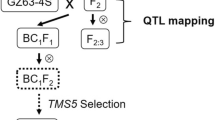
The study of 1000-grain weight (TGW) and percentage of grains with chalkiness (PGWC) is very important in rice. In this study, a set of introgression lines (ILs), derived from Sasanishiki/Habataki with Sasanishiki as the recurrent parent, were used to detect correlations and quantitative trait loci (QTL) on TGW and PGWC in two different environments. Phenotypic correlation analysis showed that there was no significant correlation between TGW and PGWC in both environments, which indicated that the linkage of TGW and PGWC traits could be broken via suitable population. A total of 20 QTL were detected in both environments, nine QTL for 1000-paddy-grain weight (PTGW), five QTL for 1000-brown-grain weight (BTGW) and six QTL for percentage of grains with chalkiness (PGWC). Moreover, five QTL, qPTGW3, qPTGW8.2, qPTGW11.1 for PTGW and qPGWC1.1, qPGWC1.2 for PGWC, were stably expressed in both environments. Phenotypic values were significantly different (P < 0.01) between the introgression lines carrying these five QTL alleles and the genetic background parent, Sasanishiki. The introgression lines carrying these QTL also represent a useful genetic resource in the context of rice yield and quality improvement via a design-breeding approach.



Similar content being viewed by others
References
Ando T., Yamamoto T., Shimizu T., Ma X. F., Shomura A., Takeuchi Y. et al. 2008 Genetic dissection and pyramiding of quantitative traits for panicle architecture by using chromosomal segment substitution lines in rice. Theor. Appl. Genet. 116, 881–890.
Bai X., Luo L., Yan W., Kovi M. R., Zhan W. and Xing Y. 2010 Genetic dissection of rice grain shape using a recombinant inbred line population derived from two contrasting parents and fine mapping a pleiotropic quantitative trait locus qGL7. BMC Genet. 11, 1471–2156.
Bai X. F., Luo L. J., Yan W. H., Kovi M. R. and Xing Y. Z. 2011 Quantitative trait loci for rice yield-related traits using recombinant inbred lines derived from two diverse cultivars. J. Genet. 90, 209–215.
Bian J., Jiang L., Liu L. L., Wei X. J., Xiao Y. H., Zhang L. J. et al. 2010 Construction of a new set of rice chromosome segment substitution lines and identification of grain weight and related traits QTLs. Breed. Sci. 60, 305–313.
Bian J., He H., Shi H., Zhu C., Peng X., Li C. et al. 2013 Dynamic QTL detection and analysis of tiller number before and after heading in ‘japonica’ rice. Aust. J. Crop Sci. 7, 1189–1197.
Cheng F. M., Zhong L. J., Wang F. and Zhang G. P. 2005 Differences in cooking and eating properties between chalky and translucent parts in rice grains. Food Chem. 90, 39–46.
Fujita N., Yoshida M., Kondo T., Saito K., Utsumi Y., Tokunaga T. et al. 2007 Characterization of SSIIIa-deficient mutants of rice: The function of SSIIIa and pleiotropic effects by SSIIIa deficiency in the rice endosperm. Plant Physiol. 144, 2009–2023.
Guo T., Liu X., Wan X., Weng J., Liu S., Liu X. et al. 2011 Identification of a stable quantitative trait locus for percentage grains with white chalkiness in rice (Oryza sativa). J. Integr. Plant Biol. 53, 598–607.
Guo L., Ma L., Jiang H., Zeng D., Hu J., Wu L. et al. 2009 Genetic analysis and fine mapping of two genes for grain shape and weight in rice. J. Integr. Plant Biol. 51, 45–51.
Hao W., Zhu M. Z., Gao J. P., Sun S. Y. and Lin H. X. 2009 Identification of quantitative trait loci for rice quality in a population of chromosome segment substitution lines. J. Integr. Plant Biol. 51, 500–512.
He P., Li S. G., Qian Q., Ma Y. Q., Li J. Z., Wang W. M. et al. 1999 Genetic analysis of rice grain quality. Theor. Appl. Genet. 98, 502–508.
Juliano B. O., Perez C. M. and Kaosa A. M. 1990 Grain quality characteristics of export rice in selected markets. Cereal Chem. 67, 192–197.
