Abstract
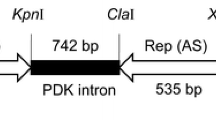
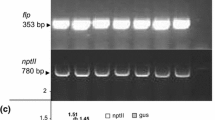
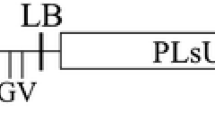
Grapevine Fanleaf virus (GFLV) is one of the most damaging and widespread nepovirus in grapevine, causing the fanleaf disease. Here, we report the development of inverted repeat (IR) constructs, consisting of GFLV-derived sequences, for genetic transformation of grapevine to induce GFLV-specific silencing. The IR construct was designed with fragments of a conserved region of the GFLV movement protein (MPc) gene from a Tunisian isolate (GFLV tun) and with cassettes containing the neomycin phosphotransferase or bar gene as selectable marker. For proof of concept, the IR construct was transferred into Nicotiana benthamiana by Agrobacterium mediated transformation. Challenge inoculation of T1 transgenic plant lines with the relevant virus resulted in plants showing resistance, recovery, retarded infection and susceptibility. Northern blot analysis, using a virus-derived sequence (MPc) as probe, could detect transgene specific small interfering RNA in resistant transgenic N. benthamiana plants after GFLV inoculation. Embryogenic cells of grapevine (Vitis vinifera cv Arich dressé) were transformed with the IR construct in combination with the bar gene using A. tumefaciens strain LBA4404. After molecular characterization of regenerated putative transgenic grapevine plants, Northern blot analysis detected different levels of IR transcripts in independent transgenic grapevine lines indicating a post-transcriptional gene silencing.







Similar content being viewed by others
Abbreviations
- dsRNA:
-
Double stranded RNA
- GFLV:
-
Grapevine Fanleaf virus
- IR:
-
Inverted repeat
- PCR:
-
Polymerase chain reaction
- PTGS:
-
Post-transcriptional gene silencing
- RT–PCR:
-
Reverse transcription–PCR
- si RNA:
-
Small interfering RNA
References
Bardonnet N, Hans F, Serghini L, Pinck L (1994) Protection against virus infection in tobacco plants expressing the coat protein of grapevine fanleaf virus. Plant Cell Rep 13:357–360. doi:10.1007/BF00232637
Baulcombe D (2004) RNA silencing in plants. Nature 431:356–363. doi:10.1038/nature02874
Bleys A, Van Houdt H, Depicker A (2006) Down regulation of endogenes mediated by a transitive silencing signal. RNA 12:1633–1639. doi:10.1261/rna.108106
Bonfim K, Faria JC, Nogueira EOPL, Rica Mendes ÉA, Aragão FJL (2007) RNAi-mediated resistance to bean golden mosaic virus in genetically engineered common bean (Phaseolus vulgaris) using an intron hairpin construction. Mol Plant Microbe Interact 20(6):717–726. doi:10.1094/MPMI-20-6-0717
Bouamama B, Jardak R, Ben Salem A, Ghorbel A, Mliki A (2009) Preservation of endangered Tunisian grapevine cultivars using embryogenic cultures. Electron J Biotechnol 12(2)
Bucher E, Lohuis D, van Poppel PMJA, Geerts-Dimitriadou C, Goldbach R, Prins M (2006) Multiple virus resistance at a high frequency using a single transgene construct. J Gen Virol 87:3697–3701. doi:10.1099/vir.0.82276-0
Chen YK, Lohuis D, Goldbach R, Prins M (2004) High frequency introduction of RNA mediated resistance against cucumber mosaic virus using inverted repeat constructs. Mol Breed 14:215–226. doi:10.1023/B:MOLB.0000047769.82881.f5
Di Nicolas-Negri E, Brunetti A, Tavazza M, Ilardi V (2005) Hairpin RNA-mediated silencing of plum pox virus P1 and HC-Pro genes for efficient and predictable resistance to the virus. Transgenic Res 14(6):989–994. doi:10.1007/s11248-005-1773-y
Ecke W, Schmitz W, Michaelis G (1990) The mitochondrial nad5 gene of sugar beet (Beta vulgaris) encoding a subunit of the respiratory NADH dehydrogenase. Curr Genet 18:133–139. doi:10.1007/BF00312601
Edwards K, Johnstone C, Thompson C (1991) A simple and rapid method for the preparation of plant genomic DNA for PCR analysis. Nucleic Acids Res 19(6):1349. doi:10.1093/nar/19.6.1349
Filichkin SA, DiFazio SP, Brunner AM, Davis JM, Yang ZK, Kalluri UC, Arias RS, Etherington E, Tuskan GA, Strauss SH (2007) Efficiency of gene silencing in Arabidopsis: direct inverted repeats vs. transitive RNAi vectors. Plant Biotechnol J 5:615–626. doi:10.1111/j.1467-7652.2007.00267.x
