Abstract
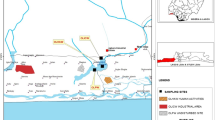
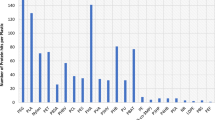
In this work the archaea and eubacteria community of a hypersaline produced water from the Campos Basin that had been transported and discharged to an onshore storage facility was evaluated by 16S recombinant RNA (rRNA) gene sequence analysis. The produced water had a hypersaline salt content of 10 (w/v), had a carbon oxygen demand (COD) of 4,300 mg/l and contains phenol and other aromatic compounds. The high salt and COD content and the presence of toxic phenolic compounds present a problem for conventional discharge to open seawater. In previous studies, we demonstrated that the COD and phenolic content could be largely removed under aerobic conditions, without dilution, by either addition of phenol degrading Haloarchaea or the addition of nutrients alone. In this study our goal was to characterize the microbial community to gain further insight into the persistence of reservoir community members in the produced water and the potential for bioremediation of COD and toxic contaminants. Members of the archaea community were consistent with previously identified communities from mesothermic reservoirs. All identified archaea were located within the phylum Euryarchaeota, with 98 % being identified as methanogens while 2 % could not be affiliated with any known genus. Of the identified archaea, 37 % were identified as members of the strictly carbon-dioxide-reducing genus Methanoplanus and 59 % as members of the acetoclastic genus Methanosaeta. No Haloarchaea were detected, consistent with the need to add these organisms for COD and aromatic removal. Marinobacter and Halomonas dominated the eubacterial community. The presence of these genera is consistent with the ability to stimulate COD and aromatic removal with nutrient addition. In addition, anaerobic members of the phyla Thermotogae, Firmicutes, and unclassified eubacteria were identified and may represent reservoir organisms associated with the conversion hydrocarbons to methane.




Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Arahal DR, Dewhirst FE, Paster BJ, Volcani BE, Ventosa A (1996) Phylogenetic analyses of some extremely halophilic Archaea isolated from Dea Sea water, determined on the basis of their 16S rRNA sequences. Appl Environ Microbiol 62:3779–3786
Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K (1989) Current protocols in molecular biology. John Wiley & Sons, New York, New York
Bonfá MRL, Grossman MJ, Mellado E, Durrant LR (2011) Biodegradation of aromatic hydrocarbons by Haloarchaea and their use for the reduction of the chemical oxygen demand of hypersaline petroleum produced water. Chemosphere 84:1671–1676
Borzenkov IA, Milekhina ET, Gotoeva MT, Rozanova EP, Beliaev SS (2006) The properties of hydrocarbon-oxidizing bacteria isolated from the oilfields of Tatarstan, Western Siberia, and Vietnam. Mikrobiologia 75:82–89
Chang HW, Nam YD, Kwon HY, Park JR, Lee JS, Yoon JH, An KG, Bae JW (2007) Marinobacterium halophilum sp. nov., a marine bacterium isolated from the Yellow Sea. Int J Syst Evol Micribiol 57:77–80
Ciobanu MC, Rabineau M, Droz L, Révillon S, Ghiglione JF, Dennielou B, Jorry SJ, Kallmeyer J, Etoubleau J, Pignet P, Crassous P, Vandenabeele-Trambouze O, Laugier J, Guégan M, Godfroy A, Alain K (2012) Sedimentological imprint on subseafloor microbial communities in Western Mediterranean Sea Quaternary sediments. Biogeosciences 9:3491–3512
Clark CE, Veil JA (2009) A white paper describing produced water volumes and managements practices in the United States. Prepared by Argonne National Laboratory, Argonne, Illinois for the US Department of Energy, National Energy Technology Laboratory
Coates JD, Lonergan DJ, Philips EJP, Jenter H, Lovley DR (1995) Desulfuromonas palmitatis sp. nov, a marine dissimilatory Fe(III) reducer that can oxidize long-chain fatty acids. Arch Microbiol 164:406–413
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ, Kulam-Syed-Mohideen AS, McGarrell DM, Marsh T, Garrity GM, Tiedje JM (2009) The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37(Database issue):D141–D145
Dahle H, Garshol F, Madsen M, Birkeland NK (2008) Microbial community structure analysis of produced water from a high-temperature North Sea oil-field. Antonie Van Leeuwenhoek 93(1–2):37–49
de la Haba RR, Arahal DR, Márquez MC, Ventosa A (2010) Phylogenetic relationships within the family Halomonadaceae based on comparative 23S and 16S rRNA gene sequence analysis. Int J Syst Evol Microbiol 60:737–748
Desper R, Gascuel O (2004) Theoretical foundations of the balanced minimum evolution method of phylogenetic inference and its relationship to weighted least-squares tree fitting. Mol Biol Evol 21:587–598
García MT, Ventosa A, Mellado E (2005) Catabolic versatility of aromatic compound-degrading halophilic bacteria. FEMS Microbiol Ecol 54:97–109
Grabowski A, Nercessian O, Fayolle F, Blanchet D, Jeanthon C (2005) Microbial diversity in production waters of a low-temperature biodegraded oil reservoir. FEMS Microbiol Ecol 54(3):427–443
Head IM, Jones DM, Larter SR (2003) Biological activity in the deep subsurface and the origin of heavy oil. Nature 426:344–352
