Abstract
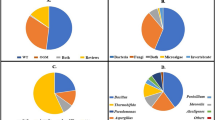
Oil pollution is a major global environmental concern. Bioremediation is considered as a suitable approach for remediation of oil-contaminated environments. In this study, two crude oil-contaminated soils were collected to isolate and identify some native and active oil-degrading bacteria to be used in remediation of contaminated sites. Five isolates were selected according to “hole-plate diffusion method” and were grown in crude oil. They were cultured in a mineral salt medium in which crude oil was employed as the sole carbon source. Biochemical, morphological and genomic identifications demonstrated the bacteria species as Pseudomonas resinovorans, Plantibacter auratus, Bacillus subtilis, Staphylococcus pasteuri and Bacillus atrophaeus. These bacteria were able to degrade 86.0%, 61.3%, 81.1%, 55.0% and 76.2% of aliphatic compounds and 58.6%, 39.4%, 55.5%, 39.0% and 49.9% of aromatic hydrocarbons in a medium containing crude oil (1% v/v) over 21 days, respectively. The degradation rates of aromatic compounds from 14 to 21 days were higher than of aliphatic hydrocarbons. This rate was 28.4% by Bacillus subtilis, 30.9% by Bacillus atrophaeus, 27.2% by Staphylococcus pasteuri and 21.3% by Pseudomonas resinovorans. In Plantibacter auratus, this rate was 16.19% which is less than aliphatic hydrocarbons. To our knowledge, it seems this is the first time to report Pseudomonas resinovorans and Plantibacter auratus as crude oil degraders. Results of this study indicated that the isolated bacteria could have a high potential to be used in bioremediation of oil-contaminated environments.






Similar content being viewed by others
References
Alharbi OML, Basheer AA, Khattab RA, Ali I (2018) Health and environment effects of persistent organic pollutants. J Mol Liq 263:442–453
Ali I, Aboul-Enein HY, Gupta VK (2009) Nano chromatography and capillary electrophoresis: pharmaceutical and environmental analyses. Wiley & Sons, Hoboken, USA
Ali I, Asim M, Khan TA (2013) Arsenite removal from water by electro-coagulation on zinc–zinc and copper–copper electrodes. Int J Environ Sci Technol 10:377–384
Ali I, Alharbi OML, Alothman ZA, Badjah AY, Alwarthan A, Basheer AA (2018) Artificial neural network modelling of amido black dye sorption on iron composite nano material: kinetics and thermodynamics studies. J Mol Liq 250:1–8. https://doi.org/10.1016/j.molliq.2017.11.163
Al-Sayegh A, Al-Wahaibi Y, Joshi S, Al-Bahry S, Elshafie A, Al-Bemani A (2017) Draft genome sequence of Bacillus subtilis AS2, a heavy crude oil-degrading and biosurfactant-producing bacterium isolated from a soil sample. Genome Announc 5:e00969-17. https://doi.org/10.1128/genomeA.00969-17
Atlas RM, Bartha R (1992) Hydrocarbon biodegradation and oil-spill bioremediation. In: Marshall KC (ed) Advances in microbial ecology, vol 12. Plenum Press, New York, pp 287–338
Azubuike CC, Chikere CB, Okpokwasili GC (2016) Bioremediation techniques–classification based on site of application: principles, advantages, limitations and prospects. World J Microbiol Biotechnol 32(11):180. https://doi.org/10.1007/s11274-016-2137-x
Basheer AA (2018) Chemical chiral pollution: impact on the society and science and need of the regulations in the 21st century. Chirality 30(4):402–406. https://doi.org/10.1002/chir.22808
Beilen V, Funhoff EG (2007) Alkane hydroxylases involved in microbial alkane degradation. Appl Microbiol Biotechnol 74:13–21. https://doi.org/10.1007/s00253-006-0748-0
Burakova E, Dyachkova T, Rukhov A, Rukhov A, Tugolukov E, Galunin E, Tkachev A, Basheer AA, Ali I (2018) Novel and economic method of carbon nanotubes synthesis on a nickel magnesium oxide catalyst using microwave radiation. J Mol Liq 253:340–346. https://doi.org/10.1016/j.molliq.2018.01.062
