Abstract
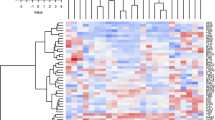
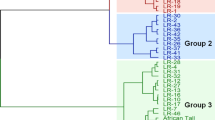
Landraces of maize represent a valuable genetic resource for breeding and genetic studies. Since 1970, landraces have been collected from all over Turkey, but the genetic diversity represented in this collection is still largely unknown. In this study, a sample of 98 landraces sampled from 45 provinces of Turkey was assessed genotypically at 28 simple sequence repeat (SSR) loci and phenotypically for 19 morphological traits. The landraces varied significantly for all the latter traits. A total of 172 SSR alleles were detected, giving a mean of 6.21 alleles per locus. The genetic distance between pairs of landraces ranged from 0.18 to 0.63, with a mean of 0.35. Positive and negative correlation exists among different morphological and agronomic traits. Positive association among different traits showed that improvement of one character may simultaneously improve the other desired trait. Based on UPGMA dendrogram and Neighbor-Net (NNET) analyses from both morphological traits and SSR data, respectively, it is obvious that maize landraces from the same geographical region were often placed in different clusters, indicating that grouping based on genetic parameters was not closely related to the geographic origin. The wide diversity present in Turkish maize landraces could be used as genetic resource in designing maize breeding program for developing new cultivars adapted to different geographic and climatic conditions, and may also contribute to worldwide breeding programs.



Similar content being viewed by others
References
Angelo M, Pinheiro de Carvalho A, Ganança JFT, Abreu I, Sousa NF, Marques dos Santos TM, Vieira MRC, Motto M (2008) Evaluation of maize (Zea mays L.) diversity on the Archipelago of Madeira. Genet Resour Crop Evol 55:221–223
Angioi SA, Rau D, Nanni L, Bellucci E, Papa R, Attene G (2011) The genetic make of the European landraces of the common bean. Plant Genetic Resources: Characterization and Utilization 1–5. doi:10.1017/S1479262111000190
Azar C, Mather DE, Hamilton RI (1997) Maize landraces of the St. Lawrance-Great lakes region of North America. Euphytica 98:141–148
Baloch FS, Kurt C, Arıoğlu H, Özkan H (2010) Assaying of diversity among soybean (Glycin max (L.) merr.) and peanut (Arachis hypogaea L.) genotypes at DNA level. Turk J Agric For 34:285–301
Bang TC, Raji AA, Ingelbrecht IL (2011) A multiplex microsattelite marker kit for diversity assessment for large cassava (Manihot esculenta Crantz) germplasm collection. Plant Mol Biol Rep. doi:10.1007/s11105-010-0273-2
Baraket G, Chatti K, Saddoud O, Abdelkarim AB, Mars M, Trifi M, Hannachi AS (2011) Comparative SSR and AFLP markers for evaluation of genetic diversity and conservation of fig. Ficus carica. Genetic resources in Tunisia. Plant Mol Biol Rep 29:171–184
Beyene Y, Botha AM, Myburg AA (2005) A comparative study of molecular and morphological methods of describing genetic relationships in traditional Ethiopian highland maize. Afr J Biotechnol 4(7):586–595
Beyene Y, Botha AM, Myburg AA (2006) Genetic diversity among traditional Ethiopian highland maize accessions assessed by simple sequence repeat (SSR) markers. Genet Resour Crop Evol 00:1–10
Bogyo BT, Proceddu E, Perrino P (1990) Analysis of sampling strategies for collecting genetic materials. Econ Bot 34:11–86
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of genetic linkage map in man using restricted fragment length polymorphism. Am J Human Genet 23:314–331
