Abstract
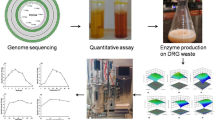
Cellulolytic bacteria from cattle rumen with ability to hydrolyze cellulose rich biomass were explored. The study selected Paenibacillus polymyxa ND24 from 847 isolates as the most potent strain, which can efficiently produce cellulase by utilizing sugarcane bagasse, rice straw, corn starch, CMC, and avicel as a sole carbon source. On annotation of P. polymyxa ND24 genome, 116 members of glycoside hydrolase (GH) family from CAZy clusters were identified and the presence of 10 potential cellulases was validated using protein folding information. Cellulase production was further demonstrated at lab-scale 5-L bioreactor exhibiting maximum endoglucanase activity up to 0.72 U/mL when cultivated in the medium containing bagasse (2% w/v) after 72 h. The bagasse hydrolysate so produced was further utilized for efficient biogas production. The presence of diverse hydrolytic enzymes and formidable cellulase activity supports the use of P. polymyxa ND24 for cost-effective bioprocessing of cellulosic biomass.





Similar content being viewed by others
References
FitzPatrick, M., Champagne, P., Cunningham, M. F., & Whitney, R. A. (2010). A biorefinery processing perspective: treatment of lignocellulosic materials for the production of value-added products. Bioresource Technology, 101, 8915–8922.
Lynd, L. R., Weimer, P. J., Van Zyl, W. H., & Pretorius, I. S. (2002). Microbial cellulose utilization: fundamentals and biotechnology. Microbiology and Molecular Biology Reviews, 66, 506–577.
Jain, K. K., Kumar, S., Deswal, D., & Kuhad, R. C. (2017). Improved Production of Thermostable Cellulase from Thermoascus aurantiacus RCKK by Fermentation Bioprocessing and its application in the hydrolysis of office waste paper, algal pulp, and biologically treated wheat straw. Applied Biochemistry and Biotechnology, 181, 784–800.
Srivastava, N., Srivastava, M., Manikanta, A., Singh, P., Ramteke, P. W., Mishra, P. K., & Malhotra, B. D. (2017). Production and optimization of physicochemical parameters of cellulase using untreated orange waste by newly isolated Emericella variecolor NS3. Applied Biochemistry and Biotechnology, 183, 601–612.
Pandey, S., Singh, S., Yadav, A. N., Nain, L., & Saxena, A. K. (2013). Phylogenetic diversity and characterization of novel and efficient cellulase producing bacterial isolates from various extreme environments. Bioscience, Biotechnology, and Biochemistry, 77, 1474–1480.
Woo, H. L., Hazen, T. C., Simmons, B. A., & DeAngelis, K. M. (2014). Enzyme activities of aerobic lignocellulolytic bacteria isolated from wet tropical forest soils. Systematic and Applied Microbiology, 37, 60–67.
Ma, J., Zhang, K., Liao, H., Hector, S. B., Shi, X., Li, J., Liu, B., Xu, T., Tong, C., Liu, X., & Zhu, Y. (2016). Genomic and secretomic insight into lignocellulolytic system of an endophytic bacterium Pantoea ananatis Sd-1. Biotechnology for Biofuels, 9(1), 25.
Irfan, M., Tayyab, A., Hasan, F., Khan, S., Badshah, M., & Shah, A. A. (2017). Production and characterization of organic solvent-tolerant cellulase from Bacillus amyloliquefaciens AK9 isolated from hot spring. Applied Biochemistry and Biotechnology, 182, 1390–1402.
Wibberg, D., Al-Dilaimi, A., Busche, T., Wedderhoff, I., Schrempf, H., Kalinowski, J., & de Orué Lucana, D. O. (2016). Complete genome sequence of Streptomyces reticuli, an efficient degrader of crystalline cellulose. Journal of Biotechnology, 222, 13–14.
Dassa, B., Borovok, I., Ruimy-Israeli, V., Lamed, R., Flint, H. J., Duncan, S. H., Henrissat, B., Coutinho, P., Morrison, M., Mosoni, P., & Yeoman, C. J. (2014). Rumen cellulosomics: divergent fiber-degrading strategies revealed by comparative genome-wide analysis of six ruminococcal strains. PLoS One, 9(7), e99221.
Suen, G., Weimer, P. J., Stevenson, D. M., Aylward, F. O., Boyum, J., Deneke, J., Drinkwater, C., Ivanova, N. N., Mikhailova, N., Chertkov, O., & Goodwin, L. A. (2011). The complete genome sequence of Fibrobacter succinogenes S85 reveals a cellulolytic and metabolic specialist. PLoS One, 6(4), e18814.
Khampa, S., Wanapat, M., Wachirapakorn, C., Nontaso, N., & Wattiaux, M. (2006). Effects of urea level and sodium DL-malate in concentrate containing high cassava chip on ruminal fermentation efficiency, microbial protein synthesis in lactating dairy cows raised under tropical condition. Asian-Australasian Journal of Animal Sciences, 19, 837–844.
