Abstract
Phosphorus (P) is obtained by plants as phosphate (Pi) from the soil and low Pi levels affects plant growth and development. Adaptation to low Pi condition entails sensing internal and external Pi levels and translating those signals to molecular and morphophysiological changes in the plant. In this review, we present findings related to local and systemin Pi sensing with focus the molecular mechanisms behind root system architectural changes and the impact of hormones and epigenetic mechanisms affecting those changes. We also present some of the recent advances in the Pi sensing and signaling mechanisms focusing on inositol pyrophosphate InsP8 and its interaction with SPX domain proteins to regulate the activity of the central regulator of the Pi starvation response, PHR.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phosphorus (P), the second most important component in plant nutrition after nitrogen, is not readily available for plants. It can be bound by calcium in alkaline soils, complexed into charged aluminum and iron oxides in acidic soils, or immobilized by organic material (Kochian et al. 2004; Seguel et al. 2013). Plants absorb P predominately in the form of inorganic phosphate (Pi) (H2PO− 4/HPO4− 2) (Marschner, 2012). In certain conditions, available P in soil solution can be as low as 10 μM (Holford, 1997; do Nascimento et al. 2015). To adapt themselves to fluctuating external and intracellular levels of Pi, plants have developed complex signaling pathways. Even though plants have mechanisms in place to adapt to low Pi condition in the soil, they are not robust enough to prevent the significant yield losses that can occur particularly in agriculturally important crops. Therefore, a deeper understanding of how plants respond to Pi starvation is in order. Extensive work has been carried out in Arabidopsis, and more in rice recently, to decipher the intricate details of the Pi starvation response (PSR) (Wu et al. 2013; López-Arredondo et al. 2014). In this review, we discuss how plants respond to Pi starvation locally as well as systemically. Under local response, we discuss the importance of root system architectural (RSA) changes, the significance of hormones and the recent advances in understanding the epigenetic component driving these responses. Under systemic response, we discuss the signaling mechanism with the inositol pyrophosphates InsP8 and the SPX domain proteins at the core along with PHR1, the central regulator of PSR. Furthermore, a brief account of the Pi responses in yeast, and a comparison of the signaling mechanisms in plants and human is presented.
Root system architecture remodeling
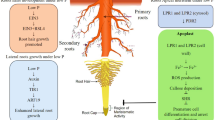
Local signals trigger adaptations of the root system architecture (RSA) such as inhibition of primary root growth and increased lateral root development in response to low external Pi to augment Pi absorption. These root developmental changes are likely governed in the rhizosphere by mechanisms that are localized to the root, as opposed to systemic sensing wherein alterations in the expression of genes involved in Pi transport and distribution, and sensors such as the SPX domain genes are triggered (Chiou and Lin, 2011; López-Arredondo et al. 2014; Wild et al. 2016; Dong et al. 2019; Zhu et al. 2019). A significant amount of work has been reported on RSA changes in Arabidopsis and this section focuses on research emanating from this model plant. The genetic interaction of the ferroxidases LOW PHOSPHATE ROOT1/2 (LPR1/2), which are expressed in the root tip (meristem and root cap), with PHOSPHATE DEFICIENCY RESPONSE2 (PDR2) under Pi-depleted condition plays a significant role in the inhibition of primary root (PR) growth as part of the local Pi sensing mechanism (Svistoonoff et al., 2007; Ticconi et al. 2009). The endoplasmic reticulum (ER)-located PDR2, a P5-type ATPase, functions as a Pi-dependent checkpoint in root development by controlling SCARECROW (SCR) expression in Pi-depleted root, which affects root patterning and stem-cell maintenance (Ticconi et al. 2009). It is likely that LPR1 is trafficked from the ER to the plasma membrane to act as a cell wall-associated ferroxidase that oxidizes Fe2+ to Fe3+ under Pi-depleted condition. In a parallel pathway, the SENSITIVE TO PROTON RHIZOTOXICITY (STOP1) transcription factor activates the ALUMINIUM ACTIVATED MALATE TRANSPORTER 1 (ALMT1), which is integrated into the plasma membrane where it functions as a malate secretion system into the apoplasm (Müller et al. 2015; Mora-Macías et al. 2017; Balzergue et al. 2017; Godon et al. 2019). LPR1-mediated oxidation of Fe2+ leads to accumulation of Fe3+ in the apoplasm, which in complex with malate activates ROS production that promotes callose deposition in the stem cell niche of the PR. Callose deposition hampers trafficking of key transcription factors such as SHR resulting in the loss of stem cell function and inhibition of primary root growth (Müller et al. 2015). A recent study has shown that STOP1 has a broader role to play as ammonium uptake by the AMT1 transporters under Pi starvation triggers rhizosphere acidification of the root surface, which results in the exudation of organic acids due to the accumulation of STOP1 (Tian et al. 2021). Presence of organic acids improves Pi acquisition from insoluble P sources. Also, this study demonstrates that ammonium acts upstream of STOP1 leading to Fe-malate accumulation inhibiting primary root growth in conjunction with LPR1 and LPR2 ferroxidases. The CLAVATA3 (CLV3)/ENDOSPERM SURROUNDING REGION14 (CLE14) peptide is induced by Pi starvation acting downstream of the PDR2–LPR1 module and its expression pattern overlaps Fe distribution in the root apical meristem under Pi-deficient condition (Gutierrez-Alanis et al., 2017). CLV2/PEP1 RECEPTOR2 (PEPR2) receptors perceive CLE14 at the plasma membrane leading to suppress the SHR–SCARECROW (SCR) and PIN-FORMED (PIN)–auxin pathways leading to callose-independent RAM exhaustion (Gutierrez-Alanis et al., 2017). These developments, therefore, underline the importance of the primary root in perceiving Pi levels and in triggering the response mechanisms that ultimately result in the reprogramming of the RSA.
Interestingly, blue light is essential, and may be enough, for primary root growth inhibition under Pi deficiency (Zheng et al. 2019). Blue light can activate the malate-mediated photo-Fenton reaction and hydroxyl radicals are produced by a Fe redox cycle, which is formed by a canonical Fenton reaction in the apoplast. The hydroxyl radicals thus formed cause primary root growth inhibition (Zheng et al. 2019). These observations were further confirmed recently when some of the molecular players involved in root growth inhibition were revealed (Gao et al. 2021). Cryptochromes and other downstream signaling components such as SPA1, COP1 and HY5 play major roles in RSA modulation. Blue light is required for the translocation of shoot-derived HY5 to the root to autoactivate root HY5, which activates LPR1 to promote primary root inhibition under Pi starvation. For a more in-depth view of the developmental responses of the root under Pi deficiency, the readers are referred to the excellent review by Professor Dong Liu (Liu, 2021).
Hormonal control
One of the most important hormones that is implicated in the responses of RSA to Pi starvation is auxin. On Pi starvation, the root tip and lateral root primordia are sites of increased auxin signaling resulting in primary root growth inhibition and enhanced formation of lateral roots (Lopez-Bucio et al., 2002; Nacry et al. 2005) and the changes in RSA could be attributed to auxin redistribution in the roots (Nacry et al. 2005). Exogenous application of auxin showed that Pi-starved plants were more sensitive than plants with adequate Pi supply, and the observed sensitivity is dependent on the auxin receptor TIR1 and the transcription factor ARF19 (Lopez-Bucio et al., 2002; Perez-Torres et al., 2008). For lateral root initiation, as both auxin and TIR1-dependent auxin signaling are important, and the fact that TIR1 is a target of PHR1, it is reasonable to assume a link between lateral root formation, auxin signaling and Pi starvation (Castrillo et al. 2017; Du and Scheres, 2018). Apart from TIR1, genes from the Aux/IAA and ARF gene families are also putative targets of PHR1 (Castrillo et al. 2017). Genes such as ARF7, ARF19 and SLR/IAA14 are involved in the Pi starvation signaling pathway in both Arabidopsis and rice and are required for the regulation of lateral root initiation (Okushima et al. 2005; Narise et al. 2010; Shen et al. 2013; Huang et al. 2018). Other genes implicated in Pi-driven auxin sensitivity are the Arabidopsis Receptor Kinase2 (ARK2)-Ubox/armadillo repeat-containing E3 ligase9 (PUB9), rice OsARF12, OsARF16, and OsPht1;8 genes, and cotton GbWRKYxu1 (Deb et al. 2014; Shen et al., 2013, Wang et al. 2014a, Jia et al. 2017; Xu et al. 2012). Under low Pi condition, auxin-dependent root hair growth is triggered by the transport of auxin from the root apex to differentiation zone with the help of TRYPTOPHAN AMINOTRANSFERASE OF ARABIDOPSIS1 (TAA1, auxin synthesis) and AUX1 (auxin transport) (Bhosale et al. 2018). Furthermore, in trichoblasts with elevated auxin levels, auxin induces the transcription of transcription factors ARF19, ROOT HAIR DEFECTIVE 6-LIKE2 (RSL2), and RSL4, which mediate a gene expression cascade promoting root hair elongation for improved Pi foraging (Bhosale et al. 2018). In a parallel study, the authors also conclude from studies in rice that auxin and OsAUX1 are important in augmenting root foraging for Pi in a mechanism similar to Arabidopsis (Giri et al. 2018).
