Abstract
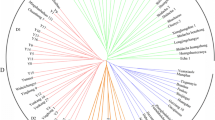
Tea (Camellia sinensis (L.) O. Kuntze) is the world’s most popular beverage crop. However, to date, no core collection has been selected from worldwide germplasm resources on the basis of genotype data. In this study, we analyzed 788 tea germplasm accessions using 23 simple sequence repeat (SSR) markers. Our population structure analysis divided the germplasms into a Japanese group and an exotic group. The latter could be divided into var. sinensis and var. assamica. The genetic diversity was higher in germplasms from China, Taiwan, India, and Sri Lanka than in those from other countries, and low in germplasms from Japan. Using the number of SSR alleles as a measure of genetic diversity, we developed a core collection consisting of 192 accessions and three subcore collections with 96, 48, and 24 accessions. Although the results might be affected by marker-selection bias, the core 192 collection adequately covered the range of variation of the 788 accessions in floral morphology, and the chemical composition of first-flush leaves. These collections will be powerful tools for breeding and genetic research in tea.






Similar content being viewed by others
References
Campana MG, Hunt HV, Jones H, White J (2011) CorrSieve: software for summarizing and evaluating Structure output. Mol Ecol Resour 11:349–352
Chakraborty R, Jin L (1993) Determination of relatedness between individuals using DNA fingerprinting. Hum Biol 65:875–895
Chen J, Wang PS, Xia YM, Xu M, Pei SJ (2005) Genetic diversity and differentiation of Camellia sinensis L. (cultivated tea) and its wild relatives in Yunnan province of China, revealed by morphology, biochemistry and allozyme studies. Genet Resour Crop Evol 52:41–52
Cockerham CC, Weir BS (1993) Estimation of gene flow from F-statistics. Evolution 47:855–863
Ercisli S (2012) The tea industry and improvement in Turkey. In: Chen L, Apostolides Z, Chen ZM (eds) Global tea breeding. Springer, Berlin, pp 309–321
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2640
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Fang W, Cheng H, Duan Y, Jiang X, Li X (2012) Genetic diversity and relationship of clonal tea (Camellia sinensis) cultivars in China as revealed by SSR markers. Plant Syst Evol 298:469–483
Goudet J (1995) FSTAT (version 1.2): a computer program to calculate F-statistics. J Hered 85:485–486
Gunasekare MTK (2012) Tea plant (Camellia sinensis) breeding in Sri Lanka. In: Chen L, Apostolides Z, Chen ZM (eds) Global tea breeding. Springer, Berlin, pp 125–176
Hashimoto M (2001) The origin of the tea plant. In: Proceedings of 2001 International Conference on O-CHA (tea) Culture and Science (Session II). Shizuoka, Japan, pp J5–J7
Hashimoto M, Takashi S (1978) Morphological studies on the origin of the tea plant V, a proposal of one place of origin by cluster analysis. Jpn J Trop Agric 21:93–101
Inazuka M, Tahira T, Hayashi K (1996) One-tube post-PCR fluorescent labeling of DNA fragments. Genome Res 6:551–557
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Kamunya SM, Wachira FN, Pathak RS, Muoki RC, Sharma RK (2012) Tea improvement in Kenya. In: Chen L, Apostolides Z, Chen ZM (eds) Global tea breeding. Springer, Berlin, pp 177–226
Kaundun SS, Matsumoto S (2002) Heterologous nuclear and chloroplast microsatellite amplification and variation in tea, Camellia sinensis. Genome 45:1041–1048
Kaundun SS, Matsumoto S (2003) Development of CAPS markers based on three key genes of the phenylpropanoid pathway in tea, Camellia sinensis (L.) O. Kuntze, and differentiation between assamica and sinensis varieties. Theor Appl Genet 106:375–383
Lai JA, Yang WC, Hsiao JY (2001) An assessment of genetic relationship in cultivated tea clones and native wild tea in Taiwan using RAPD and ISSR markers. Bot Bull Acad Sin 42:93–100
Langella O (1999) Populations 1.2.31: a population genetic software. CNRS UPR9034. http://bioinformatics.org/populations/. Accessed 8 Jan 2014
Li HL (1983) The domestication of plants in China: ecogeographical considerations. In: Keightley DN (ed) The origins of Chinese civilization. University of California Press, New Jersey, pp 21–63
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Matsumoto S, Kiriiwa Y, Takeda Y (2002) Differentiation of Japanese green tea cultivars as revealed by RFLP analysis of phenylalanine ammonia-lyase DNA. Theor Appl Genet 104:998–1002