Cai Q, Zhang B, Bing C, Zhu Z, Lin W, Tong C (2014) Screening of biosurfactant producers from petroleum hydrocarbon contaminated sources in cold marine environments. Mar Poll Bull 86:402–410. https://doi.org/10.1016/j.marpolbul.2014.06.039
Chauhan A, Green S, Pathak A, Thomas J, Venkatramanan R (2013) Whole-genome sequences of five oyster-associated bacteria show potential for crude oil hydrocarbon degradation. Genome Announc 1:e00802-13
Chen Q, Bao M, Fan X, Liang S, Sun P (2013) Rhamnolipids enhance marine oil spill bioremediation in laboratory system. Mar Poll Bull 71:269–275. https://doi.org/10.1016/j.marpolbul.2013.01.037
Chen Q, Li J, Liu M, Sun H, Bao M (2017) Study on the biodegradation of crude oil by free and immobilized bacterial consortium in marine environment. PLoS ONE 12:e0174445. https://doi.org/10.1128/genomeA.00802-13
Das N, Chandran P (2011) Microbial degradation of petroleum hydrocarbon contaminants: an overview. Biotechnol Res Int 2011:1–13. https://doi.org/10.4061/2011/941810
Das M, Das SK, Mukherjee RK (1998) Surface active properties of the culture filtrates of a Micrococcus species grown on n-alkanes and sugars. Bioresour Technol 63:231–235. https://doi.org/10.1016/S0960-8524(97)00133-8
Deschênes L, Lafrance P, Villeneuve JP, Samson R (1996) Adding sodium dodecyl sulphate and Pseudomonas aeruginosa UG2 biosurfactants inhibits polycyclic aromatic hydrocarbon biodegradation in a weathered creosotecontaminated soil. Appl Microbiol Biotechnol 46:638–646. https://doi.org/10.1007/s002530050874
Doley R, Barthakur M (2017) Biodegradation of Naphthalene by Staphylococcus pasteuri RD2 Isolated from oil contaminated soil. Int J Curr Microbiol Appl Sci 6:1310–1319. https://doi.org/10.20546/ijcmas.2017.612.148
Gentili A, Cubittoa MA, Ferrerob M, Rodriguéz MS (2006) Bioremediation of crude oil polluted seawater by a hydrocarbon degrading bacterial strain immobilized on chitin and chitosan flakes. Int Biodeterior Biodegrad 57:222–228. https://doi.org/10.1016/j.ibiod.2006.02.009
Gupta VK, Ali I (2012) Environmental water: advances in treatment, remediation and recycling. Elsevier, Netherlands
Haritash AK, Kaushik CP (2009) Biodegradation aspects of polycyclic aromatic hydrocarbons (PAHs): a review. J Hazard Mater 169:1–15. https://doi.org/10.1016/j.jhazmat.2009.03.137
Head IM, Jones DM, Röling WFM (2006) Marine microorganisms make a meal of oil. Nat Rev Microbiol 4:173–182. https://doi.org/10.1038/nrmicro1348
Ismail W, Alhamad NA, El-Sayed WS, El Nayal AM, Chiang Y, Hamzah RY (2013) Bacterial degradation of the saturate fraction of Arabian light crude oil: biosurfactant production and the effect of ZnO nanoparticles. J Pet Environ Biotechnol 4:1–11. https://doi.org/10.4172/2157-7463.1000163
Kanaly RA, Harayama S (2000) Biodegradation of high-molecular-weight polycyclic aromatic hydrocarbons by bacteria. J Bacteriol 182:2059–2067. https://doi.org/10.1128/JB.182.8.2059-2067
Kauppi S, Sinkkonen A, Romantschuk M (2011) Enhancing bioremediation of diesel-fuel-contaminated soil in a boreal climate: comparison of biostimulation and bioaugmentation. Int Biodeterior Biodegrad 65:359–368. https://doi.org/10.1016/j.ibiod.2010.10.011
Kumar AG, Vijayakumar L, Joshi G, Peter DM, Dharani G, Kirubagaran R (2014) Biodegradation of complex hydrocarbons in spent engine oil by novel bacterial consortium isolated from deep sea sediment. Bioresour Technol 170:556–564. https://doi.org/10.1016/j.biortech.2014.08.008
