Abstract
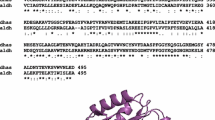
3-Hydroxypropionic acid (3-HP), a versatile and valuable platform chemical, has diverse industrial applications; but its biological production from glycerol is often limited by the capability of the enzyme aldehyde dehydrogenase (ALDH) to convert an intermediary compound, 3-hydroxypropionaldehyde (3-HPA), to 3-HP. In this study, we report a new ALDH, PuuC, from Klebsiella pneumoniae DSM 2026, that efficiently converts 3-HPA to 3-HP. The identified gene puuC was cloned, expressed in Escherichia coli, purified, and characterized for its properties. The recombinant enzyme with a molecular weight of 53.8 kDa exhibited broad substrate specificity for various aliphatic aldehydes, especially C2–C5 aldehydes. NAD+ was the preferred coenzyme for the oxidation of most aliphatic and aromatic aldehydes tested. The optimum pH and temperature for PuuC activity were pH 8.0 and 45°C. The K m values for 3-HPA and NAD+ were 0.48 and 0.09 mM, respectively. The activity of PuuC was enhanced in the presence of reducing agents such as 2-mercaptoethanol or dithiothreitol, while several metal ions, particularly Hg2+, Ag+, and Cu2+ inhibited its activity. The predicted structure of PuuC indicated the presence of K191 and E194 in close proximity to the glycine motif, suggesting that PuuC belongs to class 2 ALDHs.
Similar content being viewed by others
References
Jo, J. E., S. Mohanraj, C. Rathnasingh, E. Selvakumar, W. C. Jung, and S. H. Park (2008) Cloning, expression, and characterization of an aldehyde dehydrogenase from Escherichia coli K-12 that utilizes 3-Hydroxypropionaldehyde as a substrate. Appl. Microbiol. Biotechnol. 81: 51–60.
Raj, S. M., C. Rathnasingh, W. C. Jung, and S. H. Park (2009) Effect of process parameters on 3-hydroxypropionic acid production from glycerol using a recombinant Escherichia coli. Appl. Microbiol. Biotechnol. 84: 649–657.
Rathnasingh, C., S. M. Raj, J. E. Jo, and S. H. Park (2009) Development and evaluation of efficient recombinant Escherichia coli strains for the production of 3-hydroxypropionic acid from glycerol. Biotechnol. Bioeng. 104: 729–739.
Toraya, T., N. Tamura, T. Watanabe, M. Yamanishi, N. Hieda, and K. Mori (2008) Mechanism-based inactivation of coenzyme B12-dependant diol dehydratase by 3-unsaturated 1,2-diols and thioglycerol. J. Biochem. 144: 437–446.
Watanabe, S., M. Yamada, I. Ohtsu, and K. Makino (2007) α-ketoglutaric semialdehyde dehydrogenase isozymes involved in metabolic pathways of D-glucarate, D-galactarate, and hydroxyl-L-proline. J. Biol. Chem. 282: 6685–6695.
Hall, R. H. and E. S. Stern (1950) Acid-catalyzed hydration of acrylaldehyde: kinetics of the reaction and isolation of β-hydroxypropionaldehyde. J. Chem. Soc. 1950: 490–498.
Tamura K., J. Dudley, M. Nei, and S. Kumar (2007) MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24: 1596–1599.
Sambrook, J. and D. Russell (2001) Molecular cloning: A Laboratory Manual. 3rd ed., Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, USA.
Laemmli, U. K (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 227: 680–685.
Laskowski, R. A., M. W. MacArthur, D. S. Moss, and J. M. Thornton (1993) PROCHECK: A program to check the stereochemical quality of protein structures. J. Appl. Cryst. 26: 283–291.
Luthy, R., J. U. Bowie, and D. Eisenberg (1992) Assessment of protein models with three-dimensional profiles. Nature 356: 83–85.
Bradford, M. M. (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72: 248–254.
Hempel, J., H. Nicholas, and R. Lindahl (1993) Aldehyde dehydrogenases: widespread structural and functional diversity within a shared framework. Protein Sci. 2: 1890–1900.
Perozich, J., H. Nicholas, B. C. Wang, R. Lindahl, and J. Hempel (1999) Relationships within the aldehyde dehydrogenase extended family. Protein Sci. 8: 137–146.
Ho, K. K. and H. Weiner (2005) Isolation and characterization of an aldehyde dehydrogenase encoded by the aldB gene of Escherichia coli. J. Bacteriol. 187: 1067–1073.
Tigerstrom, R. G. V. and W. E. Razzell (1968) Aldehyde dehydrogenase. J. Biol. Chem. 243: 2691–2702.
Kurihara, S., S. Oda, K. Kato, H. G. Kim, T. Koyanagi, H. Kumagai, and H. Suzuki (2005) A novel putrescine utilization pathway involves γ-glutamylated intermediates of Escherichia coli K-12. J. Biol. Chem. 280: 4602–4608.
Steinman, C. R. and W. B. Jakoby (1967) Yeast aldehyde dehydrogenase. J. Biol. Chem. 242: 5019–5023.
Ohta, T., A. Tani, K. Kimbara, and F. Kawai (2005) A novel nictinoprotein aldehyde dehydrogenase involved in polyethylene glycol dehydration. Appl. Microbiol. Biotechnol. 68: 639–646.
Kim, H. G., Y. Kim, H. M. Kim, H. J. Shin, and S. W. Kim (2006) Purification, characterization, and cloning of trimethylamine dehydrogenase from Methylophaga sp. Strain SK1. Biotechnol. Bioprocess. Eng. 11: 337–343.
Jaureguibeitia, A., L. Saa, M. J. Llama, and J. L. Serra (2007) Purification, characterization and cloning of aldehyde dehydrogenase from Rhodococcus erythropolis UPV-1. Appl. Microbiol. Biotechnol. 73: 1073–1086.
Perozich, J., I. Kuo, R. Lindahl, and J. Hempel (2001) Coenzyme specificity in aldehyde dehydrogenase. Chem. Biol. Interact. 130–132: 115–124.
Author information
Authors and Affiliations
Corresponding author
Additional information
Both the authors contributed equally to this work.
Rights and permissions
About this article
Cite this article
Raj, S.M., Rathnasingh, C., Jung, WC. et al. A Novel NAD+-dependent aldehyde dehydrogenase encoded by the puuC gene of Klebsiella pneumoniae DSM 2026 that utilizes 3-hydroxypropionaldehyde as a substrate. Biotechnol Bioproc E 15, 131–138 (2010). https://doi.org/10.1007/s12257-010-0030-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12257-010-0030-2