Summary
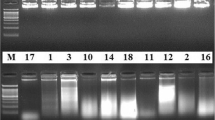
The genetic diversity of microsymbiont Frankia from Colletia hystrix (Clos.) plants growing in a restricted area, were investigated by PCR-restriction fragment length polymorphism (RFLP) technique. DNA from field-collected nodules was amplified by PCR with primers targeting two genomic regions; one included a large portion of the 3′ end of the 16S rRNA gene, the intergenic spacer, and the 5′ end of the 23S rRNA gene and the other in the nifD–nifK intergenic region in the nif operon as a means to estimate molecular diversity. A HaeIII digestion of the PCR product allowed us to identify PCR-RFLP groups or haplotypes among the Colletia-infective Frankia strains tested. An exhaustive small-scale sampling permitted us to detect haplotypes with a low frequency in the microsymbiont population and showed that Frankia microsymbionts have a higher genetic diversity than previously reported. Fifteen haplotypes were recognized on the basis of combining the restriction patterns in each region analyzed. The haplotype designated as A3 was found with a high frequency in the five microsymbiont Frankia groups studied indicating a dominant haplotype. This haplotype was also exhibited by strain ChI4, which was isolated in 1991 in the same locality suggesting that it is the most common haplotype in this area and very stable over time.
Similar content being viewed by others
References
Baker D.D., Schwintzer C.R., 1990 Actinorhizal symbioses In Stacey G., Burris R.H., Evans H.J., eds Biological Nitrogen Fixation Chapman and Hall, New York. 259–292. ISBN 0-412-02421-7
Beattie G.A., Clayton M.K., Handelsman J., 1989 Quantitative comparison of the laboratory and field Competitiveness of Rhizobium leguminosarum biovar phaseoli Applied and Environmental Microbiology 55: 2755–2761
Benson D.R., Clawson M.L., 2000 Evolution of the actinorhizal plants symbiosis In Triplett E.W., ed. Prokaryotic Nitrogen Fixation: A Model System for Analysis of a Biological Process Horizon Scientific Press Madison–Wisconsin 207–224 ISBN 1-898486-19-0
Benson D.R., Silvester W.B., 1993 Biology of Frankia strains, actinomycete symbionts of actinorhizal plants Microbiological Reviews 57: 293–319
Benson D.R., Buchholz. S.E., Hanna D.G., 1984 Identification of Frankia strains by two-dimensional polyacrylamide gel electrophoresis Applied and Environmental Microbiology 47: 489–494
Bosco M., Lumini E., Normand P., 1996 PCR-RFLP direct fingerprinting of culture Frankia microsymbiont from Dryas drummondii nodules Annali di Microbiologia ed Enzi mologia 46: 115–123
Bravo, L.A. 2002 Diversidad genética y especificidad de huésped de las cepas de Frankia aisladas de la familia Rhamnaceae. MSc. Thesis, Universidad de Chile Santiago,Chile
Callaham D., Newcomb W., Torrey J.G., 1978 Isolation and cultivation in vitro of the actinomycete causing root nodulation in Comptonia Science 199: 899–902
Carú M., Mosquera G., Bravo L., Guevara R., Sepúlveda D., Cabello A., 2003 Infectivity and effectivity of Frankia strains from the Rhamnaceae family on different actinorhizal plants Plant and Soil 251: 219–225
Carú M., 1993 Characterization of native Frankia strains isolated from Chilean shrubs (Rhamnaceae) Plant and Soil 157: 137–145
Clawson M.L., Benson D.R., 1999 Dominance of Frankia strains in stands of Alnus incana subsp. rugosa (Du Roi) R.T.Clausen and Myrica pensylvanica Loisel Canadian Journal of Botany 77: 1203–1207
Clawson M.L., Carú M., Benson D.R., 1998 Diversity of Frankia strains in root nodules of plants from the families Elaeagnaceae and Rhamnaceae Applied and Environmental Microbiology 64: 3539–3543
Dawson J.O., 1990 Interactions among actinorhizal and associated plant species In Schwintzer C.R., Tjepkema J.D., eds. The Biology of Frankia and actinorhizal plants Academic Press, Inc. San Diego 299–316 ISBN 0–12–633210-X
Franco A.A., Vincent J.M., 1976. Competition amongst rhizobial strains for the colonization and nodulation of two tropical legumes Plant and Soil 45: 27–48
George M.L., Robert F.M., 1992 Competition among Rhizobium leguminosarum bv. phaseoli strains for nodulation of common bean Canadian Journal of Microbiology 38: 157–160
Hartmann A., Giraud J.J., Catroux C., 1998 Genotypic diversity of Sinorhizobium (formely Rhizobium) meliloti strains isolated directly from a soil and from nodules of alfalfa (Medicago sativa) grown in the same soil FEMS Microbiology Ecology 25: 107–116
Jamann S., Fernández M.P., Normand P., 1993 Typing method for N2-fixing bacteria based on PCR-RFLP application to the characterization of Frankia strains Molecular Ecology 2: 17–26
Jeong S.C., Myrold D.D., 1999 Genomic fingerprinting of Frankia microsymbionts from Ceanothus copopulations using repetitive sequences and polymerase chain reactions Canadian Journal of Botany 77: 1220–1230