Ethylene (ET), cytokinin (CK), gibbberellin (GA), strigolactone (SL) and jasmonate (JA) are other hormones that play significant roles in integrating environmental cues and RSA. ET is a positive regulator of root hair growth and root elongation under Pi starvation (Borch et al. 1999; Ma et al. 2003; Zhang et al. 2003). When extracellular Pi is low, ET biosynthesis is enhanced and ETHYLENE-INSENSITIVE3 (EIN3), a key transcription factor in ET signalling, interacts with the promoters of genes targeted by RSL4 (Song et al. 2016). RSL4 being an important factor in root hair development, along with its homologs, further enhances their expression levels resulting in increased root hair formation (Song et al. 2016). Additionally, EIN3, in complex with FHY3, FAR1 and HY5, binds to the PHR1 promoter to regulate its expression and drive PSRs (Liu et al. 2017). The genes ACS2, ACS4, ACS6 and ACO encoding for enzymes necessary for the conversion of AdoMet to ACC (1-aminocyclopropane-1-carboxylate) and ACC to ethylene, are induced under Pi starvation in Arabidopsis (Morcuende et al. 2007; Thibaud et al. 2010; Lei et al. 2011). CK is critical in balancing the root:shoot ratio, and under Pi starvation, CK increases root:shoot ratio, as root growth is favored over shoot and CK concentrations are reduced under Pi starvation (Kuiper and SteingrÖVer, 1991; Franco-Zorrilla et al. 2002). Pi starvation-inducible genes are also deregulated by CK leading to an increase in Pi concentration (Martin et al. 2000; Franco-Zorrilla et al. 2002; Wang et al. 2006). Among the different types of CK, trans-zeatin is reduced under Pi starvation while cis-zeatin and cis-zeatin roboside are increased significantly (Silva-Navas et al. 2019). The two latter types are PHR1-dependent and cis-zeatin regulates genes involved in cell growth and root hair elongation, and both cause an increase in Pi concentrations in roots. Interestingly, root growth and lateral root formation is enhanced when the cis-zeatin:trans-zeatin ratio is higher, and cis-zeatin is essential for root hair elongation and Pi allocation to the root and shoot under low Pi condition (Silva-Navas et al. 2019).
SLs are plant metabolites that are derived from carotenoids and are synthesized mainly in the root (Yoneyama et al. 2013; Al-Babili and Bouwmeester 2015; Lopez-Obando et al. 2015). Under Pi starvation, SLs inhibit shoot branching and regulates important RSA changes such as lateral root formation and root hair density (Umehara et al. 2008, Kapulnik et al. 2011). Not only do the SLs move acropetally as a systemic signal while controlling shoot branching under Pi-depleted condition, they can also be synthesized in the shoot to control branching (Beveridge et al. 1997; Foo et al. 2001; Sorefan et al. 2003). Auxin-mediated control of RSA could also involve GA, and the core components of the GA signaling pathway, the DELLA proteins, contribute to anthocyanin accumulation and RSA changes and are not likely to be involved in the expression of PSI genes or Pi uptake (Jiang et al. 2007). However, a later study showed that the transcription factor MYB62 could regulate PSR through alterations in GA metabolism and signaling (Devaiah et al. 2009). JA is known to inhibit root growth and specifically controls lateral root growth and root hair development in Arabidopsis and rice (Cai et al. 2014; Wang et al. 2002). A recent study showed that overexpression of the Pi-inducible OsJAZ11 reduced the PSR under Pi-depleted condition along with increased primary add seminal root elongation (Pandey et al. 2021). OsJAZ11 is transcriptionally regulated and, interestingly, also physically interacts with OsSPX1. VIH2 has been shown to regulate the synthesis of inositol pyrophosphate InsP8 and JA-dependent defense response in Arabidopsis, and InsP8 has been implicated as a signaling molecule in driving PSR in Arabidopsis (Laha et al. 2015; Dong et al. 2019; Zhu et al. 2019). Though studies on the role of JA in Pi sensing are few and far between, with these recent developments, several more studies are expected.
Epigenetic control
Gene regulation in plants under Pi starvation is well studied at the transcriptional and posttranscriptional levels (López-Arredondo et al. 2014). Studies in Arabidopsis and rice have shown that for proper PSR, epigenetic control as an added level of regulation is essential (Yong-Villalobos et al. 2015; Secco et al. 2015; Table 1). Yong-Villalobos and co-workers showed that in the Arabidopsis drm1, drm2, and cmt3 single mutants the drm1 drm2 cmt3 triple mutant the primary root length was shorter than the wild-type under Pi-depleted condition. A similar observation was also made for lateral root density. Observations made in this study clearly showed that epigenetic marks, DNA methylation, are key to establishing proper morphophysiological PSR. ARP6 is a nuclear actin-related protein and in Arabidopsis, it is required for the proper deposition of H2A.Z at several PSR genes. Loss-of-function ARP6 mutation resulted in the loss of H2A.Z at the target loci, which led to shortening of primary roots and increased number and length of root hairs (Smith et al. 2010). Further, a reduction in H2A.Z occupancy in chromatin was reduced in atipk1–1 mutant leading to an increase in the number of Pi starvation-inducible genes (Kuo et al. 2014). Members of the histone deacetylase family, HDA6, HDA9 and HDA19, are involved in the regulation of root cell length (Chen et al. 2015). Recently, histone deacetylase 1 (HDC1), was found to affect primary root growth and LPR1/2-mediated iron deposition at the root tips under low Pi condition (Xu et al. 2020). Therefore, HDC1 is an important component of the local Pi response acting as a negative regulator of RSA changes under Pi-depleted condition. These recent studies clearly show that chromatin and DNA methylation pattern remodeling are active components of a plant’s response to Pi-depleted condition or could possibly be due to the changes that occur during the epigenetic regulation of PSR genes (Yong-Villalobos et al. 2015).
SPX domain proteins: the sensors
The SPX domain (Pfam PF03105) plays an increasingly important role in elucidating the mechanism of Pi homeostasis (Secco et al. 2012). This was initially identified in yeast and is named after the Suppressor of Yeast gpa1 (Syg1), which is a negatively affected mating pheromone signal in yeast when it is truncated, a cyclin dependent kinase (CDK) inhibitor phosphatase 81 (Pho81), and the human Xenotropic and Polytropic Retrovirus receptor1(Xpr1). Xpr1 regulates polyphosphate in platelets and was implicated in thrombosis in vivo (Mailer et al. 2021), and influences intracellular ATP levels because of the repression of IPK6 (Moritoh et al. 2021). The SPX domain is quite conserved and hydrophilic, usually located at the N-terminus of eukaryotic proteins. The SPX domain can be divided into three subdomains, each containing 30–40 amino acids, with an average length of 165 amino acids. Subdomains are separated from each other by low similarity regions (Liu et al. 2018). Proteins containing the SPX domain in plants can be divided into four groups: the first group contains only the SPX domain, such as AtSPX1 in Arabidopsis (Duan et al. 2008), OsSPX1/2 (Wang et al. 2009) and OsSPX4 (Lv et al. 2014) in rice. The second is proteins containing one SPX domain and one EXS domain, such as Arabidopsis AtPHO1, which is an ortholog of the human XPR1, regulates Pi transport from the root to the shoot (Wild et al. 2016). The third type is the protein containing an SPX domain and RING domain, such as AtNLA, a ubiquitin connection containing both SPX and RING domain, requires UBC24 to mediates polyubiquitination of Pht1;4 (Park et al. 2014). The fourth category is proteins containing one SPX domain and one MFS domain, such as OsVPE1 and OsVPE2, which localize on the tonoplast and mediate Pi efflux from the vacuole into cytosol (Wang et al. 2015).
The SPX domain has been reported as a Pi sensor and SPX proteins interact with PHR homologous proteins and regulate their function by binding to them under Pi-replete condition in the presence of inositol pyrophosphates (PP-InsPs) and prevents the PHR proteins from entering the nucleus (Liu et al. 2010; Shi et al. 2014; Wang et al. 2014b; Puga et al. 2014; Lv et al. 2014; Wild et al. 2016; Dong et al. 2019; Osorio et al. 2019). Similar mechanisms have been reported in other organisms as well. In the pathogenic process of human-pathogenic fungus Cryptococcus neoformans, InsP7 synthesized by Kcs1 regulates fungal virulence by binding to a conserved lysine surface cluster in the SPX domain of Pho81 (Desmarini et al. 2020). SPX negatively regulates Pi uptake and metabolism through PSR regulator (PHR) as an intermediate in the marine phytoplankton, Phaeodactylum tricornutum (Zhang et al. 2021). While SPX proteins have largely been implicated in the Pi response mechanism, GmSPX5 from soybean was shown to play a key role in nodule adaptation to low Pi condition (Zhuang et al. 2021) adding a new dimension to the SPX protein function.
Inositol pyrophosphates: the signaling molecules
The PP-InsPs are novel and unique signaling molecules that are implicated in a wide variety of biological processes in eukryotes (Azevedo and Saiardi 2017; Shears 2018; Lorenzo-Orts et al. 2020). PP-InsPs are high-energy pyrophosphate containing molecules obtained by the additional phosphorylation of inositol polyphosphates. The inositol polyphosphate InsP6, is considered a storage molecule, and is accumulated in the form of phytic acid particularly in the seeds (Secco et al. 2017). From InsP6, phosphorylation at positions 5 leads to the formation of PP-InsP, InsP7, catalyzed by the enzyme inositol phosphate kinase ITPK1(Riemer et al. 2021). InsP7 is then converted to InsP8 by VIH1 and VIH2, and these PP-InsPs have been reported to act as signaling molecules in plants (Laha et al. 2015; Wild et al. 2016; Dong et al. 2019; Zhu et al. 2019; Fig. 1A).
Biosynthesis pathways of PP-InsP and IP8 signalling in plants and humans. A Schematic model of InsP and PP-InsP biosynthesis pathway in humans (via Lipid-dependent pathway) and plants (via Lipid-dependent and -independent pathways). Key enzymes in InsP and PP-InsP pathways are shown for each reaction. B Pi sensing and InsP8 signaling in human/animal system is presented. C Pi sensing and InsP8 signalling under Pi-replete and Pi-depleted conditions are presented
InsP8-SPX- PHR: the interaction
PHR1, containing the MYB-like DNA-binding domain and a coiled-coil domain, belongs to the GARP transcription factor family and is homologous to the PSR1, which was shown to regulate P metabolism in Chlamydomonas reinhardtii (Wykoff et al. 1999; Rubio et al. 2001). PHR1 binds to an imperfect palindromic motif (5′-GNATATNC-3′) known as the P1BS element or the PHR1-binding sequence, which is found in most of the Pi starvation-inducible genes. PHL1 (PHR1-LIKE 1), a paralog of PHR1, functions redundantly to regulate the expression of a subset of PSR genes (Bustos et al. 2010). Orthologous genes of PHR1 have been identified in plants including rice (Zhou et al. 2008), maize (Wang et al. 2013a), wheat (Wang et al. 2013b) and soybean (Lu et al. 2020). As previously mentioned, PHR function is regulated by the SPX domain containing proteins, which act as cellular receptors for the PP-InsPs.
