis SR (2019) Rare ribosomopathies: insights into mechanisms of cancer. Nat Rev Cancer 19:228–238
Barbieri I, Kouzarides T (2020) Role of RNA modifications in cancer. Nat Rev Cancer 20:303–322
Bakin A, Lane BG, Ofengand J (1994) Clustering of pseudouridine residues around the peptidyltransferase center of yeast cytoplasmic and mitochondrial ribosomes. Biochemistry 33:13475–13483
Maden BE (1988) Locations of methyl groups in 28 S rRNA of Xenopus laevis and man. Clustering in the conserved core of molecule. J Mol Biol 201:289–314
Yang J, Sharma S, Watzinger P, Hartmann JD, Kotter P, Entian KD (2016) Mapping of complete set of ribose and base modifications of yeast rRNA by RP-HPLC and mung bean nuclease assay. PLoS One 11:e0168873
Yang J, Sharma S, Kotter P, Entian KD (2015) Identification of a new ribose methylation in the 18S rRNA of S. cerevisiae. Nucleic Acids Res 43:2342–2352
Sharma S, Marchand V, Motorin Y, Lafontaine DLJ (2017) Identification of sites of 2′-O-methylation vulnerability in human ribosomal RNAs by systematic mapping. Sci Rep 7:11490
Shi Z, Barna M (2015) Translating the genome in time and space: specialized ribosomes, RNA regulons, and RNA-binding proteins. Annu Rev Cell Dev Biol 31:31–54
Kiss L (1996) Site-specific ribose methylation of preribosomal RNA: a novel function for small nucleolar RNAs. Cell 85:1077–1088
Watkins NJ, Bohnsack MT (2012) The box C/D and H/ACA snoRNPs: key players in the modification, processing and the dynamic folding of ribosomal RNA. Wiley Interdiscip Rev RNA 3:397–414
Sharma S, Yang J, van Nues R, Watzinger P, Kotter P, Lafontaine DLJ, Granneman S, Entian KD (2017) Specialized box C/D snoRNPs act as antisense guides to target RNA base acetylation. PLoS Genet 13:e1006804
Noon KR, Guymon R, Crain PF, McCloskey JA, Thomm M, Lim J, Cavicchioli R (2003) Influence of temperature on tRNA modification in archaea: Methanococcoides burtonii (optimum growth temperature [Topt], 23 degrees C) and Stetteria hydrogenophila (Topt, 95 degrees C). J Bacteriol 185:5483–5490
Ganot P, Bortolin ML, Kiss T (1997) Site-specific pseudouridine formation in preribosomal RNA is guided by small nucleolar RNAs. Cell 89:799–809
Duan J, Li L, Lu J, Wang W, Ye K (2009) Structural mechanism of substrate RNA recruitment in H/ACA RNA-guided pseudouridine synthase. Mol Cell 34:427–439
Kozbial PZ, Mushegian AR (2005) Natural history of S-adenosylmethionine-binding proteins. BMC Struct Biol 5:19
Schubert HL, Blumenthal RM, Cheng X (2003) Many paths to methyltransfer: a chronicle of convergence. Trends Biochem Sci 28:329–335
Anantharaman V, Koonin EV, Aravind L (2002) SPOUT: a class of methyltransferases that includes spoU and trmD RNA methylase superfamilies, and novel superfamilies of predicted prokaryotic RNA methylases. J Mol Microbiol Biotechnol 4:71–75
Thomas G, Gordon J, Rogg H (1978) N4-Acetylcytidine. A previously unidentified labile component of the small subunit of eukaryotic ribosomes. J Biol Chem 253:1101–1105
Johansson MJ, Bystrom AS (2004) The Saccharomyces cerevisiae TAN1 gene is required for N4-acetylcytidine formation in tRNA. RNA 10:712–719
Kawai G, Hashizume T, Miyazawa T, McCloskey JA, Yokoyama S (1989) Conformational characteristics of 4-acetylcytidine found in tRNA. Nucleic Acids Symp Ser (21):61–62
Kumbhar BV, Kamble AD, Sonawane KD (2013) Conformational preferences of modified nucleoside N(4)-acetylcytidine, ac4C occur at “wobble” 34th position in the anticodon loop of tRNA. Cell Biochem Biophys 66:797–816
Stern L, Schulman LH (1978) The role of the minor base N4-acetylcytidine in the function of the Escherichia coli noninitiator methionine transfer RNA. J Biol Chem 253:6132–6139
Ben-Shem A, Garreau de Loubresse N, Melnikov S, Jenner L, Yusupova G, Yusupov M (2011) The structure of the eukaryotic ribosome at 3.0 A resolution. Science 334:1524–1529