Labandera C.A., Vincent J.M., 1975. Competition between an introduction strain and native Uruguayan strains of Rhizobium trifolii Plant and Soil 42: 327–347
Laguerre G., Louvrier P., Allard M.R., Amarger N., 2003 Compatibility of Rhizobial genotypes within natural populations of Rhizobium leguminosarum biovar viciae for nodulation of host legumes Applied and Environmental Microbiology 69: 2276–2283
Lumini E., Bosco M., 1996 PCR-restriction fragment length polymorphism identification and host range of single spore isolates of the flexible Frankia sp. Strain UFI 132715 Applied and Environmental Microbiology 62: 3026–3029
Maggia L., Nazaret S., Simonet P., 1992 Molecular characterization of Frankia isolates from Casuarina equisetifolia L. root nodules harvest in West Africa (Senegal and Gambia) Acta Ecologica 13: 453–461
Moawad H.A., Ellis W.R., Schmidt E.L., 1984 Rhizosphere response as a factor in competition among three serogroups of indigenous Rhizobium japonicum for nodulation of field-grown soybeans Applied and Environmental Microbiology 47: 607–612
Mosquera, G.M. 2002 Especificidad de huésped y variabilidad genética de cepas de Frankia aisladas de nódulos de radiculares de las plantas de la familia Rhamnaceae, detectada mediante análisis de marcadores moleculares RAPDs y PCR-RFLP. MSc thesis, Universidad de Chile Santiago, Chile
Murry M.A., Fontaine M.S., Torrey J.G., 1984 Growth kinetics and nitrogenase induction in Frankia sp. HFPArI3 grown in batch culture Plant and Soil 78: 61–78
Murry M.A., Konopka. A.S., Pratt S.D., Vandergon T.L., 1997 The use of PCR-based typing methods to assess the diversity of Frankia nodule endophytes of the actinorhizal shrub Ceanothus Physiologia Plantarum 99: 714–721
Nalin R., Domenach A.M., Normand P., 1995 Molecular structure of the Frankia spp. nifD-K intergenic spacer and design of Frankia genus compatible primers Molecular Ecology 4: 483–491
Nalin R., Normand P., Domenach. A., 1997 Distribution and N2-fixing activity of Frankia strains in relation to soil depth Physiologia Plantarum 99: 732–738
Nei M., 1972 Genetic distance between populations American Naturalist 106: 283–292
Ochman H., Selander R.K., 1984 Standard reference strains of Escherichia coli from natural populations Journal of Bacteriology 157: 690–693
Redell P., Bowen G.D., 1985 Do single nodules of Casuarina contain more than one Frankia strains? Plant and Soil 88: 275–279
Ritchie N.J., Myrold D.D., 1999 Geographic distribution and genetic diversity of Ceanothus-infective Frankia strains Applied and Environmental Microbiology 65: 1378–1383
Rius N., Fusté M.C., Guasp C., Lalucat J., Lorén J.G., 2001 Clonal population structure of Pseudomonas stutzeri, a species with exceptional genetic diversity Journal of Bacteriology 183: 736–744
Rouvier C., Prin Y., Redell P., Normand P., Simonet P., 1996 Genetic diversity among Frankia strains modulating members of the family Casuarinaceae in Australia revealed by PCR and restriction fragment length polymorphism analysis with crushed root nodules Applied and Environmental Microbiology 62: 979–985
Simonet P., Bosco M., Chapelon C., Moiroud A., Normand P., 1994 Molecular characterization of Frankia microsymbionts from spore-positive and spore-negative nodules in a natural alder stand Applied and Environmental Microbiology 60: 1335–1341
Simonet P., Navarro E., Rouvier C., Reddell P., Zimpfer J., Dommergues Y., Bardin R., Combarro P., Hamelin J., Domenach A.M., Gourbière F., Prin Y., Dawson J., Normand P., 1999 Co-evolution between Frankia populations and host plants in the family Casuarinaceae and consequent patterns of global dispersal Environmental Microbiology 1: 525–533
Souza V., Rocha M., Valera A., Eguiarte L., 1999 Genetic structure of natural populations of Escherichia coli in wild hosts on different continents Applied and Environmental Microbiology 65: 3373–3385
Velásquez E.P., Mateos F., Velasco N., Santos F., Burgos P.A., Villadas P., Toro N., Martínez-Molina E., 1999 Symbiotic characteristics and selection of autochthonous strains of Sinorhizobium meliloti populations in different soils Soil Biology and Biochemistry 31: 1039–1047
Weisburg W.G., Barns S.M., Pelletier D.A., Lane D.J., 1991 16S ribosomal DNA amplification for phylogenetic study Journal of Bacteriology 173: 697–703
Acknowledgements
This work was supported by Fondecyt project No. 1040880 and a Grant ENL-2001/02 of the Departamento de Investigación y Desarrollo (DID) of the Universidad de Chile. The authors thank L. Bravo for critical reading and correcting the English text.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chávez, M., Carú, M. Genetic Diversity of Frankia Microsymbionts in Root Nodules from Colletia hystrix (Clos.) Plants by Sampling at a Small-Scale . World J Microbiol Biotechnol 22, 813–820 (2006). https://doi.org/10.1007/s11274-005-9108-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-005-9108-y