VIH1 and VIH2 are homologous to the yeast and animal Vip1/PPIP5Ks (Desai et al. 2014; Laha et al. 2015) and are bifunctional kinase/phosphatase enzymes (Zhu et al. 2019). Previous studies have suggested that the preferred ligand for SPX proteins is InsP8 as the vih1 vih2 double mutants show severe defects in growth phenotype and Pi signaling, and under in vivo conditions, InsP8 restores the interaction between SPX1 and PHR1 better than 5-InsP7 (Dong et al. 2019; Zhu et al. 2019; Ried et al. 2021). The 5-InsP7 isomer, the precursor of InsP8, is generated by ITPK1, the InsP6 kinase in planta (Laha et al. 2019). We along with others showed that InsP7 was phosphorylated by VIH1 and VIH2 to generate InsP8 (Laha et al. 2015; Dong et al. 2019; Zhu et al. 2019). A recent study has shown that under Pi starvation 5-InsP7 and 1/3-InsP7 were also reduced, and recovered on Pi resupply (Riemer et al. 2021). Riemer and co-workers also showed that synthesis of 5-InsP7 and InsP8 in planta is Pi-dependent mediated by ITPK1. The authors also provide genetic evidence for the interdependence of ITPK1 and VIH2 for the maintenance of Pi homeostasis in plants. In the context of VIH1 and VIH2, under Pi-replete condition phosphatase activity is allosterically regulated by Pi resulting in an increase in InsP8 level culminating in the formation of a InsP8-SPX- PHR complex (Gu et al., 2017a; Zhu et al. 2019). Kinase activity is reduced under Pi-deficient condition as ATP levels drop leading to a decrease in the levels of InsP8 and the complex breaks down (Dong et al. 2019; Zhu et al. 2019). The released PHR can now transactivate the PSR genes by binding to the P1BS motif as a dimer (Rubio et al. 2001). Recent work has shown that the CC domain of PHRs is targeted by the InsP8-SPX1 complex and regulates the promoter binding ability of PHRs as dimers through their SPX receptors (Ried et al. 2021). Interestingly, Zhou and co-workers (Zhou et al. 2021) show that the under Pi-replete condition, InsPs bind to the SPX protein to stabilize the helix α1 structure, which enables the InsPs to allosterically decouple the PHR protein dimer. The SPX protein is then able to interact with the MYB domain and blocks PHR protein from binding to the promoters of the PSR genes. Therefore, SPX inhibition of PHR activity occurs at two levels of oligomerization and DNA binding (Zhou et al. 2021). These recent advances in Pi signaling have opened new avenues in plant research to develop crops with higher Pi use efficiency.
Pi sensing: the yeast response
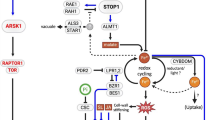
Pi sensing in yeast is a well-studied mechanism and in Saccharomyces cerevisiae, the Pi signaling pathway (also called the PHO pathway) is activated under Pi starvation and several genes that are involved in the uptake and storage of Pi are upregulated (Bergwitz and Jüppner, 2010; Secco et al. 2012; Sabbagh, 2013; Sengottaiyan et al., 2011; Conrad et al. 2014). Pho4, along with its coactivator Pho2, is a key factor in mediating PSR in yeast by regulating the expression of the Pi-responsive genes (Magbanua et al. 1997a; Fig. 2). In fact, the Pho4-Pho2 interaction enhances the affinity for Pho4-binding sites in the promoters of PHO genes (Shao et al. 1996; Magbanua et al. 1997a, b). Pho4 activity and subcellular localization is determined by the phosphorylation of its serine residues by the cyclin-dependent kinase complex Pho80-Pho85 (Toh-e et al. 1988; Kaffman et al. 1994; O’Neill et al. 1996). As a requisite transcription factor, it is dephosphorylated and accumulated in the nucleus to induce the expression of Pho4-dependent PHO genes (Zhou and O’Shea 2011).
A schematic showing Pi sensing and regulation of gene expression under Pi-replete and -depleted condition in yeast. Pho81 negatively regulates the Pho80-Pho85 complex and its binding to the complex is Pi-independent. Under Pi-replete condition, Pho81-Pho80-Pho85 complex phosphorylates Pho4 and it cannot activate the PHO genes. Under Pi-depleted condition, InsP7 interacts allosterically with the Pho81-Pho80-Pho85 tertiary complex preventing Pho4 from being phosphorylated by the complex. Lack of phosphorylation allows Pho4 to activate the genes in the PHO pathway
The cyclin-dependent kinase inhibitor Pho81 plays a key role in the PHO pathway as a negative regulator of the Pho80-Pho85 complex by binding to it independent of Pi availability and inhibits kinase activity only under Pi-limiting condition (Schneider et al. 1994; Ogawa et al. 1995; O’Neill et al. 1996; Huang et al. 2001). A conformational change is induced in Pho81 by the allosteric interaction of PP-InsPs with the Pho81-Pho85-Pho80 tertiary complex, and this change prevents Pho4 from accessing the Pho85-Pho80 kinase active site (Lee et al. 2007; Lee et al. 2008). Unphosphorylated Pho4 enters the nucleus and activates the transporters (Fig. 2).
There are five transporters in yeast viz., Pho84, Pho87, Pho89, Pho90 and Pho91. While Pho84 and Pho89 are high-affinity transporters, Pho87, Pho90 and Pho91 are low affinity transporters. Under low internal Pi, Pho84 is activated and there is an increase in Pi uptake generating a negative-feedback loop. Interestingly, Spl2 is also upregulated, which negatively regulates Pho87 and Pho90 low-affinity transporters (Wykoff et al. 2007). Levy et al. (2011) have shown that a dual-transporter system with high- and low-affinity transporters, could extend the preparation for starvation and enables yeast cells to recover subsequently. Therefore, as Pi concentration depletes gradually in the environment, the transition of yeast cells from Pi-replete to Pi-depleted condition is longer, and the cells can better adapt themselves to Pi deprivation making recovery more effective on resupplying Pi. Among the transporters, Pho84 acts as a Pi sensor and transporter, and on resupplying Pi after prolonged starvation it is internalized from the plasma membrane and degraded. Pho4 is phosphorylated on Pho84 degradation after Pi resupply by Pho80-Pho85, and exported to the cytoplasm and the PHO genes are repressed (Giots et al. 2003; Mouillon and Persson 2006; Samyn et al. 2016).
The PHO pathway is also involved in the synthesis of polyphosphates, which depends on the Vacuolar Transport Chaperone (Vtc) complex in yeast. This comprises Vtc1, Vtc2, Vtc3, and Vtc4, where Vtc4 is the catalytic subunit (Hothorn et al. 2009). Vtc1/2/4 and Vtc1/3/4 are two different Vtc complexes found in the endoplasmic reticulum and vacuole, respectively, under Pi-replete condition, and both complexes are found in the vacuole under Pi-depleted condition (Hothorn 2009). Among the various PP-InsPs, 5-InsP7 regulates VTC in vivo (Gerasimaite et al., 2017). VTC synthesizes InsPs under Pi-replete condition, and the cells use the reserve stocks under Pi-depleted condition. Higher activity of VTC leads to Pi depletion in the cells triggering the PHO pathway under Pi-replete condition (Desfougères et al. 2016). Thus the yeast Pi starvation response mechanism shares similarities with that found in the plant systems.
Plant and human InsP8 synthesis and Pi sensing: the commonalities
Inositol phosphates (InsPs) are gaining attention for broadening our understanding of energy metabolism and nutrient sensing in humans as well as plants (Chakraborty et al., 2011; Hatch and York 2010). Although they play conserved functions in various eukaryotic cellular processes, their biosynthesis and their function in Pi sensing (the most common and widely known attribute of this family) (Lee et al. 2020), differs between humans and plants with a few exceptions detailed in this section.
Biosynthesis of bioactive inositols (inositol polyphosphates) can be via lipid-dependent or lipid-independent pathway (Fig. 1A). Lipid-dependent pathway is shared between animals and plants where initiation involves phosphorylation of phosphatidylinositols (PtdIns) (Berridge and Irvine 1989; Stevenson-Paulik et al. 2002). Phospholipase C (PLC) is a conserved enzyme in humans and plants for conversion of phosphatidylinositol 4,5 bisphosphate (PtdIns(4,5)P2) into diacylglycerol (DAG) and Ins(1,4,5)P3. This initiation process is carried out in the plasmamembrane and thereafter Ins(1,4,5)P3 is released into the cytosol where successive addition of phosphates is mediated by Inositol Polyphosphate Multikinases (IPMK) in animals, and by the Inositol Pentakisphosphate 2-Kinase (IPK2) in plants (Berridge and Irvine 1989; Lee et al., 2020; Fig. 1).
Lipid-independent pathway for InsP6 synthesis has been characterized well for plants compared to animals. In the first step of the pathway, myo-inositol kinase (MIK) was reported to be involved in phosphorylation of inositol at 3 position to form InsP (Kim and Tai 2011; Shi et al. 2005). Multiple studies showed that MIK gene functionality is almost conserved in all the species of plants (Loewus and Murthy, 2000; Shi et al. 2005). Contrarily, in humans, MIK ortholog with only limited homologous regions were observed for one ribokinase as well as for various adenokinase genes. Hence, either human MIK gene is distinct compared to that of plants or they do not require MIK gene as myo-inositol phosphate synthase (MIPS) catalyzes the reaction of glucose-6-phosphate conversion to Inositol 3-phosphate (Funkhouser and Loewus 1975). Similar to MIK, another enzyme LPA1 (Low Phytic Acid) involved in the formation of InsP2 from InsP in plants also lacks a definitive ortholog in the human genome. Recently, Desfougères and colleagues reported that plants and animals both have Inositol 1,3,4-Trisphosphate 5/6-Kinases (ITPKs) to avoid the need for LPA1 enyzme for final product synthesis (Desfougères et al. 2019). Finally, the conversion of InsP5 to InsP6 requires IPK1 in plants and its counterpart in humans is HsIPPK (Kuo et al. 2018; Park et al. 2018). Inositol hexakisphosphate (InsP6) is the major source of Pi storage in seeds. Owing to Multidrug Resistance Protein 5 (MRP5) which activates InsP6 transport to the vacuolar membrane, InsP6 can be stored in higher amounts in plants (Freed et al. 2020; Raboy 2001).
