Abstract
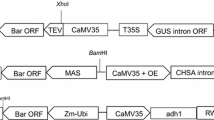
We report an Agrobacterium-mediated transformation system that can generate marker-free transgenic sorghum [Sorghum bicolor (L.) Moench] from a public line [P898012] using standard binary vectors with bar as a selectable marker. Eight co-cultivation conditions were examined for their effect on transformation. The average transformation frequencies were 0.4 and 0.7% for pZY101-TC2 and pZY101-SKRS, respectively, derived from binary vector pZY102 and containing bar and target gene(s) in separate T-DNA regions. A low selection pressure (2.5 mg l−1 DL-phosphinothrithin, PPT) was deployed during callus induction in combination with rapid selection to generate plants from 80 independent events, all but three of which were fertile and set seed. PCR and Southern analyses showed that 36 out of 80 events contained both bar and the target gene(s) (an average co-transformation frequency of 45%). Seedlings of the T1 generation transmitted T-DNAs with target gene(s) and bar gene independently, generating a fraction of progeny with only the target gene(s).




Similar content being viewed by others
References
Able JA (1998) Transformation of sorghum using the particle inflow gun (PIG). Int Sorghum Millets Newsl 39:98–100
Able JA, Rathus C, Godwin ID (2001) The investigation of optimal bombardment parameters for transient and stable transgene expression in sorghum. In Vitro Cell Dev Biol-Plant 37:341–348
Cai T, Pierce DA, Tagliani LA, Zhao ZY (2002) Agrobacterium mediated transformation of sorghum. US Patent 6369298
Carvalho CHS, Zehr UB, Gunaratna N, Anderson JM, Kononowicz HH, Hodges TK, Axtell JD (2004) Agrobacterium-mediated transformation of sorghum: factors that affect transformation efficiency. Genet Mol Biol 27:259–269
Casas AM, Kononowicz AK, Zehr UB, Tomes DT, Axtell JD, Butler LG, Bressan RA, Hasegawa PM (1993) Transgenic plants via microprojectile bombardment. Proc Natl Acad Sci USA 90:11212–11216
Casas AM, Kononowicz AK, Hann TG, Zhang L, Tomes DT, Bressan RA, Hasegawa PM (1997) Transgenic plants obtained after microprojectile bombardment of immature inflorescence. In Vitro Cell Dev Biol-Plant 33:92–100
Chilton MD, Currier TC, Farrand SK, Bendich AJ, Gordon MP, Nester EW (1974) Agrobacterium tumefaciens DNA and PS8 bacteriophage DNA not detected in crown gall tumors. Proc Natl Acad Sci USA 71:3672–3676
Dellaporta SL, Wood J, Hicks JB (1983) A plant DNA minipreparation: version 2. Plant Mol Biol Rep 1:19–22
Emani C, Sunilkumar G, Rathore KS (2002) Transgene silencing and reactivation in sorghum. Plant Sci 162:181–192
Folk WR (2003) How to improve plant protein quality: err on the side of goodness. ISB News Rep (http://www.isb.vt.edu)
Frisch DA, Harris-Haller LW, Yokubaitis NT, Thomas TL, Hardin SH, Hall TC (1995) Complete sequence of the binary vector Bin 19. Plant Mol Biol 27:405–409
Gao ZS, Jararaj J, Muthukrishnan S, Claflin L, Liang GH (2005a) Efficient genetic transformation of Sorghum using a visual screening marker. Genome 48:321–333
Gao ZS, Xie X, Ling Y, Muthukrishnan S, Liang GH (2005b) Agrobacterium tumefaciens-mediated sorghum transformation using a mannose selection system. Plant Biotechnol J 3:591–599
Girijashankar V, Sharma HC, Sharma KK, Swathisree V, Prasad LS, Bhat BV, Royer M, Secundo BS, Narasu ML, Altosaar I, Seetharama N (2005) Development of transgenic sorghum for insect resistance against the spotted stem borer (Chilo partellus). Plant Cell Rep 24:513–522
Gurel S, Gurel E, Kaur R, Wong J, Meng L, Tan HQ, Lemaux PG (2009) Efficient, reproducible Agrobacterium-mediated transformation of sorghum using heat treatment of immature embryos. Plant Cell Rep 28:429–444