Bourguiba H, Krichen L, Audergon JM, Khadari B, Trifi-Farah N (2010) Impact of mapped SSR markers on the genetic diversity of apricot (Prunus armeniaca L.) in Tunisia. Plant Mol Biol Rep 28:578–587
Bracco M, Lia VV, Gottlieb AM, Hernandez JC, Poggio L (2009) Genetic diversity in maize landraces from indigenous settlements of Northeastern Argentina. Genetica 135:39–49
Brandolini A, Brandolini A (2001) Classification of Italian maize (Zea mays L.) germplasm. Plant Genet Resour Newslett 126:1–11
Brown AHD (1978) Isozymes, plant population genetics structure and genetic conservation. Theor Appl Genet 52:145–157
CIMMYT/IBPGRI (1991) Descriptor of maize, Rome
Choukan R, Hossainzadeh A, Ghannadha MR, Talei AR, Mohammadi SA, Warburton ML (2006) Use of SSR data to determine relationships and potential heterotic groupings within medium to late maturing Iranian maize inbred lines. Field Crops Res 95:212–222
Dubreuil P, Rebourg C, Merlino M, Charcosset A (1999) The DNA pooled sampling strategy for estimating RFLP diversity of maize population. Plant Mol Biol Rep 17:123–138
Dubreuil P, Warburton M, Chastanet M, Hoisington D, Charcosset A (2006) More on the introduction of temperate maize into Europe: large scale bulk SSR genotyping and new historical elements. Maydica 51:281–291
Dice LR (1945) Measures of the amount of ecologic association between species. Ecology 26:297–302
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh leaf tissue. Focus 12:13–15
Ege H, Karahocağil P (2001) Yemlık Tahıllar Arpa, Mısır durum ve tahmin 2001/2002 TEAE Yayını No 82, Ankara
Eschholz TW, Peter R, Stamp P, Hund A (2008) Genetic diversity of Swiss maize (Zea mays L) assessed with individual and bulks on agarose gel. Genet Resour Crop Evol 55:971–983
Enoki H, Sato H, Koinuma K (2002) SSR analysis of genetic diversity among maize inbred lines adapted to cold regions of Japan. Theor Appl Genet 104:1270–1277
FAOSTAT (2009) http://faostat.fao.org/
Gouesnard B, Dallard J, Panouille A, Boyat A (1997) Classification of French maize populations based on morphological traits. Agronomie 17:491–498
Harlan JR (1975) Our vanishing genetic resources. Science 188:618–621
Hartings H, Berardo N, Mazzinelli GF, Valoti P, Verderio A, Motto M (2008) Assessment of genetic diversity and relationship among maize (Zea mays L.) Italian landraces by morphological traits and AFLP profiling. Theor Appl Genet 117:831–842
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267
Huh MK, Moon SG (2001) Chlorophyll deficient gene and morphological variations in Korean populations of maize (Zea mays). J Plant Biol 44(3):141–147
Ilarslan R, Kaya Z, Tolun AA, Bretting PK (2001) Genetic variability among Turkish Pop, Flint and Dent corn (Zea mays L. spp. mays) races: enzyme polymorphism. Euphytica 122:171–179
Ilarslan R, Kaya Z, Kandemir I, Bretting PK (2002) Genetic variability among Turkish flint, pop and dent corn (Zea mays L) races. Morphological and agronomic traits. Euphytica 128:173–182
Jambrovic A, Simic D, Ledencan T, Zdunic Z, Brkic I (2008) Genetic diversity among maize (Zea mays L.) inbred lines in Eastern Croatia. Period Biol 110(3):251–255
Janick J, Caneva G (2005) The first images of maize in Europe. Maydica 50:71–80
Jarvis DI, Meyer L, Klemick H, Guarino L, Smale M, Brown AHD, Sadiki M, Sthapit B, Hodgkin T (2000) Training guide for in situ conservation on-farm. Biodiversity International. International Plant Genetic Resources Institute, p 161
Kırtok Y (1998) Mısır Üretimi ve Kullanımı. Kocaoluk Basımı ve Yayınevi (In Turkish)
Kün E (1985) Tahıllar II (Sıcak iklim tahıllar). Ankara Üniversitesi Ziraat Fakultesi Yayınları 953. Ders Kitabı 275. Ankara Üniv. Basımevi, Ankara (In Turkish)
