Abstract
Key Message
Integrative omics approaches revealed a crosstalk among phytohormones during tuberous root development in cassava.
Abstract
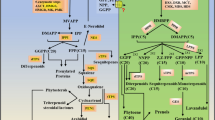
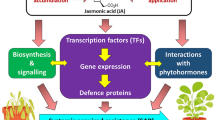
Tuberous root formation is a complex process consisting of phase changes as well as cell division and elongation for radial growth. We performed an integrated analysis to clarify the relationships among metabolites, phytohormones, and gene transcription during tuberous root formation in cassava (Manihot esculenta Crantz). We also confirmed the effects of the auxin (AUX), cytokinin (CK), abscisic acid (ABA), jasmonic acid (JA), gibberellin (GA), brassinosteroid (BR), salicylic acid, and indole-3-acetic acid conjugated with aspartic acid on tuberous root development. An integrated analysis of metabolites and gene expression indicated the expression levels of several genes encoding enzymes involved in starch biosynthesis and sucrose metabolism are up-regulated during tuberous root development, which is consistent with the accumulation of starch, sugar phosphates, and nucleotides. An integrated analysis of phytohormones and gene transcripts revealed a relationship among AUX signaling, CK signaling, and BR signaling, with AUX, CK, and BR inducing tuberous root development. In contrast, ABA and JA inhibited tuberous root development. These phenomena might represent the differences between stem tubers (e.g., potato) and root tubers (e.g., cassava). On the basis of these results, a phytohormonal regulatory model for tuberous root development was constructed. This model may be useful for future phytohormonal studies involving cassava.









Similar content being viewed by others
References
Abdala G, Castro G, Miersch O, Pearce D (2002) Changes in jasmonate and gibberellin levels during development of potato plants (Solanum tuberosum). Plant Growth Regul 36:121–126
Aksenova NP, Konstantinova TN, Golyanovskaya SA, Kossmann J, Willmitzer L, Romanov GA (2000) Transformed potato plants as a model for studying the hormonal and carbohydrate regulation of tuberization. Russian J Plant Physiol 47:370–379
Aksenova NP, Konstantinova TN, Golyanovskaya SA, Sergeeva LI, Romanov GA (2012) Hormonal regulation of tuber formation in potato. Russian J Plant Physiol 59:451–466
Asami T, Nakano T, Fujioka S (2006) Plant brassinosteroid hormones. Vitam Horm 72:479–504
Bak S, Tax F, Feldmann K, Galbraith D, Feyereisen R (2001) CYP83B1, a cytochrome P450 at the metabolic branch point in auxin and indole glucosinolate biosynthesis in Arabidopsis. Plant Cell 13:101–111
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Statist Soc B 57:289–300
Bou-Torrent J, Martinez-Garcia JF, Garcia-Martinez JL, Prat S (2011) Gibberellin A1 metabolism contributes to the control of photoperiod-mediated tuberization in potato. PLoS ONE 6:e24458
Bürger M, Chory J (2019) Stressed out about hormones: How plants orchestrate immunity. Cell Host Microbe 26:163–172
Chaweewan Y, Taylor N (2015) Anatomical assessment of root formation and tuberization in Cassava (Manihot esculenta Crantz). Tropical Plant Biol 8:1–8
Faivre-Rampant O, Cardle L, Marshall D, Viola R, Taylor M (2004) Changes in gene expression during meristem activation processes in Solanum tuberosum with a focus on the regulation of an auxin response factor gene. J Exp Bot 55:613–622
Fan M, Liu Z, Zhou L, Lin T, Liu Y, Luo L (2011) Effects of plant growth regulators and saccharide on in vitro plant and tuberous root regeneration of cassava (Manihot esculenta Crantz). J Plant Growth Regul 30:11–19
