Abstract
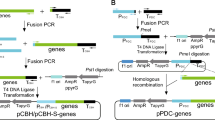
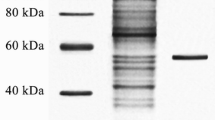
To heterologously express a Talaromyces thermophilus lipase gene in Trichoderma reesei, an efficient binary vector pChph-pCBH1sigpro-ttl which includes a newly designed cbh1 promoter and hygromycin-resistant marker was constructed. This plasmid was then transformed into T. reesei via improved Agrobacterium EHA 105-mediated transformation. After modification of co-culture conditions and enzymolysis treatment of conidia, 258 transformants were produced. A two-step screening method based on antibiotic resistance and capacity to utilize lactose and tributyrin was introduced to further select promising candidates, which would be additionally verified by PCR analysis, sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE), and lipase activity assay. Lipase production was carried out in shaking flasks, and the activity reached 241 IU/mL (7415.4 IU/mg) after 84-h fermentation. It was found that this lipase performed high alkali and thermostable tolerance with the optimal pH 9.5 and temperature 60 °C, and it could retain more than 70 % activity after being disposed in pH 11 or 70 °C for 1 h. This study herein would benefit the genetic engineering of T. reesei and the industrial application of this important fungal lipase.








Similar content being viewed by others
References
Brzozowski, A. M., Derewenda, U., Derewenda, Z. S., Dodson, G. G., Lawson, D. M., Turkenburg, J. P., Bjorkling, F., Huge-Jensen, B., Patkar, S. A., & Thim, L. (1991). A model for interfacial activation in lipases from the structure of a fungal lipase-inhibitor complex. Nature, 351, 491–494.
Stergiou, P.-Y., Foukis, A., Filippou, M., Koukouritaki, M., Parapouli, M., Theodorou, L. G., Hatziloukas, E., Afendra, A., Pandey, A., & Papamichael, E. M. (2013). Advances in lipase-catalyzed esterification reactions. Biotechnology Advances, 31, 1846–1859.
Dai, D., & Xia, L. (2006). Effect of lipase immobilization on resolution of (R, S)-2-octanol in nonaqueous media using modified ultrastable-Y molecular sieve as support. Applied Biochemistry and Biotechnology, 134, 39–50.
Gusakov, A. V. (2011). Alternatives to Trichoderma reesei in biofuel production. Trends in Biotechnology, 29, 419–425.
Amore, A., & Faraco, V. (2012). Potential of fungi as category I consolidated bioprocessing organisms for cellulosic ethanol production. Renewable and Sustainable Energy Reviews, 16, 3286–3301.
Jorgensen, M., Skovlund, D., Johannesen, P., & Mortensen, U. (2014). A novel platform for heterologous gene expression in Trichoderma reesei (Teleomorph Hypocrea jecorina). Microbial Cell Factories, 13, 33.
Nevalainen, K. M. H., Te'o, V. S. J., & Bergquist, P. L. (2005). Heterologous protein expression in filamentous fungi. Trends in Biotechnology, 23, 468–474.
Romdhane, I. B.-B., Romdhane, Z. B., Gargouri, A., & Belghith, H. (2011). Esterification activity and stability of Talaromyces thermophilus lipase immobilized onto chitosan. Journal of Molecular Catalysis B: Enzymatic, 68, 230–239.
Romdhane, I. B.-B., Fendri, A., Gargouri, Y., Gargouri, A., & Belghith, H. (2010). A novel thermoactive and alkaline lipase from Talaromyces thermophilus fungus for use in laundry detergents. Biochemical Engineering Journal, 53, 112–120.
Valero, F. (2012). Lipases and phospholipases, vol. 861. In G. Sandoval (Ed.), Methods in molecular biology (pp. 161–178) Humana Press.
Kontkanen, H., Reinikainen, T., & Saloheimo, M. (2006). Cloning and expression of a Melanocarpus albomyces steryl esterase gene in Pichia pastoris and Trichoderma reesei. Biotechnology and Bioengineering, 94, 407–415.
Jin, X., Meng, N., & Xia, L.-M. (2011). Expression of an endo-β-1, 4-glucanase gene from Orpinomyces PC-2 in Pichia pastoris. International Journal of Molecular Sciences, 12, 3366–3380.
Jin, X., & Xia, L. (2011). Heterologous expression of an endo-β-1,4-glucanase gene from the anaerobic fungus Orpinomyces PC-2 in Trichoderma reesei. World Journal of Microbiology and Biotechnology, 27, 2913–2920.
Meyer, V. (2008). Genetic engineering of filamentous fungi—progress, obstacles and future trends. Biotechnology Advances, 26, 177–185.
