Abstract
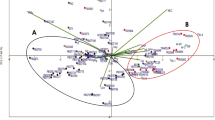
Tea plant [Camellia sinensis (L.) O.Kuntze)] is one of the most popular non-alcoholic beverage crops in the world today. In recent years, many clonal tea cultivars have been developed and widely planted to replace the diverse traditional tea populations. In this article, we study the relationships between classifications based on simple sequence repeat (SSR) markers and on morphological traits for 185 tea plant cultivars. Results show that the genetic diversity index (H) is between 0.229 and 0.803, and the mean value is 0.543; the observed heterozygosity (H o) ranges from 0.103 to 0.683, with an average of 0.340, while the genetic identity varies from 0.267 to 0.984. Based on tea-making properties, the genetic diversity in the “black-green tea” group is much higher than in the “Oolong tea” group. Based on morphological traits, cluster analysis classifies the 185 cultivars into three groups, “group I,” “group II” and “group III.” Most cultivars are related based on the geographical origin and their genetic backgrounds.







Similar content being viewed by others
References
Anderson JA, Churchill GA et al (1993) Optimizing parental selection for genetic linkage maps. Genome 36:181–186
Chen L, Chen D, Gao QK et al (1997) Isolation and appraisal of genomic DNA from tea plant [Camellia sinensis (L.)O.Kuntze]. J Tea Sci 17(2):177–181
Chen L, Yang YJ, Yu FL et al (1998) Gene diversity of 15 tea (Camellia sinensis (L.) O. Kuntze) cultivars using RAPD markers. J Tea Sci 18(1):12–27
Chen L, Gao QK, Chen DM et al (2005) The use of RAPD markers for detecting genetic diversity, relationship and molecular identification of Chinese elite tea genetic resources [Camellia sinensis (L.) O. Kuntze] preserved in tea germplasm repository. Biodivers Conserv 14:1433–1444
Chen L, Yao MZ, Zhao LP et al (2006) Recent research progresses on molecular biology of tea plant (Camellia sinensis). Floriculture, ornamental and plant biotechnology: advances and topical issues. Lond Glob Sci Books 4:425–436
Huang FP, Liang YR, Lu JL et al (2004) Evaluation of genetic diversity in oolong tea germplasms by AFLP fingerprinting. J Tea Sci 24(3):183–189
Huang M, Xie FM, Chen LY et al (2010) Comparative analysis of genetic diversity and structure in rice using ILP and SSR markers. Rice Sci 17(4):257–268
Jakše J, Martin W, McCallum J, Havey MJ (2005) Single nucleotide polymorphisms, indels, and simple sequence repeats for onion cultivar identification. J Am Soc Hort Sci 130:912–917
Labate JA, Lamkey KR, Mitchell SE et al (2003) Molecular and historical aspects of corn belt dent diversity. Crop Sci 43:80–91
Langella O. (2007) Populations 1.2.30: population genetic software (individuals or population distances, phylogenetic trees). http://bioinformatics.org/˜tryphon/populations/
Li YH, Smulders MJM, Chang RZ et al (2010) Analysis of SSRs uncovers hierarchical structure and genetic diversity in Chinese soybean landraces. Agric Sci Chin 9(12):1739–1748
Liu KJ, Muse SV (2005) Power marker: integrated analysis environment for genetic marker data. Bioinformatics 21(9):2121–2129
Luo JW, Shi ZP, Shen CW (2002) Studies on genetic relationships of tea cultivars [Camellia sinensis (L.) O. Kuntze] by RAPD analysis. J Tea Sci 22(2):140–146
Ma JQ, Zhou YH, Ma CL et al (2010) Identification and characterization of 74 novel polymorphic EST-SSR markers in the tea plant, Camellia sinensis (theaceae). Am J Bot 97(12):E153
Meng FJ, Xu XY, Huang FL (2010) Analysis of genetic diversity in cultivated and wild tomato varieties in chinese market by RAPD and SSR. Agric Sci Chin 9(10):1430–1437
Nei M (1972) Genetic distance between populations. Am Nat 106:283–292
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nybom H (2004) Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Mol Ecol 13:1143–1155
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Ming TL (1999) A systematic synopsis of the genus Camellia. Acta Botanica Yunanica 21(2):149–159
Tsukazaki H, Yamashita K, Yaguchi S, Masuzaki S et al (2008) Construction of SSR-based chromosome map in bunching onion (Allium fistulosum). Theor Appl Genet 117:1213–1223
Wang LY, Liu BY, Jiang YH et. al (2009) Phylogenetic analysis of interspecies in section Thea through SSR markers. J Tea Sci 29(5):341–346
Zhao LP, Liu Z, Chen L et al (2008) Generation and characterization of 24 novel EST derived microsatellites from tea plant (Camellia sinensis) and cross-species amplification in its closely related species and varieties. Conserv Genet 9:1327–1331
Acknowledgments
This study was supported by the earmarked fund for the Modern Agro-industry Technology Research System of China, the Priority Academic Program Development of Jiangsu Higher Education Institutions of China, National Natural Science Foundation of China (30800884, 30972403), Jiangsu Science and Technology Program of China (BE2009313-1, BE2011319), and the Suzhou Science and Technology Program of China (SZGD201067, WNZ1002).
Author information
Authors and Affiliations
Corresponding authors
Appendix
Appendix
See Table 4.
Rights and permissions
About this article
Cite this article
Fang, W., Cheng, H., Duan, Y. et al. Genetic diversity and relationship of clonal tea (Camellia sinensis) cultivars in China as revealed by SSR markers. Plant Syst Evol 298, 469–483 (2012). https://doi.org/10.1007/s00606-011-0559-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-011-0559-3