Pawar, K. D., Dar, M. A., Rajput, B. P., & Kulkarni, G. J. (2015). Enrichment and identification of cellulolytic bacteria from the gastrointestinal tract of giant African snail, Achatinafulica. Biotechnology and Applied Biochemistry, 175, 1971–1980.
Dafale, N. A. (2011). Exploration of genetic information from dynamic microbial populations for enhancing the efficiency of azo-dye-degrading systems. Environmental Reviews, 9, 310–332.
Saxton, M. A., Naqvi, N. S., Rahman, F., Thompson, C. P., Chambers, R. M., Kaste, J. M., & Williamson, K. E. (2016). Site-specific environmental factors control bacterial and viral diversity in storm water retention ponds. Aquatic Microbial Ecology, 77, 23–36.
Bohra, V., Dafale, N. A., & Purohit, H. J. (2018). Paenibacillus polymyxa ND25: candidate genome for lignocellulosic biomass utilization. 3 Biotech, 8(5), 248.
Dar, M. A., Pawar, K. D., Jadhav, J. P., & Pandit, R. S. (2015). Isolation of cellulolytic bacteria from the gastro-intestinal tract of Achatinafulica (Gastropoda: Pulmonata) and their evaluation for cellulose biodegradation. International Biodeterioration and Biodegradation, 98, 73–80.
Dafale, N., Agrawal, L., Kapley, A., Meshram, S., Purohit, H., & Wate, S. (2010). Selection of indicator bacteria based on screening of 16S rDNA metagenomic library from a two-stage anoxic–oxic bioreactor system degrading azo dyes. Bioresource Technology, 101, 476–484.
Nitisinprasert, S., & Temmes, A. (1991). The characteristics of a new non-spore-forming cellulolytic mesophilic anaerobe strain CM126 isolated from municipal sewage sludge. The Journal of Applied Bacteriology, 71, 154–161.
Miller, G. L. (1959). Use of dinitrosalicylic acid reagent for determination of reducing sugar. Analytical Chemistry, 31, 426–428.
An, C. L., Lim, W. J., Hong, S. Y., Kim, E. J., Shin, E. C., Kim, M. K., Lee, J. R., Park, S. R., Woo, J. G., Lim, Y. P., & Yun, H. D. (2004). Analysis of bgl operon structure and characterization of β-glucosidase from Pectobacterium carotovorum subsp. carotovorum LY34. Bioscience, Biotechnology, and Biochemistry, 68, 2270–2278.
Kapley, A., Tanksale, H., Sagarkar, S., Prasad, A. R., Kumar, R. A., Sharma, N., Qureshi, A., & Purohit, H. J. (2016). Antimicrobial activity of Alcaligenes sp. HPC 1271 against multidrug resistant bacteria. Functional & Integrative Genomics, 16, 57–65.
Huang, L., Zhang, H., Wu, P., Entwistle, S., Li, X., Yohe, T., Yi, H., Yang, Z., & Yin, Y. (2017). dbCAN-seq: a database of carbohydrate-active enzyme (CAZyme) sequence and annotation. Nucleic Acids Research, 46(D1), D516–D521.
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M., & Henrissat, B. (2014). The Carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Research, 42, 490–495.
Pal, R. R., Khardenavis, A. A., & Purohit, H. J. (2015). Identification and monitoring of nitrification and denitrification genes in Klebsiella pneumoniae EGD-HP19-C for its ability to perform heterotrophic nitrification and aerobic denitrification. Functional & Integrative Genomics, 15, 63–76.
Pettersen, E. F., Goddard, T. D., Huang, C. C., Couch, G. S., Greenblatt, D. M., Meng, E. C., & Ferrin, T. E. (2004). UCSF Chimera--a visualization system for exploratory research and analysis. Journal of Computational Chemistry, 25, 1605–1612.
Tikariha, H., Pal, R. R., Qureshi, A., Kapley, A., & Purohit, H. J. (2016). In silico analysis for prediction of degradative capacity of Pseudomonas putida SF1. Gene, 591(2), 382–392.
Zeng, R., Xiong, P., & Wen, J. (2006). Characterization and gene cloning of a cold-active cellulase from a deep-sea psychrotrophic bacterium Pseudoalteromonas sp. DY3. Extremophiles, 10, 79–82.
Sandgren, M., Wu, M., Karkehabadi, S., Mitchinson, C., Kelemen, B. R., Larenas, E. A., Ståhlberg, J., & Hansson, H. (2013). The structure of a bacterial cellobiohydrolase: the catalytic core of the Thermobifidafusca family GH6 cellobiohydrolase Cel6B. Journal of Molecular Biology, 425, 622–635.
Tsai, L. C., Amiraslanov, I., Chen, H. R., Chen, Y. W., Lee, H. L., Liang, P. H., & Liaw, Y. C. (2015). Structures of exoglucanase from Clostridium cellulovorans: cellotetraose binding and cleavage. Acta Crystallographica Section F Structural Biology Communications, 71(10), 1264–1272.