Martinez, D., Berka, R. M., Henrissat, B., Saloheimo, M., Arvas, M., Baker, S. E., Chapman, J., Chertkov, O., Coutinho, P. M., Cullen, D., Danchin, E. G. J., Grigoriev, I. V., Harris, P., Jackson, M., Kubicek, C. P., Han, C. S., Ho, I., Larrondo, L. F., de Leon, A. L., Magnuson, J. K., Merino, S., Misra, M., Nelson, B., Putnam, N., Robbertse, B., Salamov, A. A., Schmoll, M., Terry, A., Thayer, N., Westerholm-Parvinen, A., Schoch, C. L., Yao, J., Barabote, R., Nelson, M. A., Detter, C., Bruce, D., Kuske, C. R., Xie, G., Richardson, P., Rokhsar, D. S., Lucas, S. M., Rubin, E. M., Dunn-Coleman, N., Ward, M., & Brettin, T. S. (2008). Genome sequencing and analysis of the biomass-degrading fungus Trichoderma reesei (syn. Hypocrea jecorina). Nature Biotechnology, 26, 553–560.
Michielse, C., Hooykaas, P. J., van den Hondel, C. M. J. J., & Ram, A. J. (2005). Agrobacterium-mediated transformation as a tool for functional genomics in fungi. Current Genetics, 48, 1–17.
Su, X., Schmitz, G., Zhang, M., Mackie, R. I., & Cann, I. K. O. (2012). Heterologous gene expression in filamentous fungi. Advances in Applied Microbiology, 81, 1–61.
Zhong, Y. H., Wang, X. L., Wang, T. H., & Jiang, Q. (2007). Agrobacterium-mediated transformation (AMT) of Trichoderma reesei as an efficient tool for random insertional mutagenesis. Applied Microbiology and Biotechnology, 73, 1348–1354.
Gu, B., & Xia, L. (2013). High expression of a neutral endo-β-glucanase gene from Humicola insolens in Trichoderma reesei. Journal of Industrial Microbiology & Biotechnology, 40, 773–779.
Fang, H., & Xia, L. (2013). High activity cellulase production by recombinant Trichoderma reesei ZU-02 with the enhanced cellobiohydrolase production. Bioresource Technology, 144, 693–697.
Te’o, V. S. J., Cziferszky, A. E., Bergquist, P. L., & Nevalainen, K. M. H. (2000). Codon optimization of xylanase gene xynB from the thermophilic bacterium Dictyoglomus thermophilum for expression in the filamentous fungus Trichoderma reesei. FEMS Microbiology Letters, 190, 13–19.
Villalobos, A., Ness, J., Gustafsson, C., Minshull, J., & Govindarajan, S. (2006). Gene designer: a synthetic biology tool for constructing artificial DNA segments. BMC Bioinformatics, 7, 285.
Wang, B., & Xia, L. (2011). High efficient expression of cellobiase gene from Aspergillus niger in the cells of Trichoderma reesei. Bioresource Technology, 102, 4568–4572.
Meyers, S. A., Cuppett, S. L., & Hutkins, R. W. (1996). Lipase production by lactic acid bacteria and activity on butter oil. Food Microbiology, 13, 383–389.
Weld, R. J., Plummer, K. M., Carpenter, M. A., & Ridgway, H. J. (2006). Approaches to functional genomics in filamentous fungi. Cell Research, 16, 31–44.
Peterson, R., & Nevalainen, H. (2012). Trichoderma reesei RUT-C30—thirty years of strain improvement. Microbiology, 158, 58–68.
Mach, R., & Zeilinger, S. (2003). Regulation of gene expression in industrial fungi: Trichoderma. Applied Microbiology and Biotechnology, 60, 515–522.
Ries, L., Belshaw, N. J., Ilmén, M., Penttilä, M. E., Alapuranen, M., & Archer, D. B. (2014). The role of CRE1 in nucleosome positioning within the cbh1 promoter and coding regions of Trichoderma reesei. Applied Microbiology and Biotechnology, 98, 749–762.
Stricker, A. R., Grosstessner-Hain, K., Würleitner, E., & Mach, R. L. (2006). Xyr1 (xylanase regulator 1) regulates both the hydrolytic enzyme system and d-xylose metabolism in Hypocrea jecorina. Eukaryotic Cell, 5, 2128–2137.
Aro, N., Saloheimo, A., Ilmén, M., & Penttilä, M. (2001). ACEII, a novel transcriptional activator involved in regulation of cellulase and xylanase genes of Trichoderma reesei. Journal of Biological Chemistry, 276, 24309–24314.
Combier, J.-P., Melayah, D., Raffier, C., Gay, G., & Marmeisse, R. (2003). Agrobacterium tumefaciens-mediated transformation as a tool for insertional mutagenesis in the symbiotic ectomycorrhizal fungus Hebeloma cylindrosporum. FEMS Microbiology Letters, 220, 141–148.
Gardiner, D., & Howlett, B. (2004). Negative selection using thymidine kinase increases the efficiency of recovery of transformants with targeted genes in the filamentous fungus Leptosphaeria maculans. Current Genetics, 45, 249–255.