Pyrophosphorylation of InsP6 leads to the synthesis of PP-InsPs, which contain highly energetic phosphoanhydride bonds. As mentioned earlier, InsP6 can synthesize 1,5-InsP8 via 1-InsP7 or 5-InsP7 intermediates, with 5-InsP7 being the major route of InsP8 synthesis. Previously, PPIP5Ks (VIH/VIP in plants) were assumed to inter-convert InsP6 to InsP7 and InsP7 to InsP8 in two successive steps (Campo and San Segundo 2020; Chakraborty et al. 2011). Recently published work provided evidence that ITPKs are the responsible kinases for InsP6 conversion to InsP7 in plants in lieu of PPIP5Ks, whereas in humans this step involves IP6Ks (Adepoju et al., 2019). PPIP5Ks are mainly involved in the interconversion of 1-InsP7 to 1,5-InsP8 and as mentioned above this route is the major source of InsP8 synthesis. These kinases are attracting attention for various therapeutics in humans and also for stress alleviation in plants (Dollins et al. 2020; Gu et al. 2016).
PPIP5Ks present conserved structures in both humans and plants with KD (N-terminal ATP-grasp kinase domain), PD (C-terminal phosphatase domain) and PH (Pleckstrin-homology) domains. Despite limited sequence identity of AtITPKs and PPIP5Ks, KD domain is conserved in these two kinases (Wang et al. 2012, 2014c). PH domains are common among signaling proteins and have affinity for phospholipids and their derived head groups. Arginine residue is required for ligand binding in human PH domain of PPIP5Ks whereas in Arabidopsis, the purpose is served by a lysine residue (Gokhale et al. 2011). Moreover, AtPIPP5Ks and HsPIPP5Ks also share a C-terminal intrinsically disordered region, crucial for protein-protein interactions (Machkalyan et al. 2016).
PD-KD domains render unique features to PPIP5Ks and their balanced activities are critical for growth, development and Pi sensing in plants (Zhu et al. 2019). Under certain conditions in humans, PD can restrict the KD activity and PD mutation of HsPIPP5K resulted in autosomal recessive non-syndromic hearing loss. Hence, the importance of PD-KD balance in humans was demonstrated and this balance can be disrupted by Pi, which is a strong inhibitor of PD domain in one of HsPPIP5K2 isoform (Gu et al., 2017a). Briefly, PPIP5Ks activity is affected by external Pi in humans. In plants, this is not proven yet as the PD domain of PPIP5Ks is recalcitrant to enzymatic observations. However, in plants AtPPIP5K loss-of-function mutations attributed to increased InsP7 and decreased InsP8 levels with stunted growth and Pi sensing defects (Dong et al. 2019; Zhu et al. 2019).
As discussed above, InsPs and PP-InsPs synthesis pathways exhibit similarities and differences among humans and plants at various levels with common downstream InsP8 synthesis (Fig. 1B, C). InsP8 is a proven metabolic messenger of intra-cellular Pi status across humans and plants (Dong et al. 2019; Li et al. 2020). Yet, this signaling pathway also differs in terms of enzymatic players in both kingdoms (Lee et al. 2020). Among all eukaryotes PP-InsP-SPX interaction is a crucial signaling module for the control of cellular Pi homeostasis. SPX domain contains 135–380 amino acids and there are four groups of SPX domain-containing proteins in plants (Secco et al. 2012; Wang et al. 2021). Contrarily, in the human genome, the only SPX domain-containing protein with cellular Pi efflux control is the Xenotropic and Polytropic Retrovirus Receptor 1 (XPR1).
Prior to the Wild et al. study, the SPX domain was considered to have a strong binding affinity directly to Pi. This report changed this notion and showed higher binding affinity of the SPX domain to InsP and PP-InsP compared to Pi and PPi (Wild et al. 2016). In plants, under Pi-sufficient condition, PP-InsPs expedite SPX interaction with PHR1 and its homologs to prevent PSR genes expression. From available data, it is safe to assume that high cellular Pi concentrations led to VIH-mediated InsP8 synthesis, which eventually facilitated SPX-PHR1 binding and finally transcription of PSR genes is turned off. On the other hand, low cellular Pi leads to a decrease in InsP8 levels and PHR1 is available for the activation of PSR genes (Dong et al. 2019; Zhu et al. 2019).
In earlier studies on human cells, cellular InsP7 as well as InsP8 levels were considered to be correlated with levels of cellular Pi (Lonetti et al. 2011; Gu et al., 2017b). Recent genetic studies with PPIPK5s knock out mutants in humans alleviated XPR- mediated Pi efflux by reduction in InsP8 levels. Restoration of Pi efflux resulted in Pi efflux rescue, hence, highlighting InsP8 as the main cellular Pi-sensing signal (Li et al. 2020). Therefore, studies from humans and plants argue that InsP8 is the ligand that modulates SPX domain interaction which in turn controls PSR gene expression by interacting with a transcription factor or controls Pi efflux directly in humans.
Challenges and perspectives
PSR is multifaceted. Therefore, understanding the mechanisms and making use of the knowledge gained in addressing problems of nutrient stress necessitates a multifaceted approach. With significant progress being made in Pi signaling vis-a-vis the signaling molecules, PP-InsPs, and the sensor, SPX domain, the diversity of SPX domain-containing proteins in plants need to be explored further as presence of an expanded SPX family compared to humans would help define other functions in this sessile group of organisms. Similarly, PP-InsP-SPX interaction should be characterized to define interaction components of other SPX-domain containing protein members. Another challenge is whether a single enzyme should be targeted in PP-InP biosynthesis pathway or PP-InP interactions with other proteins should be focused to have a better view of PP-InsP targets and their interacting mechanisms (Furkert et al. 2020; Lee et al. 2021). Moreover, a deeper understanding of these mechanisms would be beneficial for the development of new therapeutics in human disease as well as in understanding plant mechanisms related to these PP-InsP interactions. In recent times, work on epigenetic regulation in PSR is gathering momentum. Reduced cost of sequencing has enabled large-scale sequencing studies particularly on DNA methylation. Since most work has focused on Arabidopsis and/or rice to a large extent, epigenomic studies should be extended to other crops as well and better analysis methods are required to obtain a better understanding of the mechanisms that drive epigenetic changes in crops under low Pi stress. This would require concerted efforts as most of the breakthrough studies are still being made in Arabidopsis and it is difficult to fathom how it would translate to other crops, and in particular, how knowledge on these discovered mechanisms would translate under field conditions, which is a completely different ecosystem compared to controlled laboratory conditions. The other aspect of utilizing epigenetic mechanisms in plant breeding efforts is the need to understand transgenerational inheritance of the epigenetic marks. Therefore, a thorough understanding mechanisms that underlie low Pi-stress resilience is required to develop crops that are tolerant to such nutrient stress by engineering the epigenome.
Availability of data and materials
Not applicable.
Abbreviations
- P:
-
Phosphorus
- Pi:
-
Phosphate
- PSR:
-
Phosphate stress response
- RSA:
-
Root system architecture
- Fe:
-
Iron
- ET:
-
Ethylene
- CK:
-
Cytokinin
- GA:
-
Gibberellin
- SL:
-
Strigolactone
- JA:
-
Jasmonate
- InsP:
-
Inositol polyphosphates
- PP-InsP:
-
Inositol pyrophosphates
- TF:
-
Transcription factor
References
Adepoju O, Williams SP, Craige B, Cridland CA, Sharpe AK, Brown AM, Land E, Perera IY, Mena D, Sobrado P, Gillaspy GE (2019) Inositol trisphosphate kinase and Diphosphoinositol Pentakisphosphate kinase enzymes constitute the inositol pyrophosphate synthesis pathway in plants. bioRxiv. https://doi.org/10.1101/724914
Al-Babili S, Bouwmeester HJ (2015) Strigolactones a novel carotenoid-derived plant hormone. Annu Rev Plant Biol 66:161–186. https://doi.org/10.1146/annurev-arplant-043014-114759
Azevedo C, Saiardi A (2017) Eukaryotic phosphate homeostasis: the inositol pyrophosphate perspective. Trends Biochem Sci 42(3):219–231. https://doi.org/10.1016/j.tibs.2016.10.008
Balzergue C, Dartevelle T, Godon C, Laugier E, Meisrimler C, Teulon JM, Creff A, Bissler M, Brouchoud C, Hagège A, Müller J, Chiarenza S, Javot H, Becuwe-Linka N, David P, Péret B, Delannoy E, Thibaud MC, Armengaud J, Abel S, Pellequer JL, Nussaume L, Desnos T (2017) Low phosphate activates STOP1-ALMT1 to rapidly inhibit root cell elongation. Nat Commun 8(1):15300. https://doi.org/10.1038/ncomms15300