Gao J, Cao X, Shi S, Ma Y, Wang K, Liu S, Chen D, Chen Q, Ma H (2016) Genome-wide survey of Aux/IAA gene family members in potato (Solanum tuberosum): identification, expression analysis, and evaluation of their roles in tuber development. Biochem Biophys Res Commun 471:320–327
Gopal J, Chamail A, Sarkar D (2004) In vitro production of microtubers for conservation of potato germplasm: effect of genotype, abscisic acid, and sucrose. Vitro Cellular Devel Biol-Plant 40:485–490
Hannapel DJ, Banerjee AK (2017) Multiple mobile mRNA signals regulate tuber development in potato. Plants (Basel) 6:e8
Hannapel DJ, Sharma P, Lin T, Banerjee AK (2017) The multiple signals that control tuber formation. Plant Physiol 174:845–856
Hay A, Tsiantis M (2010) KNOX genes: versatile regulators of plant development and diversity. Development 137:3153–3165
Hirose N, Makita N, Kojima M, Kamada-Nobusada T, Sakakibara H (2007) Overexpression of a type-A response regulator alters rice morphology and cytokinin metabolism. Plant Cell Physiol 48:523–539
Ito Y, Kurata N (2006) Identification and characterization of cytokinin-signalling gene families in rice. Gene 382:57–65
Ivana M, Lidiya S, Miloš O, Oksana Z, Tatyana K, Josef E, Jaroslava O, Svetlana G, Yurii R, Nina A (1997) Growth pattern, tuber formation and hormonal balance in in vitro potato plants carrying ipt gene. Plant Growth Regul 21:27–36
Jain M, Tyagi AK, Khurana JP (2006) Molecular characterization and differential expression of cytokinin-responsive type-A response regulators in rice (Oryza sativa). BMC Plant Biol 6:1
Jiang Z, Liu X, Peng Z, Wan Y, Ji Y, He W, Wan W, Luo J, Guo H (2011) AHD2.0: an update version of Arabidopsis hormone database for plant systematic studies. Nucleic Acids Res 39:D1123–D1129
Jiang Z, Li J, Qu L-J (2017) Hormone metabolism and signaling in plants. In: Jiang Z, Li C, Smit S (eds) Auxins. Elsevier, Amsterdam, pp 39–51
Kasahara H (2016) Current aspects of auxin biosynthesis in plants. Biosci Biotechnol Biochem 80:34–42
Kiba T, Takei K, Kojima M, Sakakibara H (2013) Side-chain modification of cytokinins controls shoot growth in Arabidopsis. Dev Cell 27:452–461
Kloosterman B, Navarro C, Bijsterbosch G, Lange T, Prat S, Visser RG, Bachem CW (2007) StGA2ox1 is induced prior to stolon swelling and controls GA levels during potato tuber development. Plant J 52:362–373
Koda Y, Okazawa Y (1983) Characteristic changes in the levels of endogenous plant hormones in relation to the onset of potato tuberization. Jpn J Crop Sci 52:592–597
Kojima M, Kamada-Nobusada T, Komatsu H, Takei K, Kuroha T, Mizutani M, Ashikari M, Ueguchi-Tanaka M, Matsuoka M, Suzuki K, Sakakibara H (2009) Highly sensitive and high-throughput analysis of plant hormones using MS-probe modification and liquid chromatography-tandem mass spectrometry: an application for hormone profiling in Oryza sativa. Plant Cell Physiol 50:1201–1214
Kolachevskaya OO, Alekseeva VV, Sergeeva LI, Rukavtsova EB, Getman IA, Vreugdenhil D, Buryanov YI, Romanov GA (2015) Expression of auxin synthesis gene tms1 under control of tuber-specific promoter enhances potato tuberization in vitro. J Integr Plant Biol 57:734–744
Kolachevskaya OO, Sergeeva LI, Floková K, Getman IA, Lomin SN, Alekseeva VV, Rukavtsova EB, Buryanov YI, Romanov GA (2017) Auxin synthesis gene tms1 driven by tuber-specific promoter alters hormonal status of transgenic potato plants and their responses to exogenous phytohormones. Plant Cell Rep 36:419–435