Kang H. G., Park S. H., Matsuoka M. and An G. H. 2005 White-core endosperm floury endosperm-4 in rice is generated by knockout mutations in the C4-type pyruvate orthophosphate dikinase gene (OsPPDKB). Plant J. 42, 901–911.
Kubo T., Aida Y., Nakamura K., Tsunematsu H., Doi K. and Yoshimura A. 2002 Reciprocal chromosome segment substitution series derived from japonica and indica cross of rice (Oryza sativa L.). Breed. Sci. 52, 319–325.
Li H. H., Ye G. Y. and Wang J. K. 2007 A modified algorithm for the improvement of composite interval mapping. Genetics 175, 361–374.
Li H., Li J. M., Ribaut Z. and Wang J. 2008 Inclusive composite interval mapping (ICIM) for digenic epistasis of quantitative traits in biparental populations. Theor. Appl. Genet. 116, 243–260.
Mackill D. J. and Ni J. J. 2001 Molecular mapping and marker assisted selection for major gene traits in rice. In Rice genetics IV (G. S. Khush, D. S. Brar and B. Hardy), pp. 137–151. Science Publisher, Enfield, USA.
Marathi B., Guleria S., Mohapatra T., Parsad R., Mariappan N., Kurungara V. K. et al. 2012 QTL analysis of novel genomic regions associated with yield and yield related traits in new plant type based recombinant inbred lines of rice (Oryza sativa L.). BMC Plant Biol. 12, 137.
McCouch S. R. 2008 Gene nomenclature system for rice. Rice 1, 72–84.
Nelson J. C., McClung A. M., Fjellstrom R. G., Moldenhauer K. A. K., Boza E., Jodari F. et al. 2011 Mapping QTL main and interaction influences on milling quality in elite US rice germplasm. Theor. Appl. Genet. 122, 291–309.
Song X. J., Huang W., Shi M., Zhu M. Z. and Lin H. X. 2007 A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat. Genet. 39, 623–630.
Wang J. 2009 Inclusive composite interval mapping of quantitative trait genes. Acta. Agron. Sin. 35, 239–245.
Wang J. K., Wan X. Y., Li H. H., Pfeiffer W. H., Crouch J. and Wan J. M. 2007 Application of identified QTL-marker associations in rice quality improvement through a design breeding approach. Theor. Appl. Genet. 115, 87–100.
Yamakawa H., Hirose T., Kuroda M. and Yamaguchi T. 2007 Comprehensive expression profiling of rice grain filling-related genes under high temperature using DNA microarray. Plant Physiol. 144, 258–277.
Zhou L., Jiang L., Liu X., Chen H., Chen L., Liu S. and Wan J. 2009 QTL mapping and interaction analysis for 1000-grain weight and percentage of grains with chalkiness in rice. Acta. Agron. Sin. 35, 255–261.
Acknowledgements
We thank Dr Song Yan for providing research materials and molecular-marker data. We thank Tsuyu Ando, Toshio Yamamoto, Masahiro Yano et al. for construction of the ILs population. We also thank undergraduate students Jianyong Hu, Guangxu Zhang and Ningpan Zhang in the agronomy department for helping to collect the phenotypic data. This research is financially supported by Jiangxi Province Youth Science Fund Project (20122BAB214014), Doctoral Fund of Ministry of Education of China (20123603120001) and Jiangxi Provincial Department of Education Science and Technology Project (GJJ12222).
Author information
Authors and Affiliations
Corresponding author
Additional information
[Bian J.-M., Shi H., Li C.-J., Zhu C.-L., Yu Q.-Y., Peng X.-S., Fu J.-R., He X.-P., Chen X.-R., Hu L.-F., Ouyang L.-J. and He H.-H. 2013 QTL mapping and correlation analysis for 1000-grain weight and percentage of grains with chalkiness in rice. J. Genet. 92, xx–xx]
Jian-Min Bian and Huan Shi contributed equally to this work.
Rights and permissions
About this article
Cite this article
BIAN, JM., SHI, H., LI, CJ. et al. QTL mapping and correlation analysis for 1000-grain weight and percentage of grains with chalkiness in rice. J Genet 92, 281–287 (2013). https://doi.org/10.1007/s12041-013-0267-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12041-013-0267-6