Liu, T., Liu, L., Jiang, X., Hou, J., Fu, K., Zhou, F., & Chen, J. (2010). Agrobacterium-mediated transformation as a useful tool for the molecular genetic study of the phytopathogen Curvularia lunata. European Journal of Plant Pathology, 126, 363–371.
Abuodeh, R. O., Orbach, M. J., Mandel, M. A., Das, A., & Galgiani, J. N. (2000). Genetic transformation of Coccidioides immitis facilitated by Agvobactevium tumefaciens. Journal of Infectious Diseases, 181, 2106–2110.
Zhang, Y., Li, G., He, D., Yu, B., Yokoyama, K., & Wang, L. (2011). Efficient insertional mutagenesis system for the dimorphic pathogenic fungus Sporothrix schenckii using Agrobacterium tumefaciens. Journal of Microbiological Methods, 84, 418–422.
Giese, H., Kruithof, P., Meier, K., Sieben, M., Antonov, E., Hommes, R. W. J., & Büchs, J. (2014). Improvement and scale-down of a Trichoderma reesei shake flask protocol to microtiter plates enables high-throughput screening. Journal of Bioscience and Bioengineering, 118, 702–709.
Li, C., Yang, Z., He Can Zhang, R., Zhang, D., Chen, S., & Ma, L. (2013). Effect of pH on cellulase production and morphology of Trichoderma reesei and the application in cellulosic material hydrolysis. Journal of Biotechnology, 168, 470–477.
Prasetyo, J., Sumita, S., Okuda, N., & Park, E. (2010). Response of cellulase activity in pH-controlled cultures of the filamentous fungus Acremonium cellulolyticus. Applied Biochemistry and Biotechnology, 162, 52–61.
Qin, L.-N., Cai, F.-R., Dong, X.-R., Huang, Z.-B., Tao, Y., Huang, J.-Z., & Dong, Z.-Y. (2012). Improved production of heterologous lipase in Trichoderma reesei by RNAi mediated gene silencing of an endogenic highly expressed gene. Bioresource Technology, 109, 116–122.
Rajesh, E. M., Arthe, R., Rajendran, R., Balakumar, C., Pradeepa, N., & Anitha, S. (2010). Investigation of lipase production by Trichoderma reesei and optimization of production parameters. Electronic Journal of Environmental, Agricultural and Food Chemistry, 9, 1177–1189.
Gouka, R. J., Punt, P. J., & van den Hondel, C. A. M. J. J. (1997). Efficient production of secreted proteins by Aspergillus: progress, limitations and prospects. Applied Microbiology and Biotechnology, 47, 1–11.
Ahamed, A., & Vermette, P. (2008). Culture-based strategies to enhance cellulase enzyme production from Trichoderma reesei RUT-C30 in bioreactor culture conditions. Biochemical Engineering Journal, 40, 399–407.
Dutta, S., & Ray, L. (2009). Production and characterization of an alkaline thermostable crude lipase from an isolated strain of Bacillus cereus C7. Applied Biochemistry and Biotechnology, 159, 142–154.
Gupta, R., Gupta, N., & Rathi, P. (2004). Bacterial lipases: an overview of production, purification and biochemical properties. Applied Microbiology and Biotechnology, 64, 763–781.
Papaparaskevas, D., Christakopoulos, P., Kekos, D., & Macris, B. (1992). Optimizing production of extracellular lipase from Rhodotorula glutinis. Biotechnology Letters, 14, 397–402.
Costa, M. A. F., & Peralta, R. M. (1999). Production of lipase by soil fungi and partial characterization of lipase from a selected strain (Penicillium wortmannin). Journal of Basic Microbiology, 39, 11–15.
Singh, A., & Mukhopadhyay, M. (2012). Overview of fungal lipase: a review. Applied Biochemistry and Biotechnology, 166, 486–520.
Romdhane, I. B.-B., Frikha, F., Maalej-Achouri, I., Gargouri, A., & Belghith, H. (2012). Gene cloning and molecular characterization of the Talaromyces thermophilus lipase catalyzed efficient hydrolysis and synthesis of esters. Gene, 494, 112–118.
Fickers, P., Marty, A., & Nicaud, J. M. (2011). The lipases from Yarrowia lipolytica: genetics, production, regulation, biochemical characterization and biotechnological applications. Biotechnology Advances, 29, 632–644.
Acknowledgments
This work was supported by the National High-tech R&D Program (2007AA05Z401) and the Program for Zhejiang Leading Team of S&T Innovation (2011R50002) of China.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
ESM 1
Sequence alignment of the original and optimized lipase sequence (GIF 498 kb)
Rights and permissions
About this article
Cite this article
Zhang, X., Li, X. & Xia, L. Heterologous Expression of an Alkali and Thermotolerant Lipase from Talaromyces thermophilus in Trichoderma reesei . Appl Biochem Biotechnol 176, 1722–1735 (2015). https://doi.org/10.1007/s12010-015-1673-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-015-1673-4