Kolachevskaya OO, Lomin SN, Arkhipov DV, Romanov GA (2019) Auxins in potato: molecular aspects and emerging roles in tuber formation and stress resistance. Plant Cell Rep 38:681–698
Ku AT, Huang YS, Wang YS, Ma D, Yeh KW (2008) IbMADS1 (Ipomoea batatas MADS-box 1 gene) is involved in tuberous root initiation in sweet potato (Ipomoea batatas). Ann Bot 102:57–67
Ku YS, Sintaha M, Cheung MY, Lam HM (2018) Plant hormone signaling crosstalks between biotic and abiotic stress responses. Int J Mol Sci 17:19
Kumar D, Wareing PF (1974) Studies on tuberization of Solanum andigena. II. Growth hormone and tuberization. New Phytol 73:833–840
Kusano M, Fukushima A, Kobayashi M, Hayashi N, Jonsson P, Moritz T, Ebana K, Saito K (2007) Application of a metabolomic method combining one-dimensional and two-dimensional gas chromatography-time-of-flight/mass spectrometry to metabolic phenotyping of natural variants in rice. J Chromatogr B Analyt Technol Biomed Life Sci 855:71–79
Kusano M, Redestig H, Hirai T, Oikawa A, Matsuda F, Fukushima A, Arita M, Watanabe S, Yano M, Hiwasa-Tanase K, Ezura H, Saito K (2011) Covering chemical diversity of genetically-modified tomatoes using metabolomics for objective substantial equivalence assessment. PLoS ONE 6:e16989
Kusano K, Baxter I, Fukushima A, Oikawa A, Okazaki Y, Nakabayashi R, Bouvrette DJ, Achard F, Jakubowski AR, Ballam JM, Phillips JR, Culler AH, Saito K (2015) Assessing metabolomic and chemical diversity of a soybean lineage representing 35 years of breeding. Metabolomics 11:261–270
Li SB, Xie ZZ, Hu CG, Zhang JZ (2016) A review of auxin response factors (ARFs) in plants. Front Plant Sci 7:47
Ludwig-Muller J (2011) Auxin conjugates: their role for plant development and in the evolution of land plants. J Exp Bot 62:1757–1773
Maruyama K, Urano K, Yoshiwara K, Morishita Y, Sakurai N, Suzuki H, Kojima M, Sakakibara H, Shibata D, Saito K, Shinozaki K, Yamaguchi-Shinozaki K (2014) Integrated analysis of the effects of cold and dehydration on rice metabolites, phytohormones, and gene transcripts. Plant Physiol 164:1759–1771
McNeil SD, Nuccio ML, Ziemak MJ, Hanson AD (2001) Enhanced synthesis of choline and glycine betaine in transgenic tobacco plants that overexpress phosphoethanolamine N-methyltransferase. Proc Natl Acad Sci U S A 98:10001–10005
Medina RD, Faloci MM, Gonzalez AM, Mroginski LA (2007) In vitro cultured primary roots derived from stem segments of cassava (Manihot esculenta) can behave like storage organs. Ann Bot 99:409–423
Mehdi R, Lamm CE, Bodampalli Anjanappa R, Müdsam C, Saeed M, Klima J, Kraner ME, Ludewig F, Knoblauch M, Gruissem W, Sonnewald U, Zierer W (2019) Symplasmic phloem unloading and radial post-phloem transport via vascular rays in tuberous roots of Manihot esculenta. J Exp Bot 70:5559–5573
Mendez-Hernandez HA, Ledezma-Rodriguez M, Avilez-Montalvo RN, Juarez-Gomez YL, Skeete A, Avilez-Montalvo J, De-la-Pena C, Loyola-Vargas VM (2019) Signaling overview of plant somatic embryogenesis. Front Plant Sci 10:77
Mockaitis K, Estelle M (2008) Auxin receptors and plant development: a new signaling paradigm. Annu Rev Cell Dev Biol 24:55–80
Mussig C, Shin GH, Altmann T (2003) Brassinosteroids promote root growth in Arabidopsis. Plant Physiol 133:1261–1271
Ohdan T, Francisco PB Jr, Sawada T, Hirose T, Terao T, Satoh H, Nakamura Y (2005) Expression profiling of genes involved in starch synthesis in sink and source organs of rice. J Exp Bot 56:3229–3244