Bergwitz C, Jüppner H (2010) Regulation of phosphate homeostasis by PTH, vitamin D, and FGF23. Annu Rev Med 61:91–104
Berridge MJ, Irvine RF (1989) Inositol phosphates and cell signalling. Nature 341:197–205
Beveridge CA, Symons GM, Murfet IC, Ross JJ, Rameau C (1997) The rms1 mutant of pea has elevated indole-3-acetic acid levels andreduced root-sap zeatin riboside content but increased branching con-trolled by graft-transmissible signal(s). Plant Physiol 115:1251–1258
Bhosale R, Giri J, Pandey BK, Giehl RFH, Hartmann A, Traini R et al (2018) A mechanistic framework for auxin dependent Arabidopsis root hair elongation to low external phosphate. Nat Commun 9:1409
Borch K, Bouma TJ, Lynch JP, Brown KM (1999) Ethylene: a regulator of root architectural responses to soil phosphorus availability. Plant Cell Environ 22(4):425–431. https://doi.org/10.1046/j.1365-3040.1999.00405.x
Bustos R, Castrillo G, Linhares F, Puga MI, Rubio V, Pérez-Pérez J, Solano R, Leyva A, Paz-Ares J (2010) A central regulatory system largely controls transcriptional activation and repression responses to phosphate starvation in Arabidopsis. PLoS Genet 6(9):e1001102. https://doi.org/10.1371/journal.pgen.1001102
Cai XT, Xu P, Zhao PX, Liu R, Yu LH, Xiang CB (2014) Arabidopsis ERF109 mediates cross-talk between jasmonic acid and auxin biosynthesis during lateral root formation. Nat Commun 5(1):5833. https://doi.org/10.1038/ncomms6833
Campo S, San Segundo B (2020) Systemic induction of phosphatidylinositol-based signaling in leaves of arbuscular mycorrhizal rice plants. Sci Rep 10(1):1–17. https://doi.org/10.1038/s41598-020-72985-6
Castrillo G, Teixeira PJPL, Paredes SH, Law TF, de Lorenzo L, Feltcher ME, Finkel OM, Breakfield NW, Mieczkowski P, Jones CD, Paz-Ares J, Dangl JL (2017) Root microbiota drive direct integration of phosphate stress and immunity. Nature 543:513–518. https://doi.org/10.1038/NATURE21417
Chakraborty A, Kim S, Snyder SH (2011) Inositol pyrophosphates as mammalian cell signals. Sci Signal 4(188):re1
Chandrika NN, Sundaravelpandian K, Yu SM, Schmidt W (2013) ALFIN-LIKE 6 is involved in root hair elongation during phosphate deficiency in Arabidopsis. New Phytol 198(3):709–720. https://doi.org/10.1111/nph.12194
Chen CY, Wu K, Schmidt W (2015) The histone deacetylase HDA19 controls root cell elongation and modulates a subset of phosphate starvation responses in Arabidopsis. Sci Rep 5(1):15708. https://doi.org/10.1038/srep15708
Chiou T-J, Lin S-I (2011) Signaling network in sensing phosphate availability in plants. Annu Rev Plant Biol 62:185–206. https://doi.org/10.1146/annurev-arplant-042110-103849
Conrad M, Schothorst J, Kankipati HN, Van Zeebroeck G, Rubio-Texeira M, Thevelein JM (2014) Nutrient sensing and signaling in the yeast Saccharomyces cerevisiae. FEMS Microbiol Rev 38:254–299. https://doi.org/10.1111/1574-6976.12065
Deb S, Sankaranarayanan S, Wewala G, Widdup E, Samuel MA (2014) The S-Domain Receptor Kinase Arabidopsis Receptor Kinase2 and the U Box/Armadillo Repeat-Containing E3 Ubiquitin Ligase9 module mediates lateral root development under phosphate starvation in Arabidopsis. Plant Physiol 165:1647–1656
Desai M, Rangarajan P, Donahue JL, et al (2014) Two inositol hexakisphosphate kinases drive inositol pyrophosphate synthesis in plants. Plant Journal 80:642–653. https://doi.org/10.1111/tpj.12669
Desfougères Y, Gerasimaitë RU, Jessen HJ, Mayer A (2016) Vtc5, a novel subunit of the vacuolar transporter chaperone complex, regulates polyphosphate synthesis and phosphate homeostasis in yeast. J Biol Chem. 291:22262–22275. https://doi.org/10.1074/jbc.M116.746784
Desfougères Y, Wilson MS, Laha D, Miller GJ, Saiardi A (2019) ITPK1 mediates the lipid-independent synthesis of inositol phosphates controlled by metabolism. Proc Natl Acad Sci 116(49):24551–24561
Desmarini D, Lev S, Furkert D, Crossett B, Saiardi A, Kaufman-Francis K, Li C, Sorrell TC, Wilkinson-White L, Matthews J, Fiedler D, Djordjevic JT (2020) IP(7)-SPX domain interaction controls fungal virulence by stabilizing phosphate signaling machinery. mBio 11(5):e01920
Devaiah BN, Madhuvanthi R, Karthikeyan AS, Raghothama KG (2009) Phosphate starvation responses and gibberellic acid biosynthesis are regulated by the MYB62 transcription factor in Arabidopsis. Mol Plant 2(1):43–58. https://doi.org/10.1093/mp/ssn081
do Nascimento C, Pagliari P, Schmitt D et al (2015) Phosphorus concentrations in sequentially fractionated soil samples as affected by digestion methods. Sci Rep 5:17967. https://doi.org/10.1038/srep17967
Dollins DE, Bai W, Fridy PC, Otto JC, Neubauer JL, Gattis SG, Mehta KPM, York JD (2020) Vip1 is a kinase and pyrophosphatase switch that regulates inositol diphosphate signaling. Proc Natl Acad Sci U S A 117:9356–9364
Dong J, Ma G, Sui L, Wei M, Satheesh V, Zhang R, Ge S, Li J, Zhang T-E, Wittwer C, Jessen HJ, Zhang H, An GY, Chao DY, Liu D, Lei M (2019) Inositol pyrophosphate InsP8 acts as an intracellular phosphate signal in Arabidopsis. Mol Plant 12(11):1463–1473. https://doi.org/10.1016/j.molp.2019.08.002
Du Y, Scheres B (2018) Lateral root formation and the multiple roles of auxin. J Exp Bot 69(2):155–167. https://doi.org/10.1093/JXB/ERX223
Duan K, Yi K, Dang L, Huang H, Wu W, Wu P (2008) Characterization of a sub-family of Arabidopsis genes with the SPX domain reveals their diverse functions in plant tolerance to phosphorus starvation. Plant J 54(6):965–975. https://doi.org/10.1111/j.1365-313X.2008.03460.x
Foo E, Turnbull CG, Beveridge CA (2001) Long-distance signaling and the control of branching in the rms1 mutant of pea. Plant Physiol 126:203–209
Franco-Zorrilla JM, Martin AC, Solano R, Rubio V, Leyva A, Paz-Ares J (2002) Mutations at CRE1 impair cytokinin-induced repression ofphosphate starvation responses in Arabidopsis. Plant J 32:353–360
Freed C, Adepoju O, Gillaspy G (2020) Can inositol pyrophosphates inform strategies for developing low phytate crops? Plants 9:115. https://doi.org/10.3390/plants9010115
Furkert D, Hostachy S, Nadler-Holly M, Fiedler D (2020) Triplexed affinity reagents to sample the mammalian inositol pyrophosphate interactome. Cell Chem Biol 27:1097–1108
Funkhouser EA, Loewus FA (1975) Purification of myo-Inositol 1-Phosphate Synthase from rice cell culture by affinity chromatography. Plant Physiol 56:786–90. https://doi.org/10.1104/pp.56.6.786
Gao Y-Q, Bu L-H, Han M-L, Wang Y-L, Li Z-Y, Liu H-T, Chao D-Y (2021) Long-distance blue light signalling regulates phosphate deficiency-induced primary root growth inhibition. Mol Plant 14(9):1539–1553. https://doi.org/10.1016/j.molp.2021.06.002
Gerasimaite R, Pavlovic I, Capolicchio S, Hofer A, Schmidt A, Jessen HJ, Mayer A (2017) Inositol pyrophosphate specificity of the SPX-dependent polyphosphate polymerase VTC. ACS Chem Biol 12:648–653
Giots F, Donaton MCV, Thevelein JM (2003) Inorganic phosphate is sensed by specific phosphate carriers and acts in concert with glucose as a nutrient signal for activation of the protein kinase A pathway in the yeast Saccharomyces cerevisiae. Mol Microbiol 47:1163–1181
Giri J, Bhosale R, Huang G, Pandey BK, Parker H, Zappala S, Yang J, Dievart A, Bureau C, Ljung K, Price A, Rose T, Larrieu A, Mairhofer S, Sturrock CJ, White P, Dupuy L, Hawkesford M, Perin C, Liang W, Peret B, Hodgman CT, Lynch J, Wissuwa M, Zhang D, Pridmore T, Mooney SJ, Guiderdoni E, Swarup R, Bennett MJ (2018) The rice auxin influx carrier OsAUX1 is required to facilitate root hair elongation in response to low external phosphate. Nat Commun 9(1):1408. https://doi.org/10.1038/s41467-018-03850-4
Godon C, Mercier C, Wang X, David P, Richaud P, Nussaume L, Liu D, Desnos T (2019) Under phosphate starvation conditions, Fe and Al trigger accumulation of the transcription factor STOP1 in the nucleus of Arabidopsis root cells. Plant J 99(5):937–949
Gokhale NA, Zaremba A, Shears SB (2011) Receptor-dependent compartmentalization of PPIP5K1, a kinase with a cryptic polyphosphoinositide binding domain. Biochem J 434:415–426
Gu C, Nguyen HN, Ganini D, Chen Z, Jessen HJ, Gu Z, Wang H, Shears SB (2017b) KO of 5-InsP7 kinase activity transforms the HCT116 colon cancer cell line into a hypermetabolic, growth-inhibited phenotype. Proc Natl Acad Sci U S A 2017(114):11968–11973
Gu C, Nguyen H-N, Hofer A, Jessen HJ, DaiX WH et al (2017a) The significance of the bifunctional kinase/phosphatase activities of PPIP5Ks for coupling inositol pyrophosphate cell-signaling to cellular phosphate homeostasis. J Biol Chem 292(11):4544–4555. https://doi.org/10.1074/jbc.M116.765743