Kün E (1994) Tahıllar II (Sıcak iklim tahıllar). Ankara Üniversitesi Ziraat Fakultesi Yayınları 1360:141–206 (In Turkish)
Laborda PR, Oliveira KM, Garcia AAF, Paterniani MEAGZ, de Souza AP (2005) Tropical maize germplasm: what can we say about its genetic diversity in the light of molecular markers? Theor Appl Genet 111:1288–1299
Legesse BW, Myburg AA, Pixley KV, Botha AM (2007) Genetic diversity of African maize inbred lines revealed by SSR markers. Hereditas 144:10–17
Leng ER, Tavcar A, Trifunovic V (1962) Maize of Southeastern Europe and its potential value in breeding programs elsewhere. Euphytica 11:263–272
Liu YJ, Huang YB, Rong TZ, Tian ML, Yang JP (2005) Comparative analysis of genetic diversity in land-races of waxy maize from Yunnan and Guizhou using SSR markers. Sci Agric Sin 4:648–653
Matsuoka Y, Mitchell SE, Kresovich S, Goodman M, Doebley J (2002) Microsatellites in Zea—variability, patterns of mutations, and use for evolutionary studies. Theor Appl Genet 104:436–450
Magorokosho C (2006). Genetic diversity and field performance of maize varieties from Zimbave, Zambia and Malawi. Dissertation, Texas A&M University, South Africa
Okumus A (2007) Genetic variation and relationship between Turkish flint maize landraces by RAPD markers. Am J Agri Biol Sci 2(2):49–53
Ögel B (2000) Türk Kültür tarihine Giriş Cilt-II. ‘Türk köy ve Şehir hayatı GökTürklerden Osmanlılara’. T.C. Kültür Bakanlığı Yayınları/638 Yayınlar Başkalığı Kültür Eserleri Dizisi/46 (in Turkish)
Özkan H, Kafkas S, Ozer MS, Brandolini A (2005) Genetic relationship among South-East Turkey wild barley population and sampling strategies of Hordeum spontaneum. Theor Appl Genet 112:12–20
Pressoir G, Berhaud J (2004) Patterns of population structure in maize landraces from the Central Valleys of Oaxaca in Mexico. Heredity 92:88–94
Qi-Lun Y, Ping F, Shu-Xian Z (2008) Constructing a core collection for maize (Zea mays L.) landrace from Wuling mountain region in China. Agr Sci China 7(12):1423–1432
Rao NK (2004) Plant genetic resources: advancing conservation and use through biotechnology. Afr J Biotechnol 3:136–145
Rebourg C, Gousnard B, Charcosset A (2001) Large scale molecular analysis of European maize populations. Relationship with morphological variation. Heredity 86:576–584
Reif JC, Hamrit S, Heckenberger M, Schipprack W, Peter Maurer H, Bohn M, Melchinger AE (2005) Genetic structure and diversity of European flint maize populations determined with SSR analysis of individual and bulks. Theor Appl Genet 111:906–913
Rohlf FJ (2004) NTSYS-pc ver 2.11 T. Exter Software. Satauket, NY
Ruiz de Galarreta JI, Alvarez A (2001) Morphological classification of maize landraces from Northern Spain. Genet Resour Crop Evol 48:391–400
Saeed A, Hovsepyan H, Darvishzadeh R, Imtiaz M, Panguluri SK, Nazaryan R (2011) Genetic diversity of Iranian accession, improved lines of chickpea (Cicer arietinum L.) and their wild relatives using simple sequence repeats. Plant Mol Biol Rep. doi:10.1007/s11105-011-0294-5
SAS Institute (2002) The SAS system for windows, release 9.0. SAS Institute Inc, Cary
Senior ML, Murphy JP, Goodman MM, Stuber CW (1998) Utility of SSRs for determining genetic similarities and relationships in maize using agarose gel system. Crop Sci 38:1088–1098
Sharma L, Prasanna BM, Ramesh B (2010) Analysis of phenotypic and microsattelite-based diversity of maize landraces in India, especially from the North East Himalayan region. Genetica. doi:10.1007/s10709-010-9436-1