Pelacho AM, Mingo-Castel AM (1991) Jasmonic acid induces tuberization of potato stolons cultured in vitro. Plant Physiol 97:1253–1255
Redestig H, Fukushima A, Stenlund H, Moritz T, Arita M, Saito K, Kusano M (2009) Compensation for systematic cross-contribution improves normalization of mass spectrometry based metabolomics data. Anal Chem 81:7974–7980
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43:e47
Rodríguez-Falcón M, Bou J, Prat S (2006) Seasonal control of tuberization in potato: conserved elements with the flowering response. Annu Rev Plant Biol 57:151–180
Roitsch T, Ehneß R (2000) Regulation of source/sink relations by cytokinins. Plant Growth Regul 32:359–367
Romanov GA, Aksenova NP, Konstantinova TN, Golyanovskaya SA, Kossmann J, Willmitzer L (2000) Effect of indole-3-acetic acid and kinetin on tuberization parameters of different cultivars and transgenic lines of potato in vitro. Plant Growth Regul 32:245–251
Roumeliotis E, Visser RG, Bachem CW (2012) A crosstalk of auxin and GA during tuber development. Plant Signal Behav 7:1360–1363
Roumeliotis E, Kloosterman B, Oortwijn M, Visser RG, Bachem CW (2013) The PIN family of proteins in potato and their putative role in tuberization. Front Plant Sci 4:524
Sanchez Carranza AP, Singh A, Steinberger K, Panigrahi K, Palme K, Dovzhenko A, Dal Bosco C (2016) Hydrolases of the ILR1-like family of Arabidopsis thaliana modulate auxin response by regulating auxin homeostasis in the endoplasmic reticulum. Sci Rep 6:24212
Schnorr K, Gaillard C, Biget C, Nygaard P, Laioue M (1996) A second form of adenine phosphoribosyltransferase in Arabiodpsis thaliana with relative specificity towards cytokinins. Plant Journal 9:891–898
Shigenaga AM, Argueso CT (2016) No hormone to rule them all: interactions of plant hormones during the responses of plants to pathogens. Semin Cell Dev Biol 56:174–189
Simm S, Scharf K-D, Jegadeesan S, Chiusano ML, Firon N, Schleiff E (2016) Survey of genes involved in biosynthesis, transport, and signaling of phytohormones with focus on Solanum lycopersicum. Bioinfor Biol Insights 10:185–207
Smehilova M, Dobruskova J, Novak O, Takac T, Galuszka P (2016) Cytokinin-specific glycosyltransferases possess different roles in cytokinin homeostasis maintenance. Front Plant Sci 7:1264
Smith SM, Fulton DC, Chia T, Thorneycroft D, Chapple A, Dunstan H, Hylton C, Zeeman SC, Smith AM (2004) Diurnal changes in the transcriptome encoding enzymes of starch metabolism provide evidence for both transcriptional and posttranscriptional regulation of starch metabolism in Arabidopsis leaves. Plant Physiol 136:2687–2699
Sojikul P, Saithong T, Kalapanulak S, Pisuttinusart N, Limsirichaikul S, Tanaka M, Utsumi Y, Sakurai T, Seki M, Narangajavana J (2015) Genome-wide analysis reveals phytohormone action during cassava storage root initiation. Plant Mol Biol 88:531–543
Sung S, Xu D, Black C (1989) Idenfitication of activiely filling sucrose sinks. Plant Physiol 89:1117–1121
Teerawanichpan P, Lertpanyasampatha M, Netrphan S, Varavinit S, Boonseng O, Narangajavana J (2008) Influence of cassava storage root development and environmental conditions on starch granule size distribution. Starch 60:696–705
Tian T, Liu Y, Yan H, You Q, Yi X, Du Z, Xu W, Su Z (2017) agriGO v2.0: a GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res 45:W122–W129
Tokunaga H, Kojima M, Kuroha T, Ishida T, Sugimoto K, Kiba T, Sakakibara H (2012) Arabidopsis lonely guy (LOG) multiple mutants reveal a central role of the LOG-dependent pathway in cytokinin activation. Plant J 69:355–365