Gu C, Wilson MSC, Jessen HJ, Saiardi A, Shears SB (2016) Inositol pyrophosphate profiling of two HCT116 cell lines uncovers variation in InsP8 levels. PLoS One 11(10):e0165286. https://doi.org/10.1371/journal.pone.0165286
Gutierrez-Alanis D, Yong-Villalobos L, Jimenez-Sandoval P, Alatorre-Cobos F, Oropeza-Aburto A, Mora-Macias J, Sanchez-Rodriguez F, Cruz-Ramirez A, Herrera-Estrella L (2017) Phosphate starvation-dependent iron mobilization induces CLE14 expression to trigger root meristem differentiation through CLV2/PEPR2 signaling. Dev Cell 41:555–570.e3
Ham BK, Lucas WJ (2017) Phloem-mobile RNAs as systemic signaling agents. Annu Rev Plant Biol 68:173–195
Hatch AJ, York JD (2010) SnapShot: inositol phosphates. Cell 143(6):1030–1030.e1. https://doi.org/10.1016/j.cell.2010.11.045
Holford ICR (1997) Soil phosphorus: its measurement, and its uptake by plants. Aust J Soil Res 35:227–239
Hothorn M, Neumann H, Lenherr ED, Wehner M, Rybin V, Hassa PO, Uttenweiler A, Reinhardt M, Schmidt A, Seiler J, Ladurner AG, Herrmann C, Scheffzek K, Mayer A (2009) Catalytic core of a membrane-associated eukaryotic polyphosphate polymerase. Science 324(5926):513–516
Huang KL, Ma GJ, Zhang ML, Xiong H, Wu H, Zhao CZ, Liu CS, Jia HX, Chen L, Ren F (2018) The ARF7 and ARF19 transcription factors positively regulate PHOSPHATE STARVATION RESPONSE1 in Arabidopsis roots. Plant Physiol 178:413–427
Huang S, Jeffery DA, Anthony MD, O’Shea EK (2001) Functional analysis of the cyclin-dependent kinase inhibitor Pho85p protein kinase, in the transduction pathway of pi signals in Saccharomyces cerevisiae. Mol Cell Biol 15:997–1004
Jia H, Zhang S, Wang L, Yang Y, Zhang H, Cui H, Shao H, Xu G (2017) OsPht1;8, a phosphate transporter, is involved in auxin and phosphate starvation response in rice. J Exp Bot 68(18):5057–5068. https://doi.org/10.1093/jxb/erx317
Jiang C, Gao X, Liao L, Harberd NP, Fu X (2007) Phosphate starvation root architecture and anthocyanin accumulation responses are modulated by the gibberellin–DELLA signaling pathway in Arabidopsis. Plant Physiol 145:1460–1470
Kaffman A, Herskowitz I, Tjian R, O’Shea EK (1994) Phosphorylation of the transcription factor PHO4 by a cyclin-CDK complex, PHO80-PHO85. Science 263:1153–1156
Kapulnik Y, Delaux PM, Resnick N, Mayzlish-Gati E, Wininger S, Bhattacharya C, Séjalon-Delmas N, Combier JP, Bécard G, Belausov E, Beeckman T, Dor E, Hershenhorn J, Koltai H (2011) Strigolactones affect lateral root formation and root-hair elongation in Arabidopsis. Planta 233(1):209–216. https://doi.org/10.1007/s00425-010-1310-y
Kim S-I, Tai TH (2011) Identification of genes necessary for wild-type levels of seed phytic acid in Arabidopsis thaliana using a reverse genetics approach. Mol Genet Genomics 286(2):119–133
Kochian LV, Hoekenga OA, Pinero MA (2004) How do crop plants tolerate acid soils? Mechanisms of aluminum tolerance and phosphorus efficiency. Annu Rev Plant Biol 55(1):459–493. https://doi.org/10.1146/annurev.arplant.55.031903.141655
Kuiper D, SteingrÖVer E (1991) Responses of growth, shoot to root ratio and Cytokinin concentrations in root tissue of two barley varieties, differing if their salt resistance. In: McMichael BL, Persson H (eds) Developments in agricultural and managed forest ecology. Elsevier, NY, USA, pp 463–471
Kuo HF, Chang TY, Chiang SF, Wang WD, Charng YY, Chiou TJ (2014) Arabidopsis inositol pentakisphosphate 2-kinase, AtIPK1, is required for growth and modulates phosphate homeostasis at the transcriptional level. Plant J 80:503–515
Kuo HF, Hsu YY, Lin WC, Chen KY, Munnik T, Brearley CA, Chiou TJ (2018) Arabidopsis inositol phosphate kinases IPK1 and ITPK1 constitute a metabolic pathway in maintaining phosphate homeostasis. Plant J 95:613–630
Laha D, Johnen P, Azevedo C, Dynowski M, Weiss M, Capolicchio S, Mao H, Iven T, Steenbergen M, Freyer M et al (2015) VIH2 regulates the synthesis of inositol pyrophosphate InsP8 and jasmonate-dependent defenses in Arabidopsis. Plant Cell 27:1082–1097
Laha D, Parvin N, Hofer A, Giehl RF, Fernandez-Rebollo N, von Wire’n N, Saiardi A, Jessen HJ, Schaaf G (2019) Arabidopsis ITPK1 and ITPK2 have an evolutionarily conserved phytic acid kinase activity. ACS Chem Biol 14:2127–2133
Lee B, Park SJ, Hong S, Kim K, Kim S (2021) Inositol polyphosphate multikinase signaling: multifaceted functions in health and disease. Mol Cells 44:187
Lee S, Kim MG, Ahn H, Kim S (2020) Inositol pyrophosphates: signaling molecules with pleiotropic actions in mammals. Molecules 25(9):2208. https://doi.org/10.3390/molecules25092208
Lee Y-S, Huang K, Quiocho FA, O’Shea EK (2008) Molecular basis of cyclin-CDK-CKI regulation by reversible binding of an inositol pyrophosphate. Nat Chem Biol 4(1):25–32. https://doi.org/10.1038/nchembio.2007.52
Lee Y-S, Mulugu S, York JD, O’Shea EK (2007) Regulation of a cyclin-CDK-CDK inhibitor complex by inositol pyrophosphates. Science 316:109–112. https://doi.org/10.1126/science.1139080
Lei M, Zhu C, Liu Y, Karthikeyan AS, Bressan RA, Raghothama KG et al (2011) Ethylene signalling is involved in regulation of phosphate starvation-induced gene expression and production of acid phosphatases and anthocyanin in Arabidopsis. New Phytol 189:1084–1095
Levy S, Kafri M, Carmi M, Barkai N (2011) The competitive advantage of a dual-transporter system. Science 334:1408–1412
Li X, Gu C, Hostachy S, Sahu S, Wittwer C, Jessen HJ, Shears SB (2020) Control of XPR1-dependent cellular phosphate efflux by InsP8 is an exemplar for functionally-exclusive inositol pyrophosphate signaling. Proc Natl Acad Sci 117(7):3568–3574
Liu D (2021) Root developmental responses to phosphorus nutrition. J Int Plant Biol 63(6):1065–1090. https://doi.org/10.1111/jipb.13090
Liu F, Wang Z, Ren H et al (2010) OsSPX1 suppresses the function of OsPHR2 in the regulation of expression of OsPT2 and phosphate homeostasis in shoots of rice. Plant J 62:508–517. https://doi.org/10.1111/j.1365-313X.2010.04170.x
Liu N, Shang W, Li C, Jia L, Wang X, Xing G, Zheng W (2018) Evolution of the SPX gene family in plants and its role in the response mechanism to phosphorus stress. Open Biol 8(1):170231
Liu Y, Xie Y, Wang H, Ma X, Yao W, Wang H (2017) Light and ethylene coordinately regulate the phosphate starvation response through transcriptional regulation of PHOSPHATE STARVATION RESPONSE1. Plant Cell 29(9):2269–2284. https://doi.org/10.1105/TPC.17.00268
Loewus FA, Murthy PP (2000) Myo-inositol metabolism in plants. Plant Sci 150(1):1–19. https://doi.org/10.1016/S0168-9452(99)00150-8
Lonetti A, Szijgyarto Z, Bosch D, Loss O, Azevedo C, Saiardi A (2011) Identification of an evolutionarily conserved family of inorganic polyphosphate endopolyphosphatases. J Biol Chem 286(37):31966–31974. https://doi.org/10.1074/jbc.M111.266320
López-Arredondo DL, Leyva-González MA, González-Morales SI, López-Bucio J, Herrera-Estrella L (2014) Phosphate nutrition: improving low-phosphate tolerance in crops. Annu Rev Plant Biol 65:95–123. https://doi.org/10.1146/annurev-arplant-050213-035949
Lopez-Bucio J, Hernandez-Abreu E, Sanchez-Calderon L, Nieto-Jacobo M, Simpson J, Herrera-Estrella L (2002) Phosphate availability alters architecture and causes changes in hormone sensitivity in the Arabidopsis root system. Plant Physiol 129:244–256
Lopez-Obando M, Ligerot Y, Bonhomme S, Boyer FD, Rameau C (2015) Strigolactone biosynthesis and signaling in plant development. Development 142:3615–3619
Lorenzo-Orts L, Couto D, Hothorn M (2020) Identity and functions of inorganic and inositol polyphosphates in plants. New Phytol 225:637–652. https://doi.org/10.1111/nph.16129
Lu M, Cheng Z, Zhang X-M et al (2020) Spatial divergence of PHR-PHT1 modules maintains phosphorus homeostasis in soybean nodules. Plant Physiol 184:236–250. https://doi.org/10.1104/pp.19.01209
Lv Q, Zhong Y, Wang Y, Wang Z, Zhang L, Shi J, Wu Z, Liu Y, Mao C, Yi K, Wu P (2014) SPX4 negatively regulates phosphate signaling and homeostasis through its interaction with PHR2 in Rice. Plant Cell 26(4):1586–1597
Ma Z, Baskin TI, Brown KM, Lynch JP (2003) Regulation of root elongation under phosphorus stress involves changes in ethylene responsiveness. Plant Physiol 131:1381–1390
Machkalyan G, Trieu P, Pétrin D, Hébert TE (2016) Miller GJ PPIP5K1 interacts with the exocyst complex through a C-terminal intrinsically disordered domain and regulates cell motility. Cell Signal 28:401–411
Magbanua JP, Fujisawa K, Ogawa N, Oshima Y (1997b) The homeodomain protein Pho2p binds at an A/T-rich segment flanking the binding site of the basic-helix-loop-helix protein Pho4p in the yeast PHO promoters. Yeast 13:1299–1308