Fuchs M, Cambra M, Capote N, Jelkmann W, Kundu J, Laval V, Martelli GP, Minafra A, Petrovic N, Pfeiffer P, Pompe Novak M, Ravelonandro M, Saldarelli P, Stussi-Garaud C, Vigne E, Zagrai I (2007) Safety assessment of transgenic plums and grapevines expressing viral coat protein genes: new insights into real environmental impact of perennial plants engineered for virus resistance. J Plant Pathol 89(1):5–12
Furutani N, Yamagishi N, Hidaka S, Shizukawa Y, Kanematsu S, Kosaka Y (2007) Soybean mosaic virus resistance in transgenic soybean caused by post-transcriptional gene silencing. Breed Sci 57:123–128. doi:10.1270/jsbbs.57.123
Gambino G, Gribaudo I, Leopold S, Schartl A, Laimer M (2005) Molecular characterization of grapevine plants transformed with GFLV resistance genes: I. Plant Cell Rep 24:655–662. doi:10.1007/s00299-005-0006-4
Hajdukiewicz P, Zvab Z, Maliga P (1994) The small, versatile pPZP family of Agrobacterium binary vector for plant transformation. Plant Mol Biol 25:989–994. doi:10.1007/BF00014672
Hily JM, Scorza R, Ravelonandro M, Damsteegt V, Bassett C, Petri C, Lui Z (2007) Plum pox virus coat protein gene intron-hairpin-RNA (ihpRNA) constructs provide resistance to plum pox virus in Nicotiana benthamiana and Prunus domestica. J Am Soc Hortic Sci 132(6):580–858
Himber C, Dunoyer P, Moissiard G, Ritzenthaler C, Voinnet O (2003) Transitivity dependent cell to cell movement of RNA silencing. EMBO J 22(17):4523–4533. doi:10.1093/emboj/cdg431
Horsch RB, Hofmann NF, Eichholtz D, Roger SG, Fraley RT (1985) A simple and general method for transferring genes into plants. Science 227:1229–1231. doi:10.1126/science.227.4691.1229
Jardak-Jamoussi R, Bouamama B, Mliki A, Ghorbel A, Reustle GM (2008) The use of phosphinothricin resistance as selectable marker for genetic transformation of grapevine. Vitis 47(1):35–37
Johansen EI (1996) Intron insertion facilitates amplification of cloned virus cDNA in Escherichia coli while biological activity is the re-established after transcription in vivo. Proc Natl Acad Sci USA 93:12400–12405. doi:10.1073/pnas.93.22.12400
Kalantidis K, Psaradakis S, Tabler M, Tsargis M (2002) The occurrence of CMV-Specific RNAs in transgenic tobacco expressing virus derived double stranded RNA is indicative of resistance to the virus. Mol Plant Microbe Interact 15(8):826–833. doi:10.1094/MPMI.2002.15.8.826
Krastanova S, Perrin M, Barbier P, Demangeat G, Cornuet P, Bardonnet N, Otten L, Pinck L, Walter B (1995) Transformation of grapevine rootstocks with the coat protein gene of grapevine fanleaf nepovirus. Plant Cell Rep 14:550–554. doi:10.1007/BF00231936
Lechtenberg B, Schubert D, Forsbach A, Gils M, Schmidt R (2003) NT-DNA configurations nor arrangements of tandemly repeated transgenes are sufficient to trigger transgene silencing. Plant J 34:507–517. doi:10.1046/j.1365-313X.2003.01746.x
Lipardi C, Wei Q, Paterson BM (2001) RNAi as random degradative PCR: siRNA primers convert mRNA into dsRNAs that are degraded to generate new siRNAs. Cell 107:297–307. doi:10.1016/S0092-8674(01)00537-2
Lodhi MA, Ye JN, Weeden NF, Reisch BI (1994) A simple and efficient method for DNA extraction from grapevine cultivars and Vitis species. PMB Rep 12(1):6–13
Lui HM, Zhu CX, Zhu XP, Guo X, Song YZ, Wen FJ (2007) A link between PVYN CP gene-mediated virus resistance and transgene arrangement. J Phytopathol 155:676–682
Maghuly F, Leopold S, da Câmara Machado A, Borroto Fernandez E, Ali Khan M, Gambino G, Gribaudo I, Schartl A, Laimer M (2006) Molecular characterization of grapevine plants transformed with GFLV resistance genes: II. Plant Cell Rep 25(6):546–553. doi:10.1007/s00299-005-0087-0
Marchuk D, Drumm M, Saulino A, Collins FS (1991) Construction of T-vectors, a rapid and general system for direct cloning of unmodified PCR products. Nucleic Acids Res 19(5):1154. doi:10.1093/nar/19.5.1154
Missiou A, Kalantidis K, Boutla A, Tzortzakaki S, Tabler M, Tsagris M (2004) Generation of transgenic potato plants highly resistant to potato virus Y (PVY) through RNA silencing. Mol Breed 14:185–197. doi:10.1023/B:MOLB.0000038006.32812.52