Huson DH, Richter DC, Rausch C, Dezulian T, Franz M, Rupp R (2007) Dendroscope: an interactive viewer for large phylogenetic trees. BMC Bioinforma 8(1):460
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (Ed), Mammalian protein metabolism, Academic Press, New York, 1969, vol. 3 pp. 21-132)
Kaye JZ, Sylvan JB, Edwards KJ, Baross JA (2011) Halomonas and Marinobacter ecotypes from hydrothermal vent, subseafloor and deep-sea environments. FEMS Microbiol Ecol 75(1):123–133
Korenblum E, Souza DB, Penna M, Seldin L (2012) Molecular analysis of the bacterial communities in crude oil samples from two Brazilian offshore petroleum platforms. Int J Microbiol 2012:156537. doi:10.1155/2012/156537
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, New York, pp 115–175
Lever MA, Rouxel O, Alt JC, Shimizu N, Ono S, Coggon RM, Shanks WC III, Lapham L, Elvert M, Prieto-Mollar X, Hinrichs K-U, Inagaki F, Teske A (2013) Evidence for microbial carbon and sulfur cycling in deeply buried ridge flank basalt. Science 339:1305–1308
Magot M (2005) Indigenous microbial communities in oil fields. In: Ollivier B, Magot M (eds) Petroleum microbiology. ASM Press, Washington, pp 21–33
Mbadinga SM, Li KP, Zhou L, Wang LY, Yang SZ, Liu JF, Gu JD, Mu BZ (2012) Analysis of alkane-dependent methanogenic community derived from production water of a high-temperature petroleum reservoir. Appl Microbiol Biotechnol 96(2):531–542
McCormack P, Jones P, Hetheridge MJ, Rowland SJ (2001) Analysis of oilfield produced waters and production chemicals by electrospray ionisation multi-stage mass spectrometry (ESI-MSn). Water Res 35:3567–3578
Mohriak WU, Szatmari P, Anjos S (2012) Salt: geology and tectonics of selected Brazilian basins in their global context. Geol Soc Lond Spec Publ 363:131–158
Nesbo CL, Kumaraswamy R, Dlutek M, Doolittle WF, Foght J (2010) Searching for mesophilic Thermotogales bacteria: “mesotogas” in the wild. Appl Environ Microbiol 76:4896–4900
Oie CSI, Albaugh CE, Peyton BM (2007) Benzoate and salicylate degradation by Halomonas campisalis, an alkaliphilic and moderately halophilic microorganism. Water Res 41:1235–1242
Onstott TC, Hinton SM, Silver BJ, King HE Jr (2010) Coupling hydrocarbon degradation to anaerobic respiration and mineral diagenesis: theoretical constraints. Geobiology 8:69–88
Pham VD, Hnatow LL, Zhang S, Fallon RD, Jackson SC, Tomb JF, DeLong EF, Keeler SJ (2009) Characterizing microbial diversity in production water from an Alaskan mesothermic petroleum reservoir with two independent molecular methods. Environ Microbiol 11(1):176–187
Piubeli F, Grossman MJ, Fantinatti-Garboggini F, Durrant LR (2012) Enhanced reduction of COD and aromatics in petroleum-produced water using indigenous microorganisms and nutrient addition. Int Biodeterior Biodegrad 68:78–84
Reysenbach AL, Longnecker K, Kirshtein J (2000) Novel bacterial and archaeal lineages from an in situ growth chamber deployed at a mid-atlantic ridge hydrothermal vent. Appl Environ Microbiol 66(9):3798–3806
Satter A, Iqbal GM, Buchwalter J (2008) Practical enhanced reservoir engineering: assisted with simulation software. PennWell, Tulsa, Oklahoma
Speight JG (2007) The chemistry and technology of petroleum, 4th edn. Marcel Dekker, New York
Turner S, Pryer KM, Miao VPW, Palmer JD (1999) Investigating deep phylogenetic relationships among cyanobacteria and plastids by small subunit rRNA sequence analysis. J Eukaryot Microbiol 46:327–338
van der Kraan GM, Bruining J, Lomans BP, van Loosdrecht MCM, Muyzer G (2010) Microbial diversity of an oil-water processing site and its associated oil field: the possible role of microorganisms as information carriers from oil-associated environments. FEMS Microbiol Ecol 71:428–443
Vasconcellos SP, Crespim E, da Cruz GF, Senatore DB, Simioni KCM, dos Santos Neto EV, Marsaioli AJ, de Oliveira VM (2009) Isolation, biodegradation ability and molecular detection of hydrocarbon degrading bacteria in petroleum samples from a Brazilian offshore basin. Org Geochem 40(5):574–588
Wang LY, Duan RY, Liu JF, Yang SZ, Gu JD, Mu BZ (2012) Molecular analysis of the microbial community structures in water-flooding petroleum reservoirs with different temperatures. Biogeosciences 9:4645–4659
Wright ES, Yilmaz LS, Noguera DR (2012) DECIPHER, a search-based approach to chimera identification for 16 s rRNA sequences. Appl Environ Microbiol 78(3):717–725
Acknowledgments
We thank the Coordinator for the Improvement of Personnel in Higher Education (CAPES, Brazil), the National Council for the Development of Science and Technology (CNPq), and the Foundation for the Support of Science in São Paulo State (FAPESP) for financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Robert Duran
Rights and permissions
About this article
Cite this article
Piubeli, F., Grossman, M.J., Fantinatti-Garboggini, F. et al. Phylogenetic analysis of the microbial community in hypersaline petroleum produced water from the Campos Basin. Environ Sci Pollut Res 21, 12006–12016 (2014). https://doi.org/10.1007/s11356-014-3155-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-014-3155-6