Magbanua JP, Ogawa N, Harashima S, Oshima Y (1997a) The transcriptional activators of the PHO regulon, Pho4p and Pho2p, interact directly with each other and with components of the basal transcription machinery in Saccharomyces cerevisiae. J Biochem 121:1182–1189. https://doi.org/10.1093/oxfordjournals.jbchem.a021713
Mailer RK, Allende M, Heestermans M, Schweizer M, Deppermann C, Frye M, Pula G, Odeberg J, Gelderblom M, Rose-John S, Sickmann A, Blankenberg S, Huber TB, Kubisch C, Maas C, Gambaryan S, Firsov D, Stavrou EX, Butler LM, Renné T (2021) Xenotropic and polytropic retrovirus receptor 1 regulates procoagulant platelet polyphosphate. Blood 137(10):1392–1405
Marschner P (2012) Marschner’s mineral nutrition of higher plants, 3rd edn. Academic Press, Cambridge, p 649
Martin AC, del Pozo JC, Iglesias J, Rubio V, Solano R, de La Pena A et al (2000) Influence of cytokinins on the expression of phosphate starvation responsive genes in Arabidopsis. Plant J 24:559–567
Mora-Macías J, Ojeda-Rivera JO, Gutiérrez-Alanís D, Yong-Villalobos L, Oropeza-Aburto A, Raya-González J, Jiménez-Domínguez G, Chávez-Calvillo G, Rellán-Álvarez R, Herrera-Estrella L (2017) Malate-dependent Fe accumulation is a critical checkpoint in the root developmental response to low phosphate. Proc Natl Acad Sci 114:E3563–E3572
Morcuende R, Bari R, Gibon Y, Zheng W, Pant BD, Bläsing O, Usadel B, Czechowski T, Udvardi MK, Stitt M, Scheible WR (2007) Genome-wide reprogramming of metabolism and regulatory networks of Arabidopsis in response to phosphorus. Plant Cell Environ 30:85–112
Moritoh Y, Abe SI, Akiyama H, Kobayashi A, Koyama R, Hara R, Kasai S, Watanabe M (2021) The enzymatic activity of inositol hexakisphosphate kinase controls circulating phosphate in mammals. Nat Commun 12(1):4847
Mouillon J-M, Persson BL (2006) New aspects on phosphate sensing and signalling in Saccharomyces cerevisiae. FEMS Yeast Res 6:171–176
Müller J, Toev T, Heisters M, Teller J, Moore KL, Hause G, Dinesh DC, Bürstenbinder K, Abel S (2015) Iron-dependent Callose deposition adjusts root meristem maintenance to phosphate availability. Dev Cell 33:216–230
Nacry P, Canivenc G, Muller B et al (2005) A role for auxin redistribution in the responses of the root system architecture to phosphate starvation in Arabidopsis. Plant Physiol 138:2061–2074. https://doi.org/10.1104/pp.105.060061
Narise T, Kobayashi K, Baba S, Shimojima M, Masuda S, Fukaki H et al (2010) Involvement of auxin signaling mediated by IAA14 and ARF7/19 in membrane lipid remodeling during phosphate starvation. Plant Mol Biol 72:533–544
O’Neill EM, Kaffman A, Jolly ER, O’Shea EK (1996) Regulation of PHO4 nuclear localization by the PHO80–PHO85 cyclin-CDK complex. Science 271(5246):209–212. https://doi.org/10.1126/science.271.5246.209
Ogawa N, Noguchi K, Sawai H, Yamashita Y, Yompakudee C, Oshima Y (1995) Functional domains of Pho81p, an inhibitor of the Pho85p protein kinase, in the transduction pathway for pi signals in Saccharomyces cerevisiae. Mol Cell Biol 15:997–1004
Okushima Y, Overvoorde PJ, Arima K, Alonso JM, Chan A, Chang C, Ecker JR, Hughes B, Lui A, Nguyen D, Onodera C, Quach H, Smith A, Yu G, Theologis A (2005) Functional genomic analysis of the AUXIN RESPONSE FACTOR gene family members in Arabidopsis thaliana: unique and overlapping functions of ARF7 and ARF19. Plant Cell 17:444–463. https://doi.org/10.1105/TPC.104.028316
Osorio MB, Ng S, Berkowitz O et al (2019) SPX4 acts on PHR1-dependent and -independent regulation of shoot phosphorus status in Arabidopsis. Plant Physiol 181:332–352. https://doi.org/10.1104/pp.18.00594
Pandey BK, Verma L, Prusty A, Singh AP, Bennett MJ, Tyagi AK, Giri J, Mehra P (2021) OsJAZ11 regulates phosphate starvation responses in rice. Planta 254:8
Park BS, Seo JS, Chua NH (2014) Nitrogen limitation adaptation recruits phosphate2 to target the phosphate transporter pt2 for degradation during the regulation of Arabidopsis phosphate homeostasis. Plant Cell 26(1):454–464. https://doi.org/10.1105/tpc.113.120311
Park SJ, Lee S, Park SE, Kim S (2018) Inositol pyrophosphates as multifaceted metabolites in the regulation of mammalian signaling networks. Animal Cells Syst 22:1–6
Perez-Torres CA, Lopez-Bucio J, Cruz-Ramirez A, Ibarra-Laclette E, Dharmasiri S, Estelle M, Herrera-Estrella L (2008) Phosphate availability alters lateral root development in Arabidopsis by modulating auxin sensitivity via a mechanism involving the TIR1 auxin receptor. Plant Cell 20:3258–3272
Puga MI, Mateos I, Charukesi R, Wang Z, Franco-Zorrilla JM, de Lorenzo L, Irigoyen ML, Masiero S, Bustos W, Rodríguez J, Leyva A, Rubio V, Sommer H, Paz-Ares J (2014) SPX1 is a phosphate-dependent inhibitor of phosphate starvation response 1 in Arabidopsis. Proc Natl Acad Sci U S A 111(41):14947–14952
Raboy V (2001) Seeds for a better future: “low phytate” grains help to overcome malnutrition and reduce pollution. Trends Plant Sci 6:458–462
Ried MK, Wild R, Zhu J, Pipercevic J, Sturm K, Broger L et al (2021) Inositol pyrophosphates promote the interaction of SPX domains with the coiled-coil motif of PHR transcription factors to regulate plant phosphate homeostasis. Nat Commun 12:384
Riemer E, Qiu D, Laha D et al (2021) ITPK1 is an InsP6/ADP pohsphotransferase that controls phosphate signaling in Arabidopsis. Mol Plant 14:1864–1880
Rubio V, Linhares F, Solano R, Martin AC, Iglesias J, Leyva A et al (2001) A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev 15(16):2122–2133. https://doi.org/10.1101/gad.204401
Sabbagh Y (2013) Phosphate as a sensor and signaling molecule. Clin Nephrol 79:57–65. https://doi.org/10.5414/CN107322
Samyn DR, Van der Veken J, Van Zeebroeck G, Persson BL, Karlsson BC (2016) Key residues and phosphate release routes in the Saccharomyces cerevisiae Pho84 transceptor. J Biol Chem 291(51):26388–26398. https://doi.org/10.1074/jbc.M116.738112
Schneider KR, Smith RL, O’Shea EK (1994) Phosphate-regulated inactivation of the kinase PHO80–PHO85 by the CDK inhibitor PHO81. Science 266:122–126
Secco D, Wang C, Arpat BA, Wang Z, Poirier Y, Tyerman SD, Wu P, Shou H, Whelan J (2012) The emerging importance of the SPX domain-containing proteins in phosphate homeostasis. New Phytol 193(4):842–851. https://doi.org/10.1111/j.1469-8137.2011.04002.x
Secco D, Wang C, Shou H, Schultz MD, Chiarenza S, Nussaume L, Ecker JR, Whelan J, Lister R (2015) Stress induced gene expression drives transient DNA methylation changes at adjacent repetitive elements. Elife 4:e09343. https://doi.org/10.7554/eLife.09343
Secco D, Whelan J, Rouached H, Lister R (2017) Nutrient stress-induced chromatin changes in plants. Curr Opin Plant Biotech 39:1–7. https://doi.org/10.1016/j.pbi.2017.04.001
Seguel A, Cumming J, Klugh-Stewart K, Cornejo P, Borie P (2013) The role of arbuscular mycorrhizas in decreasing aluminium phytotoxicity in acidic soils: a review. Mycorrhiza. 23:167–183
Sengottaiyan P, Petrlova J, Lagerstedt JO, Ruiz-Pavon L, Budamagunta MS, Voss JC, Persson BL (2011) Characterization of the biochemical and biophysical properties of the Saccharomyces cerevisiae phosphate transporter Pho89. Biochem Biophys Res Commun 436:551–556. https://doi.org/10.1016/j.bbrc.2013.06.011
Shao D et al (1996) Interaction of Saccharomyces cerevisiae Pho2 with Pho4 increases the accessibility of the activation domain of Pho4. Mol Gen Genet 251(3):358–364
Shears SB (2018) Intimate connections: Inositol pyrophosphates at the interface of metabolic regulation and cell signaling. J Cell Physiol 233:1897–1912. https://doi.org/10.1002/jcp.26017
Shen C, Wang S, Zhang S, Xu Y, Qian Q, Qi Y et al (2013) OsARF16, a transcription factor, is required for auxin and phosphate starvation response in rice (Oryza sativaL.). Plant Cell Environ 36:607–620
Shi J, Hu H, Zhang K et al (2014) The paralogous SPX3 and SPX5 genes redundantly modulate pi homeostasis in rice. J Exp Bot 65:859–870. https://doi.org/10.1093/jxb/ert424
Shi J, Wang H, Hazebroek J, Ertl DS, Harp T (2005) The maize low-phytic acid 3 encodes a myo-inositol kinase that plays a role in phytic acid biosynthesis in developing seeds. Plant J 42(5):708–719
Silva-Navas J, Conesa CM, Saez A, Navarro-Neila S, Garcia-Mina JM, Zamarreño AM, Baigorri R, Swarup R, del Pozo JC (2019) Role of cis-zeatin in root responses to phosphate starvation. New Phytol 224(1):242–257. https://doi.org/10.1111/NPH.16020
Smith AP, Jain A, Deal RB, Nagarajan VK, Poling MD, Raghothama KG, Meagher RB (2010) Histone H2A.Z regulates the expression of several classes of phosphate starvation response genes but not as a transcriptional activator. Plant Physiol 152(1):217–225. https://doi.org/10.1104/pp.109.145532