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Ohsako T, Ohgushi T, Motosugi H, Oka K (2008) Microsatellite variability within and among local landrace populations of tea, Camellia sinensis (L.) O. Kuntze, in Kyoto, Japan. Genet Resour Crop Evol 55:1047–1053
Peakall R, Smouse P (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 82:2537–2539
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Raina SN, Ahuja PS, Sharma RK, Das SC, Bhardwaj P, Negi R, Sharma V, Singh SS, Sud RK, Kalia RK et al (2012) Genetic structure and diversity of India hybrid tea. Genet Resour Crop Evol 59:1527–1541
Rosenberg NA (2004) Distruct: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Takeda Y (2002) Studies on variations in genetic resources of tea in Japan and application to tea breeding. Bull Natl Inst Veg Tea Sci 1:97–180
Takeda Y, Toyao T (1980) Identification and classification of the green tea varieties by the morphological characters of flower organs. Tea Res J 52:1–6
Tanaka J (2012) Japanese tea breeding history and the future perspective. In: Chen L, Apostolides Z, Chen ZM (eds) Global tea breeding. Springer, Berlin, pp 227–239
Tanaka J, Ikeda S (2002) Rapid and efficient DNA extraction method from various plant species using diatomaceous earth and a spin filter. Breed Sci 52:151–155
Tanaka J, Sawai Y (2005) Mat locus control the existence of an aromatic substance methyl anthranilate in tea leaves, derived from clonal strain ‘Shizu-Inzatsu 131’. In: Res. Results of Veg. & Tea Sci., National Agricultural and Bio-oriented Research Organization, Tsukuba, Japan, 75–76 (in Japanese)
Taniguchi F, Yoshida K, Tanaka J, Ogino A (2010) Identification of quantitative trait loci for resistance to anthracnose in the tea cultivar ‘Minamisayaka’ using an SSR-based linkage map. In: Proceedings of 2010 International Conference on O-CHA (tea) Culture and Science (Session II). Shizuoka, Japan, pr-p-04
Taniguchi F, Furukawa K, Ohta-Metoku S, Yamaguchi N, Ujihara T, Kono I, Fukuoka H, Tanaka J (2012) Construction of a high-density reference linkage map of tea (Camellia sinensis). Breed Sci 62:263–273
Toyao T, Takeda Y (1999) Studies on geographical diversity of floral morphology of tea plant (Camellia sinensis (L.) O. Kuntze) using the method of numerical taxonomy. Tea Res J 87:39–57
Vigouroux Y, Glaubitz JC, Matsuoka Y, Goodman MM, Sánchez GJ, Doebley J (2008) Population structure and genetic diversity of new world maize races assessed by DNA microsatellites. Am J Bot 95:1240–1253
Wachira FN, Tanaka J, Takeda Y (2001) Genetic variation and differentiation in tea germplasm revealed by RAPD and AFLP variation. J Hortic Sci Biotechnol 76:557–563
Wang XC, Chen L, Yang Y (2011) Establishment of core collection for Chinese tea germplasm based on cultivated region grouping and phenotypic data. Front Agric China 5:344–350
Yao MZ, Chen L, Liang YR (2008) Genetic diversity among tea cultivars from China, Japan and Kenya revealed by ISSR markers and its implication for parental selection in tea breeding programs. Plant Breed 127:166–172
Yao MZ, Ma CL, Qiao TT, Jin JQ, Chen L (2012) Diversity distribution and population structure of tea germplasms in China revealed by EST-SSR markers. Tree Genet Genomes 8:205–220
Acknowledgments
We thank R. Ohsawa at University of Tsukuba and N. Aoki at National Agriculture and Food Research Organization for their helpful comments on the manuscript. We also thank Y. Kuramae and M. Iwata at NARO Institute of Vegetable and Tea Science for their technical assistance.
Data archiving statement
We followed standard Tree Genetics and Genomes policy. All genotype data are deposited in the Dryad Repository doi:10.5061/dryad.5jn04.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Y. Tsumura
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
(XLSX 34 kb)
Supplementary Table 2
(XLSX 10 kb)
Supplementary Table 3
(DOCX 21 kb)
Supplementary Table 4
(DOCX 19 kb)
Supplementary Table 5
(DOCX 67 kb)
Supplementary Fig. 1
DeltaK values and average maximum correlations in the structure result (PPT 159 kb)
Rights and permissions
About this article
Cite this article
Taniguchi, F., Kimura, K., Saba, T. et al. Worldwide core collections of tea (Camellia sinensis) based on SSR markers. Tree Genetics & Genomes 10, 1555–1565 (2014). https://doi.org/10.1007/s11295-014-0779-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-014-0779-0