Ulmasov T, Hagen G, Guilfoyle T (1999) Activation and repression of transcription by auxin-response factors. Proc Natl Acad Sci USA 96:5844–5849
Utsumi Y, Utsumi C, Sawada T, Fujita N, Nakamura Y (2011) Functional diversity of isoamylase oligomers: the ISA1 homo-oligomer is essential for amylopectin biosynthesis in rice endosperm. Plant Physiol 156:61–77
Utsumi Y, Tanaka M, Morosawa T, Kurotani A, Yoshida T, Mochida K, Matsui A, Umemura Y, Ishitani M, Shinozaki K, Sakurai T, Seki M (2012) Transcriptome analysis using a high-density oligomicroarray under drought stress in various genotypes of cassava: an important tropical crop. DNA Res 19:335–345
Utsumi Y, Tanaka M, Kurotani A, Yoshida T, Mochida K, Matsui A, Ishitani M, Sraphet S, Whankaew S, Asvarak T, Narangajavana J, Triwitayakorn K, Sakurai T, Seki M (2016) Cassava (Manihot esculenta) transcriptome analysis in response to infection by the fungus Colletotrichum gloeosporioides using an oligonucleotide-DNA microarray. J Plant Res 129:711–726
Utsumi Y, Utsumi C, Tanaka M, Ha VT, Matsui A, Takahashi S, Seki M (2017) Formation of friable embryogenic callus in cassava is enhanced under conditions of reduced nitrate, potassium and phosphate. PLoS ONE 12:e0180736
Wang B, Bailly A, Zwiewka M, Henrichs S, Azzarello E, Mancuso S, Maeshima M, Friml J, Schulz A, Geisler M (2013) Arabidopsis TWISTED DWARF1 functionally interacts with auxin exporter ABCB1 on the root plasma membrane. Plant Cell 25:202–214
Wang D, Cheng L, Wang Y, Zhang F (2018) Comparative proteomic analysis of potato (Solanum tuberosum L.) tuberization in vitro regulated by IAA. Am J Potato Res 95:395–412
Xu X, van Lammeren AAM, Vermeer E, Vreugdenhil D (1998) The role of gibberellin, abscisic acid, and sucrose in the regulation of potato tuber formation in vitro. Plant Physiol 117:575–584
Yang J, An D, Zhang P (2011) Expression profiling of cassava storage roots reveals an active process of glycolysis/gluconeogenesis. J Integr Plant Biol 53:193–211
Acknowledgements
We thank Dr. Salome Prato (Departamento de Genética Molecular de Plantas, Centro Nacional de Biotecnología, Madrid, Spain.) and Dr. Uwe Sonnewald (Biochemistry, Department of Biology, Friedrich-Alexander University Erlangen-Nuremberg, Erlangen, Germany) for their critical suggestions and discussion. This work was financially supported by the East Asia Science and Innovation Area Joint Research Program (to M.S.), the Strategic Funds for the Promotion of Science and Technology (to M.S.), and the EIG CONCERT-Japan (JPMJSC16C4) 4th Call under the Strategic International Research Cooperative Program (to M.S.) of the Japan Science and Technology Agency (JST) as well as the Science and Technology Research Partnership for Sustainable Development (to M.S.) of the JST/Japan International Cooperation Agency and the RIKEN Center for Sustainable Resource Science (to M.S.). We thank Edanz Group (https://en-author-services.edanzgroup.com/) for editing a draft of this manuscript.
Author information
Authors and Affiliations
Contributions
YU and MS supervised the experiments and wrote the manuscript. YU analyzed the data. YU and MT completed the gene expression analysis. YU and CU were responsible for plant management. AM and ST assisted with the transcriptome analysis. AF, Makoto K, RS, Miyako K, AO, and KS completed the metabolome analysis. HS and MK completed the phytohormone analysis. PS and JN prepared the cassava samples.