Lin TC, Shen FT, Chang JS, Young CC, Arun AB, Lin SY, Chen TL (2009) Hydrocarbon degrading potential of bacteria isolated from oil-contaminated soil. J Taiwan Inst Chem Eng 40:580–582. https://doi.org/10.1016/j.jtice.2009.03.001
Luqueño FF, Encinas CV, Marsch R, Martínez-Suárez C, Vázquez-Núñez E, Dendooven L (2011) Microbial communities to mitigate contamination of PAHs in soil possibilities and challenges: a review. Environ Sci Pollut Res Int 18:12–30. https://doi.org/10.1007/s11356-010-0371-6
Mojarad M, Alemzadeh A, Golafarin G, Javaheri M (2016) Kerosene biodegradation ability and characterization of bacteria isolated from oil-polluted soil and water. J Environ Chem Eng 4:4323–4329
Pasumarthi R, Chandrasekaran S, Mutnuri S (2013) Biodegradation of crude oil by Pseudomonas aeruginosa, and Escherichia fergusonii, isolated from the Goan Coast. Marine Poll Bull 76:276–282. https://doi.org/10.1016/j.marpolbul.2013.08.026
Perelo LW (2010) Review: in situ and bioremediation of organic pollutants in aquatic sediments. J Hazard Mater 177:81–89. https://doi.org/10.1016/j.jhazmat.2009.12.090
Saadoun I (2002) Isolation and characterization of bacteria from crude petroleum oil contaminated soil and their potential to degrade diesel fuel. J Basic Microbiol 42:420–428. https://doi.org/10.1002/1521-4028
Schippers A, Bosecker K, Spröer C, Schumann P (2005) Microbacterium oleivorans sp. nov. and Microbacterium hydrocarbonoxydans sp. nov. novel crude-oil-degrading gram-positive bacteria. Int J Syst Evol Microbiol 55:655–660. https://doi.org/10.1099/ijs.0.63305-0
Soleimani M, Afyuni M, Hajabbasi MA, Nourbakhsh F, Sabzalian MR, Christensen JH (2010) Phytoremediation of an aged petroleum contaminated soil using endophyte infected and non-infected grasses. Chemosphere 81:1084–1090. https://doi.org/10.1016/j.chemosphere.2010.09.034
Suja F, Rahim F, Taha MR, Nuraini H, Razali MR, Khalid A, Hamzah A (2014) Effects of local microbial bioaugmentation and biostimulation on the bioremediation of total petroleum hydrocarbons (TPH) in crude oil contaminated soil based on laboratory and field observations. Int Biodeterior Biodegrad 90:115–122. https://doi.org/10.1016/j.ibiod.2014.03.006
Wang H, Wang C, Lin M, Sun X, Wang C, Hu X (2013) Phylogenic diversity of bacterial communities associated with bioremediation of crude oil in microcosms. Int Biodeterior Biodegrad 85:400–406. https://doi.org/10.1016/j.ibiod.2013.07.015
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703. https://doi.org/10.1128/jb.173.2.697-703
Zhang X, Xu D, Zhu C, Lundaa T, Scherr KE (2012) Isolation and identification of biosurfactant producing and crude oil degrading pseudomonas aeruginosa strains. Chem Eng J 209:138–146. https://doi.org/10.1016/j.cej.2012.07.110
Zhao D, Liu C, Liu L, Zhang Y, Liu Q, Wu WM (2011) Selection of functional consortium for crude oil-contaminated soil remediation. Int Biodeterior Biodegrad 65:1244–1248. https://doi.org/10.1016/j.ibiod.2011.07.008
Acknowledgements
The authors appreciate Mrs. Hélène Dailly at the Catholic University of Louvain for her helpful comments on analyzing samples by GC–MS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors confirm that there are no known conflicts of interest associated with this publication.
Additional information
Editorial responsibility: M. Abbaspour.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kiamarsi, Z., Soleimani, M., Nezami, A. et al. Biodegradation of n-alkanes and polycyclic aromatic hydrocarbons using novel indigenous bacteria isolated from contaminated soils. Int. J. Environ. Sci. Technol. 16, 6805–6816 (2019). https://doi.org/10.1007/s13762-018-2087-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13762-018-2087-y