Sharma SS, Negi MS, Sinha P, Kumar K, Tripathi SB (2011) Assessment of genetic diversity of biodiesel species pongamia pinnata accessions using AFLP and three endonuclease-AFLP. Plant Mol Biol Rep 29:12–18
Schuelke M (2000) An economic method for the fluorescentlabeling of PCR fragments. Nat Biotechnol 18:233–234
Smith JSC, Smith OS (1992) Fingerprinting crop varieties. Adv Agron 47:85–140
Taşdan K (2005) Maize market in turkey. PhD thesis, Institute of Natural and Applied Sciences University of Çukurova, Adana, Turkey
Uzun A, Yesiloglu T, Polat I, Kacar YA, Gulsen O, Yildirim B, Tuzcu O, Tepe S, Canan I, Anil S (2011) Evaluation of genetic diversity in lemon and some of their relatives based on SRAP and SSR markers. Plant Mol Biol Rep. doi:10.1007/s11105-010-0277-y
Warburton ML, Xia XC, Crossa J, Franco J, Melchinger AE, Frisch M, Bohn M, Hoisintong D (2002) Genetic characterization of CIMMYT maize inbred lines and open pollinated populations using large scale fingerprinting methods. Crop Sci 42:1832–1840
Wu YS, Zheng YL, Sun R, Wu SY, Gu HB, Bi YH (2004) Genetic diversity of waxy corn and popcorn landraces in Yunan by SSR markers. Acta Agron Sin 30:36–42
Xia X, Reif J, Hoisington D, Melchinger A, Frisch M, Warburton M (2004) Genetic diversity among CIMMYT maize inbred lines investigated with SSR markers: I. Lowland tropical maize. Crop Sci 44:2230–2237
Xie C, Warburton M, Li M, Li X, Xiao M, Hao Z, Zhao Q, Zhang S (2008) An analysis of population structure and linkage disequilibrium using multilocus data in 187 maize inbred lines. Mol Breed 21:407–418
Yao Q, Yang K, Pan G, Rong T (2007) Genetic diversity of maize (Zea mays L.) landraces from Southwest China based on SSR data. J Genet Genom 34(9):851–860
Yücel C, Baloch FS, Özkan H (2009) Genetic analysis of some physical properties of bread wheat grain (Triticum aestivum L. em Thell). Turk J Agric For 33:525–535
Zeven AC (1998) Landraces: a review of definition and classification. Euphytica 104:127–139
Acknowledgement
We thank the Menemen gene bank (Aegean Agricultural Research Institute, Izmir, Turkey) for the kind provision of landrace seed stocks. The authors express their gratitude to TÜBİTAK (The Scientific and Technological Research Council of Turkey, TOVAG-104O186) and University of Cukurova, Scientific Research Projects Unit (ZF2004BAP17) for their financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is dedicated to our dear colleague, the late Prof. Dr. Ahmet Can Ülger.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
Means and standard deviation for: days to tasseling (DTS), plant height (PHT), ear length (EHT), stem diameter (SDI), angel of uppermost leaf (AUL), number of leaves (NL) and leaf area index (LEA) of Turkish maize landraces (DOC 158 kb)
Supplementary Table 2
Means and standard deviation for: tassels on primary branches (PBNT), ear length (ELG), ear diameter (EDI), ear rows number (ERN), number of kernels per row (NKR) and total number of kernels per ear (TNKE) of Turkish maize landraces (DOC 152 kb)
Supplementary Table 3
Means and standard deviation for: ear peduncle length (EPLG), ear weight (EWE), individual plant yield (PYL), kernel length (KLG), kernel width (KWD) and kernel thickness (KT) of Turkish maize landraces (DOC 161 kb)
Rights and permissions
About this article
Cite this article
Cömertpay, G., Baloch, F.S., Kilian, B. et al. Diversity Assessment of Turkish Maize Landraces Based on Fluorescent Labelled SSR Markers. Plant Mol Biol Rep 30, 261–274 (2012). https://doi.org/10.1007/s11105-011-0332-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-011-0332-3