Ikeuchi Y, Kitahara K, Suzuki T (2008) The RNA acetyltransferase driven by ATP hydrolysis synthesizes N4-acetylcytidine of tRNA anticodon. EMBO J 27:2194–2203
Chimnaronk S, Suzuki T, Manita T, Ikeuchi Y, Yao M, Suzuki T, Tanaka I (2009) RNA helicase module in an acetyltransferase that modifies a specific tRNA anticodon. EMBO J 28:1362–1373
Ito S, Akamatsu Y, Noma A, Kimura S, Miyauchi K, Ikeuchi Y, Suzuki T, Suzuki T (2014) A single acetylation of 18 S rRNA is essential for biogenesis of the small ribosomal subunit in Saccharomyces cerevisiae. J Biol Chem 289:26201–26212
Sharma S, Langhendries JL, Watzinger P, Kotter P, Entian KD, Lafontaine DL (2015) Yeast Kre33 and human NAT10 are conserved 18S rRNA cytosine acetyltransferases that modify tRNAs assisted by the adaptor Tan1/THUMPD1. Nucleic Acids Res 43:2242–2258
Ohashi Z, Maeda M, McCloskey JA, Nishimura S (1974) 3-(3-Amino-3-carboxypropyl)uridine: a novel modified nucleoside isolated from Escherichia coli phenylalanine transfer ribonucleic acid. Biochemistry 13:2620–2625
Meyer B, Immer C, Kaiser S, Sharma S, Yang J, Watzinger P, Weiss L, Kotter A, Helm M, Seitz HM et al (2020) Identification of the 3-amino-3-carboxypropyl (acp) transferase enzyme responsible for acp3U formation at position 47 in Escherichia coli tRNAs. Nucleic Acids Res 48:1435–1450
Meyer B, Wurm JP, Sharma S, Immer C, Pogoryelov D, Kotter P, Lafontaine DL, Wohnert J, Entian KD (2016) Ribosome biogenesis factor Tsr3 is the aminocarboxypropyl transferase responsible for 18S rRNA hypermodification in yeast and humans. Nucleic Acids Res 44:4304–4316
Lafontaine DL, Preiss T, Tollervey D (1998) Yeast 18S rRNA dimethylase Dim1p: a quality control mechanism in ribosome synthesis? Mol Cell Biol 18:2360–2370
Lafontaine D, Delcour J, Glasser AL, Desgres J, Vandenhaute J (1994) The DIM1 gene responsible for the conserved m6(2)Am6(2)A dimethylation in the 3′-terminal loop of 18 S rRNA is essential in yeast. J Mol Biol 241:492–497
White J, Li Z, Sardana R, Bujnicki JM, Marcotte EM, Johnson AW (2008) Bud23 methylates G1575 of 18S rRNA and is required for efficient nuclear export of pre-40S subunits. Mol Cell Biol 28:3151–3161
Figaro S, Wacheul L, Schillewaert S, Graille M, Huvelle E, Mongeard R, Zorbas C, Lafontaine DL, Heurgue-Hamard V (2012) Trm112 is required for Bud23-mediated methylation of the 18S rRNA at position G1575. Mol Cell Biol 32:2254–2267
Sardana R, White JP, Johnson AW (2013) The rRNA methyltransferase Bud23 shows functional interaction with components of the SSU processome and RNase MRP. RNA 19:828–840
Liang XH, Liu Q, Fournier MJ (2009) Loss of rRNA modifications in the decoding center of the ribosome impairs translation and strongly delays pre-rRNA processing. RNA 15:1716–1728
Taylor AB, Meyer B, Leal BZ, Kotter P, Schirf V, Demeler B, Hart PJ, Entian KD, Wohnert J (2008) The crystal structure of Nep1 reveals an extended SPOUT-class methyltransferase fold and a pre-organized SAM-binding site. Nucleic Acids Res 36:1542–1554
Wurm JP, Duchardt E, Meyer B, Leal BZ, Kotter P, Entian KD, Wohnert J (2009) Backbone resonance assignments of the 48 kDa dimeric putative 18S rRNAmethyltransferase Nep1 from Methanocaldococcus jannaschii. Biomol NMR Assign 3:251-254. [Epub 2009, Sep 25, https://doi.org/10.1007/s12104-009-9187-z]
Buchhaupt M, Kotter P, Entian KD (2007) Mutations in the nucleolar proteins Tma23 and Nop6 suppress the malfunction of the Nep1 protein. FEMS Yeast Res 7:771–781
Eschrich D, Buchhaupt M, Kotter P, Entian KD (2002) Nep1p (Emg1p), a novel protein conserved in eukaryotes and archaea, is involved in ribosome biogenesis. Curr Genet 40:326–338
Buchhaupt M, Meyer B, Kotter P, Entian KD (2006) Genetic evidence for 18S rRNA binding and an Rps19p assembly function of yeast nucleolar protein Nep1p. Mol Gen Genomics 276:273–284