Isorna, P., Polaina, J., Latorre-García, L., Cañada, F. J., González, B., & Sanz-Aparicio, J. (2007). Crystal structures of Paenibacillus polymyxa β-glucosidase B complexes reveal the molecular basis of substrate specificity and give new insights into the catalytic machinery of family I glycosidases. Journal of Molecular Biology, 371, 1204–1218.
Gulhane, M., Khardenavis, A. A., Karia, S., Pandit, P., Kanade, G. S., Lokhande, S., Vaidya, A. N., & Purohit, H. J. (2016). Biomethanation of vegetable market waste in an anaerobic baffled reactor: Effect of effluent recirculation and carbon mass balance analysis. Bioresource Technology, 215, 100–109.
Lee, Y. J., Kim, B. K., Lee, B. H., Jo, K. I., Lee, N. K., Chung, C. H., Lee, Y. C., & Lee, J. W. (2008). Purification and characterization of cellulase produced by Bacillus amyoliquefaciens DL-3 utilizing rice hull. Bioresource Technology, 99, 378–386.
Gao, Y., Xu, J., Zhang, Y., Yu, Q., Yuan, Z., & Liu, Y. (2013). Effects of different pretreatment methods on chemical composition of sugarcane bagasse and enzymatic hydrolysis. Bioresource Technology, 144, 396–400.
Sethi, S., Datta, A., Gupta, B. L., & Gupta, S. (2013). Optimization of cellulase production from bacteria isolated from soil. ISRN Biotechnology, 2013, 985685. https://doi.org/10.5402/2013/985685.
Padilha, I. Q. M., Carvalho, L. C. T., Dias, P. V. S., Grisi, T. C. S. L., Silva, F. L., Santos, S. F. M., & Araújo, D. A. M. (2015). Production and characterization of thermophilic carboxymethyl cellulase synthesized by Bacillus sp. growing on sugarcane bagasse in submerged fermentation. Brazilian Journal of Chemical Engineering, 32, 35–42.
Waghmare, P. R., Kshirsagar, S. D., Saratale, R. G., Govindwar, S. P., & Saratale, G. D. (2014). Production and characterization of cellulolytic enzymes by isolated Klebsiella sp. PRW-1 using agricultural waste biomass. Emirates Journal of Food and Agriculture, 26, 44–59.
Goyal, V., Mittal, A., Bhuwal, A. K., Singh, G., Yadav, A., & Aggarwal, N. K. (2014). Parametric optimization of cultural conditions for carboxymethyl cellulase production using pretreated rice straw by Bacillus sp. 313SI under stationary and shaking conditions. Biotechnology Research International, 2014, 651839. https://doi.org/10.1155/2014/651839.
Dantur, K. I., Enrique, R., Welin, B., & Castagnaro, A. P. (2015). Isolation of cellulolytic bacteria from the intestine of Diatraea saccharalis larvae and evaluation of their capacity to degrade sugarcane biomass. AMB Express, 5(1), 15.
Ladeira, S. A., Cruz, E., Delatorre, A. B., Barbosa, J. B., & Leal Martins, M. L. (2015). Cellulase production by thermophilic Bacillus sp: SMIA-2 and its detergent compatibility. Electronic Journal of Biotechnology, 18, 110–115.
Gastelum-Arellanez, A., Paredes-López, O., & Olalde-Portugal, V. (2014). Extracellular endoglucanase activity from Paenibacillus polymyxa BEb-40: production, optimization and enzymatic characterization. World Journal of Microbiology and Biotechnology, 30, 2953–2965.
Ghio, S., Insani, E. M., Piccinni, F. E., Talia, P. M., Grasso, D. H., & Campos, E. (2016). GH10 XynA is the main xylanase identified in the crude enzymatic extract of Paenibacillus sp. A59 when grown on xylan or lignocellulosic biomass. Microbiological Research, 186, 16–26.
Ekperigin, M. M. (2007). Preliminary studies of cellulase production by Acinetobacter anitratus and Branhamella sp. African Journal of Biotechnology, 6, 028–033.
Deka, D., Bhargavi, P., Sharma, A., Goyal, D., Jawed, M., & Goyal, A. (2011). Enhancement of cellulase activity from a new strain of Bacillus subtilis by medium optimization and analysis with various cellulosic substrates. Enzyme Research, 2011, 151656. https://doi.org/10.4061/2011/151656.
Acknowledgements
Miss Varsha Bohra acknowledges Department of Science and Technology of India for awarding Junior Research Fellowship (IF150088) for carrying out the work. The manuscript has been checked for plagiarism by Knowledge Resource Centre, CSIR-NEERI, Nagpur, India, and assigned KRC No. CSIR-NEERI/KRC/2017/AUG/EBGD/2.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Electronic Supplementary Material
ESM 1
(DOCX 13 kb)
Rights and permissions
About this article
Cite this article
Bohra, V., Tikariha, H. & Dafale, N.A. Genomically Defined Paenibacillus polymyxa ND24 for Efficient Cellulase Production Utilizing Sugarcane Bagasse as a Substrate. Appl Biochem Biotechnol 187, 266–281 (2019). https://doi.org/10.1007/s12010-018-2820-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-018-2820-5