Song L, Yu H, Dong J, Che X, Jiao Y, Liu D (2016) The molecular mechanism of ethylene-mediated root hair development induced by phosphate starvation. PLoS Genet 12(7):e1006194. https://doi.org/10.1371/journal.pgen.1006194
Sorefan K, Booker J, Haurogne K, Goussot M, Bainbridge K, Foo E et al (2003) MAX4 and RMS1 are orthologous dioxygenase-like genesthat regulate shoot branching in Arabidopsis and pea. Genes Dev 17:1469–1474
Stevenson-Paulik J, Odom AR, York JD (2002) Molecular and biochemical characterization of two plant inositol polyphosphate 6−/3−/5-kinases. J Biol Chem 277:42711–42718
Svistoonoff S, Creff A, Reymond M, Sigoillot-Claude C, Ricaud L, Blanchet A, Nussaume L, Desnos T (2007) Root tip contact with low-phosphate media reprograms plant root architecture. Nat Genet 39(6):792–796. https://doi.org/10.1038/ng2041
Thibaud MC, Arrighi JF, Bayle V, Chiarenza S, Creff A, Bustos R et al (2010) Dissection of local and systemic transcriptional responses to phosphate starvation in Arabidopsis. Plant J 64:775–789
Tian WH, Ye JY, Cui MQ, Chang JB, Liu Y, Li GX, Wu YR, Xu JM, Harberd NP, Mao CZ, Jin CW, Ding ZJ, Zheng SJ (2021) A transcription factor STOP1-centered pathway coordinates ammonium and phosphate acquisition in Arabidopsis. Mol Plant 14(9):1554–1568. https://doi.org/10.1016/j.molp.2021.06.024
Ticconi CA, Lucero RD, Sakhonwasee S, Adamson AW, Creff A, Nussaume L, Desnos T, Abel S (2009) ER-resident proteins PDR2 and LPR1 mediate the developmental response of root meristems to phosphate availability. Proc Natl Acad Sci U S A 106:14174–14179
Toh-e A, Tanaka K, Uesono Y, Wickner RB (1988) PHO85, a negative regulator of the PHO system, is a homolog of the protein kinase gene, CDC28, of Saccharomyces cerevisiae. Mol Gen Genet 214:162–164
Umehara M, Hanada A, Yoshida S, Akiyama K, Arite T, Takeda-Kamiya N et al (2008) Inhibition of shoot branching by new terpenoid plant hormones. Nature 455:195–200
Wang C, Yue W, Ying Y, Wang S, Secco D, Liu Y, Whelan J, Tyerman SD, Shou H (2015) Rice SPX-major facility Superfamily3, a vacuolar phosphate efflux transporter, is involved in maintaining phosphate homeostasis in Rice. Plant Physiol 169(4):2822–2831
Wang H, Derose EF, London RE, Shears SB (2014c) IP6K structure and the molecular determinants of catalytic specificity in an inositol phosphate kinase family. Nat Commun 5:4178
Wang H, Falck JR, Hall TMT, Shears SB (2012) Structural basis for an inositol pyrophosphate kinase surmounting phosphate crowding. Nat Chem Biol 8:111–116
Wang J, Sun J, Miao J et al (2013b) A phosphate starvation response regulator ta-PHR1 is involved in phosphate signalling and increases grain yield in wheat. Ann Bot 111:1139–1153. https://doi.org/10.1093/aob/mct080
Wang S, Ichii M, Taketa S, Xu L, Xia K, Zhou X (2002) Lateral root formation in rice (Oryza sativa): promotion effect of jasmonic acid. J Plant Physiol 159(8):827–832. https://doi.org/10.1078/0176-1617-00825
Wang S, Zhang S, Sun C, Xu Y, Chen Y, Yu C, Qian Q, Jiang DA, Qi YH (2014a) Auxin response factor (OsARF12), a novel regulator for phosphate homeostasis in rice (Oryza sativa). New Phytol 201(1):91–103. https://doi.org/10.1111/nph.12499
Wang X, Bai J, Liu H, Sun Y, Shi X, Ren Z (2013a) Overexpression of a maize transcription factor ZmPHR1 improves shoot inorganic phosphate content and growth of Arabidopsis under low-phosphate conditions. Plant Mol Biol Rep 31(3):665–677. https://doi.org/10.1007/s11105-012-0534-3
Wang X, Yi K, Tao Y, Wang F, Wu Z, Jiang D et al (2006) Cytokinin represses phosphate-starvation response through increasing of intracellular phosphate level. Plant Cell Environ 29(10):1924–1935. https://doi.org/10.1111/j.1365-3040.2006.01568.x
Wang Z, Hu H, Huang H, Duan K, Wu Z, Wu P (2009) Regulation of OsSPX1 and OsSPX3 on expression of OsSPX domain genes and pi-starvation signaling in rice. J Integr Plant Biol 51:663–674
Wang Z, Kuo HF, Chiou TJ (2021) Intracellular phosphate sensing and regulation of phosphate transport systems in plants. Plant Physiol 187(4):2043–2055
Wang Z, Ruan W, Shi J, Zhang L, Xiang D, Yang C et al (2014b) RiceSPX1 and SPX2 inhibit phosphate starvation responses through inter-acting with PHR2 in a phosphate-dependent manner. Proc Natl Acad Sci U S A 111:14953–14958
Wild R, Gerasimaite R, Jung JY, Truffault V, Pavlovic I, Schmidt A, Saiardi A, Jessen HJ, Poirier Y, Hothorn M, Mayer A (2016) Control of eukaryotic phosphate homeostasis by inositol polyphosphate sensor domains. Science 352:986–990
Wu P, Shou H, Xu G, Lian X (2013) Improvement of phosphorus efficiency in rice on the basis of understanding phosphate signaling and homeostasis. Curr Opin Plant Biol 16:205–212. https://doi.org/10.1016/j.pbi.2013.03.002
Wykoff DD, Grossman AR, Weeks DP et al (1999) Psr1, a nuclear localized protein that regulates phosphorus metabolism in Chlamydomonas. PNAS 96:15336–15341. https://doi.org/10.1073/pnas.96.26.15336
Wykoff DD, Rizvi AH, Raser JM, Margolin B, O'Shea EK (2007) Positive feedback regulates switching of phosphate transporters in S. cerevisiae. Mol Cell 27:1005–1013
Xu JM, Wang ZQ, Wang JY, Li PF, Jin JF, Chen WW, Fan W, Kochian LV, Zheng SJ, Yang JL (2020) Low phosphate represses histone deacetylase complex1 to regulate root system architecture remodeling in Arabidopsis. New Phytol 225:1732–1745
Xu L, Jin L, Long L, Liu L, He X, Gao W, Zhu L, Zhang X (2012) Overexpression ofGbWRKY1 positively regulates the pi starvation response by alterationof auxin sensitivity in Arabidopsis. Plant Cell Rep 31(12):2177–2188. https://doi.org/10.1007/s00299-012-1328-7
Yoneyama K, Kisugi T, Xie X, Yoneyama K (2013) Chemistry of strigolactones: why and how do plants produce so many strigolactones? In: de Bruijn FJ (ed) Molecular microbial ecology of the rhizosphere. John Wiley & Sons, Inc., Hoboken, NJ
Yong-Villalobos L, Gonzalez-Morales SI, Wrobel K, Gutierrez-Alanis D, Cervantes-Perez SA, Hayano-Kanashiro C, Oropeza-Aburto A, Cruz-Ramirez A, Martinez O, Herrera-Estrella L (2015) Methylome analysis reveals an important role for epigenetic changes in the regulation of the Arabidopsis response to phosphate starvation. Proc Natl Acad Sci U S A 112(52):E7293–EE730. https://doi.org/10.1073/pnas.1522301112
Zhang K, Zhou Z, Li J, Wang J, Yu L, Lin S (2021) SPX-related genes regulate phosphorus homeostasis in the marine phytoplankton, Phaeodactylum tricornutum. Commun Biol 4(1):797
Zhang YJ, Lynch JP, Brown KM (2003) Ethylene and phosphorus availability have interacting yet distinct effects on root hair development. J Exp Bot 54:2351–2361
Zhang ZL, Zheng Y, Ham BK, Chen JY, Yoshida A, Kochian LV, Fei ZJ, Lucas WJ (2016) Vascular-mediated signalling involved in early phosphate stress response in plants. Nat Plants 2(4):16033. https://doi.org/10.1038/nplants.2016.33
Zheng Z, Wang Z, Wang X, Liu D (2019) Blue light-triggered chemical reactions underlie phosphate deficiency-induced inhibition of root elongation of Arabidopsis seedlings grown in petri dishes. Mol Plant 12(11):1515–1523. https://doi.org/10.1016/j.molp.2019.08.001
Zhou J, Hu Q, Xiao X, Yao D, Ge S, Ye J, Li H, Cai R et al (2021) Mechanism of phosphate sensing and signaling revealed by rice SPX1-PHR2 complex structure. Nat Commun 12(1):7040. https://doi.org/10.1038/s41467-021-27391-5
Zhou J, Jiao FC, Wu Z et al (2008) OsPHR2 is involved in phosphate-starvation signaling and excessive phosphate accumulation in shoots of plants. Plant Physiol 146:1673–1686. https://doi.org/10.1104/pp.107.111443
Zhou X, O’Shea EK (2011) Integrated approaches reveal determinants of genome-wide binding and function of the transcription factor Pho4. Mol Cell 42(6):826–836
Zhu J, Lau K, Puschmann R, Harmel RK, Zhang Y, Pries V, Gaugler P, Broger L, Dutta AK, Jessen HJ et al (2019) Two bifunctional inositol pyrophosphate kinases/phosphatases control plant phosphate homeostasis. ELife 8:1–25
Zhuang Q, Xue Y, Yao Z et al (2021) Phosphate starvation responsive GmSPX5 mediates nodule growth through interaction with GmNF-YC4 in soybean (Glycine max). Plant J 108:1422–1438. https://doi.org/10.1111/tpj.15520
Acknowledgements
Not applicable.
Funding
This work was supported by the Chinese Academy of Sciences.
Author information
Authors and Affiliations
Contributions
ML, VS: conceived the review; AT: drafted the section on the comparison between plant and human InsP8 signaling; JL: drafted the section on the InsP-SPX-PHR interaction; ML, VS: Wrote, revised and finalized the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Satheesh, V., Tahir, A., Li, J. et al. Plant phosphate nutrition: sensing the stress. Stress Biology 2, 16 (2022). https://doi.org/10.1007/s44154-022-00039-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44154-022-00039-0