Strunk BS, Karbstein K (2009) Powering through ribosome assembly. RNA 15:2083–2104
Bousquet-Antonelli C, Vanrobays E, Gelugne JP, Caizergues-Ferrer M, Henry Y (2000) Rrp8p is a yeast nucleolar protein functionally linked to Gar1p and involved in pre-rRNA cleavage at site A2. RNA 6:826–843
Peifer C, Sharma S, Watzinger P, Lamberth S, Kotter P, Entian KD (2013) Yeast Rrp8p, a novel methyltransferase responsible for m1A 645 base modification of 25S rRNA. Nucleic Acids Res 41:1151–1163
Sharma S, Hartmann JD, Watzinger P, Klepper A, Peifer C, Kotter P, Lafontaine DLJ, Entian KD (2018) A single N(1)-methyladenosine on the large ribosomal subunit rRNA impacts locally its structure and the translation of key metabolic enzymes. Sci Rep 8:11904
Sharma S, Watzinger P, Kotter P, Entian KD (2013) Identification of a novel methyltransferase, Bmt2, responsible for the N-1-methyl-adenosine base modification of 25S rRNA in Saccharomyces cerevisiae. Nucleic Acids Res 41:5428–5443
Postma L, Lehrach H, Ralser M (2009) Surviving in the cold: yeast mutants with extended hibernating lifespan are oxidant sensitive. Aging (Albany NY) 1:957–960
Motorin Y, Lyko F, Helm M (2010) 5-methylcytosine in RNA: detection, enzymatic formation and biological functions. Nucleic Acids Res 38:1415–1430
Sharma S, Yang J, Watzinger P, Kotter P, Entian KD (2013) Yeast Nop2 and Rcm1 methylate C2870 and C2278 of the 25S rRNA, respectively. Nucleic Acids Res 41:9062–9076
Foster PG, Nunes CR, Greene P, Moustakas D, Stroud RM (2003) The first structure of an RNA m5C methyltransferase, Fmu, provides insight into catalytic mechanism and specific binding of RNA substrate. Structure 11:1609–1620
Hong B, Brockenbrough JS, Wu P, Aris JP (1997) Nop2p is required for pre-rRNA processing and 60S ribosome subunit synthesis in yeast. Mol Cell Biol 17:378–388
Schosserer M, Minois N, Angerer TB, Amring M, Dellago H, Harreither E, Calle-Perez A, Pircher A, Gerstl MP, Pfeifenberger S et al (2015) Methylation of ribosomal RNA by NSUN5 is a conserved mechanism modulating organismal lifespan. Nat Commun 6:6158
Sharma S, Yang J, Duttmann S, Watzinger P, Kotter P, Entian KD (2014) Identification of novel methyltransferases, Bmt5 and Bmt6, responsible for the m3U methylations of 25S rRNA in Saccharomyces cerevisiae. Nucleic Acids Res 42:3246–3260
Basturea GN, Rudd KE, Deutscher MP (2006) Identification and characterization of RsmE, the founding member of a new RNA base methyltransferase family. RNA 12:426–434
Nordlund ME, Johansson JO, von Pawel-Rammingen U, Bystrom AS (2000) Identification of the TRM2 gene encoding the tRNA(m5U54)methyltransferase of Saccharomyces cerevisiae. RNA 6:844–860
Taoka M, Nobe Y, Hori M, Takeuchi A, Masaki S, Yamauchi Y, Nakayama H, Takahashi N, Isobe T (2015) A mass spectrometry-based method for comprehensive quantitative determination of post-transcriptional RNA modifications: the complete chemical structure of Schizosaccharomyces pombe ribosomal RNAs. Nucleic Acids Res 43:e115
Acknowledgments
We thank Jun Yang, Peter Kötter, and Britta Meyer for fruitful discussions and suggestions. We would also like to thank Deutsche Forschungsgemeinschaft (DFG), Goethe-University, Frankfurt/M., and the state of Hesse for financial support.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2022 The Author(s)
About this protocol
Cite this protocol
Sharma, S., Entian, KD. (2022). Chemical Modifications of Ribosomal RNA. In: Entian, KD. (eds) Ribosome Biogenesis. Methods in Molecular Biology, vol 2533. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2501-9_9
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2501-9_9
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2500-2
Online ISBN: 978-1-0716-2501-9
eBook Packages: Springer Protocols