Mitter N, Mitchell R, Dietzgen RG (2006) Fate of hairpin transcript components during RNA silencing and its suppression in transgenic virus-resistant tobacco. J Biotechnol 126(2):115–122. doi:10.1016/j.jbiotec.2006.03.036
Ramesh SV, Mishra AK, Praveen S (2007) Hairpin RNA-mediated strategies for silencing of tomato leaf curl virus AC1 and AC4 genes for effective resistance in plants. Oligonucleotides 17:251–257. doi:10.1089/oli.2006.0063
Renault AS, Deloire A, Letinois I, Kraeva E, Tesniere C, Ageorges A, Redon C, Brierne J (2000) β-1, 3-glucanase gene expression in grapevine leaves as a response to infection with Botrytis cinerea. Am J Enol Vitic 5:81–87
Sijen T, Fleenor J, Simmer F, Thijssen KL, Parrish S, Timmons L, Plasterk RHA, Fire A (2001) On the role of RNA amplification in dsRNA triggered gene silencing. Cell 107:465–476. doi:10.1016/S0092-8674(01)00576-1
Smith NA, Singh SP, Wang MB, Stoutjesdijk PA, Green AJ, Waterhouse PM (2000) Total silencing by intron spliced hairpin RNAs. Nature 407:319–320. doi:10.1038/35036500
Stam M, Mol JNM, Kooter JM (1997) The silence of genes in transgenic plants. Ann Bot (Lond) 79:3–12. doi:10.1006/anbo.1996.0295
Szittya G, Molnar A, Sihavy D, Hornyik C, Burgyan J (2002) Short defective interfering RNAs of tombusviruses are not targeted but trigger post-transcriptional gene silencing against their helper virus. Plant Cell 14:359–372
Tenllado F, Barajas D, Vargas M, Atencio FA, Gonzalez-Jara P, Diaz-Ruiz JR (2003) Transient expression of homologous hairpin RNA causes interference with plant virus infection and is overcome by a virus encoded suppressor of gene silencing. Mol Plant Microbe Interact 16(2):149–158. doi:10.1094/MPMI.2003.16.2.149
Thomas MR, Matsumoto S, Cain P, Scott NS (1993) Repetitive DNA of grapevine: classes present and sequences suitable for cultivar identification. Theor Appl Genet 86:173–180
Tougou M, Furutani N, Yamagishi N, Shizukawa Y, Takahata Y, Hidaka S (2006) Development of resistant transgenic soybeans with inverted repeat-coat protein genes of soybean dwarf virus. Plant Cell Rep 25(11):1213–1218. doi:10.1007/s00299-006-0186-6
Vaistij FE, Jones L, Baulcombe DC (2002) Spreading of RNA targeting and DNA methylation in RNA silencing requires transcription of the target gene and a putative RNA-dependent RNA polymerase. Plant Cell 14:857–867. doi:10.1105/tpc.010480
Valat L, Fuchs M, Burrus M (2006) Transgenic grapevine rootstock clones expressing the coat protein or movement protein genes of grapevine fanleaf virus: characterization and reaction to virus infection upon protoplast electroporation. Plant Sci 170:739–747. doi:10.1016/j.plantsci.2005.11.005
Van Houdt H, Bleys A, Depicker A (2003) RNA target sequences promote spreading of RNA silencing. Plant Physiol 131:245–253. doi:10.1104/pp.009407
Vigne E, Komar V, Fuchs M (2004) Field safety assessment of recombination in transgenic grapevines expressing the coat protein gene of grapevine fanleaf virus. Transgenic Res 13(2):165–179. doi:10.1023/B:TRAG.0000026075.79097.c9
Wang Q, Li P, Hanania Sahar N, Mawassi M, Gafny R, Sela I, Tanne E, Perl A (2000) Improvement of Agrobacterium-mediated transformation efficiency and transgenic plant regeneration of Vitis vinifera L. by optimizing selection regimes and utilizing cryopreserved cell suspensions. Plant Sci 168:565–571
Wang MB, Wesley SV, Finnegan EJ, Smith NA, Waterhouse PM (2001) Replicating satellite RNA induces sequence-specific DNA methylation and truncated transcripts in plants. RNA 7:16–28. doi:10.1017/S1355838201001224
Wetzel T, Jardak R, Meunier L, Ghorbel A, Reustle GM, Krczal G (2002) Simultaneous RT/PCR detection and differentiation of arabis mosaic and grapevine fanleaf nepoviruses in grapevines with a single pair of primers. J Virol Methods 101:63–69. doi:10.1016/S0166-0934(01)00422-0
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jardak-Jamoussi, R., Winterhagen, P., Bouamama, B. et al. Development and evaluation of a GFLV inverted repeat construct for genetic transformation of grapevine. Plant Cell Tiss Organ Cult 97, 187–196 (2009). https://doi.org/10.1007/s11240-009-9514-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-009-9514-1