Hood EE, Helmer GL, Fraley RT, Chilton MD (1986) The hypervirulence of Agrobacterium tumefaciens A281 is encoded in a region of pTiBo542 outside of T-DNA. J Bacteriol 168:1291–1301
Howe A, Sato S, Dweikat I, Fromm M, Clemente TE (2006) Rapid and reproducible Agrobacterium-mediated transformation of sorghum. Plant Cell Rep 25:784–791
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Jeoung JM, Krishnaveni S, Muthukrishnan S, Trick HN, Liang GH (2002) Optimization of sorghum transformation parameters using genes for green fluorescent protein and β-glucuronidase as visual markers. Hereditas 137:20–28
Lee BK, Kennon AR, Chen X, Jung TW, Ahn BO, Lee JY, Zhang Z (2007) Recovery of transgenic events from two highly recalcitrant maize (Zea mays L.) genotypes using Agrobacterium-mediated standard-binary-vector transformation. Maydica 52:457–469
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Nguyen TV, Thu TT, Claeys M, Angenon G (2007) Agrobacterium-mediated transformation of sorghum using an improved in vitro regeneration system. Plant Cell Tiss Organ Cult 91:155–164
Southern EM (1975) Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol 98:503–517
Tadesse Y, Sagi L, Swennen R, Jacobs M (2003) Optimization of transformation condition and production of transgenic sorghum (Sorghum bicolor) via microparticle bombardment. Plant Cell Tissue Organ Cult 75:1–18
Vega J, Yu W, Kennon A, Chen X, Zhang Z (2008) Improvement of Agrobacterium-mediated transformation in Hi-II maize (Zea mays L.) using standard binary vectors. Plant Cell Rep 27:297–305
Wu XR, Kenzior A, Wilmot D, Scanlon S, Chen Z, Topin A, He S, Acevedo A, Folk WR (2007) Altered expression of plant lysyl tRNA synthetase promotes tRNA misacylation and translational recoding of lysine. Plant J 50:627–636
Zeng P, Vadnais D, Zhang Z, Polacco J (2004) Refined glufosinate selection in Agrobacterium-mediated transformation of soybean [Glycine max (L.) Merr.]. Plant Cell Rep 22:478–482
Zhao ZY, Cai T, Tagliani L, Miller M, Wang N, Pang H, Rudert M, Schroeder S, Hondred D, Seltzer J, Pierce D (2000) Agrobacterium-mediated sorghum transformation. Plant Mol Biol 44:789–798
Zhao ZY, Glassman K, Sewalt V, Wang N, Miller M, Chang S, Thompson T, Catron S, Wu E, Bidney D, Kedebe Y, Jung R (2003) Nutritionally improved transgenic sorghum. In: Vasil IK (ed) Plant biotechnology 2002 and beyond. Proceedings of the 10th IAPTC & B Congress. Kluwer Academic Publishers, pp 413–416
Zhu H, Muthukrishana S, Krishnaveni S, Jeoung JM, Liang GH (1998) Biolistic transformation of sorghum using a rice chitinase gene. J Genet Breed 52:243–252
Acknowledgments
The authors thank: Scanlon S and other colleagues from Dr. Folk’s lab for their assistance; Z. Zhao, N. Wang, S. Zheng and H. Cline (Pioneer Hi-Bred Intl. Inc.) for helpful suggestions and for P898012 and Drs. G. Liang and Z. S. Gao (Kansas State University) for helpful comments; Aventis CropScience (Research Triangle Park, NC) for providing glufosinate-ammonium and Liberty® as generous gifts; Dr. Seth D. Findley (University of Missouri, Columbia, MO) for critical review of the manuscript. All sorghum transformation experiments were conducted in the Plant Transformation Core Facility at the University of Missouri. Financial support was provided by the University of Missouri Agricultural Experiment Station, the Provost’s office and the University of Missouri Food for the twenty-first Century Eminence Program.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lu, L., Wu, X., Yin, X. et al. Development of marker-free transgenic sorghum [Sorghum bicolor (L.) Moench] using standard binary vectors with bar as a selectable marker. Plant Cell Tiss Organ Cult 99, 97–108 (2009). https://doi.org/10.1007/s11240-009-9580-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-009-9580-4