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11103_2020_1033_MOESM1_ESM.tif
Supplementary Figure 1. Data obtained from metabolome, simultaneous analysis of phytohormones, and transcriptome analyses. We collected 59,135 data points from our transcriptome analysis, 1,125 data points from our metabolome analysis (221 data points by GC-MS and 904 data points by CE-MS), 25 data points from our phytohormone analysis and data regarding the starch content after the filtering process. Integrated data are presented in this report. Supplementary file1 (TIF 742 kb)
11103_2020_1033_MOESM2_ESM.tif
Supplementary Figure 2. Identification of differentially accumulated metabolites from the metabolome data of the pre-tuberous root, parenchyma, and cortex samples. Supplementary file2 (TIF 2610 kb)
11103_2020_1033_MOESM3_ESM.tif
Supplementary Figure 3. Heat map analysis of 135 metabolites in all samples. Red and blue represent increasing and decreasing metabolite contents, respectively, relative to the levels in F4 root or L4 leaf samples. Supplementary file3 (TIF 2671 kb)
11103_2020_1033_MOESM4_ESM.tif
Supplementary Figure 4. Heat map of Pearson’s correlation coefficients for the characteristic metabolites. The correlation coefficients represent the relationships between the metabolites at each stage. Correlation coefficients are presented with a continuous color gradient, where significant positive and negative correlations are marked in red and blue, respectively. Yellow indicates no significant correlation. Color bars represent the significance of the correlation coefficients. Supplementary file4 (TIF 5799 kb)
11103_2020_1033_MOESM5_ESM.tif
Supplementary Figure 5. Selection of differentially expressed genes (DEGs) predominantly induced in parenchyma and cortex (bark) at 8 weeks and 12 weeks after cutting. a Venn diagrams presenting the co-occurrence of DEGs (BH FDR < 0.05, fold change ≥ 2.0 or ≤ 2.0) in the parenchyma at 8 and 12 weeks. DEGs were compared to the gene expression of each gene in F4 and that of each parenchyma sample (I8_Pa, S8_Pa, I12_Pa and S12_Pa). The upregulated and downregulated genes compared with the gene expression of F4 was represented as UP and DOWN, respectively. b Venn diagrams presenting the co-occurrence of DEGs (BH FDR < 0.05, fold change ≥ 2.0 or ≤ 2.0) in the cortex at 8 and 12 weeks. DEGs were compared to the gene expression of each gene in F4 and that of each cortex (bark) sample (I8_Co, S8_Co, I12_Co and S12_Co). The upregulated and downregulated genes compared with the gene expression of F4 was represented as UP and DOWN, respectively. Supplementary file5 (TIF 1834 kb)
11103_2020_1033_MOESM6_ESM.tif
Supplementary Figure 6. GO functional categories of up-regulated DEGs in the parenchyma (Figure 6b). GO biological process functional categories for 2,848 genes were identified with the Singular Enrichment Analysis (SEA) of agriGO. The first pair of numerals represents the number of genes in the input list associated with the GO term and the number of genes in the input list. The second pair of numerals represents the number of genes associated with the GO term in the Arabidopsis database, the total number of Arabidopsis genes with GO annotations, and the p-value in parentheses. The color of the arrows indicates the relationship among the GO terms. Black stands for ‘is a’, which means that the whole of genes in upper GO term is included in the lower GO term on hierarchical tree. Orange stands for ‘part of’, which means that the part of genes in upper GO term is included in lower GO term on hierarchical tree. Red stands for ‘positive regulate’. Purple stands for ‘regulate’. Green indicates ‘negative regulate’. Long dashed lines indicate ‘two significant nodes’, and short dashed lines stands for ‘one significant node’. Long dashed, short dashed, and solid lines represent two-, one- and zero- enriched terms between connected boxes, respectively. Box colors indicate the levels of statistical significance: yellow < 0.01; orange < 0.001. White boxes indicate non-significant terms. Supplementary file6 (TIF 2063 kb)
11103_2020_1033_MOESM7_ESM.tif
Supplementary Figure 7. GO functional categories of up-regulated DEGs in the cortex (Figure 6b). GO biological process functional categories for 548 genes were identified with the Singular Enrichment Analysis (SEA) of agriGO. The first pair of numerals represents the number of genes in the input list associated with the GO term and the number of genes in the input list. The second pair of numerals represents the number of genes associated with the GO term in the Arabidopsis database, the total number of Arabidopsis genes with GO annotations, and the p-value in parentheses. The color of the arrows indicates the relationship among the GO terms. Black stands for ‘is a’, which means that the whole of genes in upper GO term is included in the lower GO term on hierarchical tree. Orange stands for ‘part of’, which means that the part of genes in upper GO term is included in lower GO term on hierarchical tree. Red stands for ‘positive regulate’. Purple stands for ‘regulate’. Green indicates ‘negative regulate’. Long dashed lines indicate ‘two significant nodes’, and short dashed lines stands for ‘one significant node’. Long dashed, short dashed, and solid lines represent two-, one- and zero- enriched terms between connected boxes, respectively. Box colors indicate levels of statistical significance: yellow < 0.01; orange < 0.001. White boxes indicate non-significant terms. Supplementary file7 (TIF 1471 kb)
11103_2020_1033_MOESM8_ESM.tif
Supplementary Figure 8. GO functional categories of up-regulated DEGs in both the cortex and parenchyma (Figure 6b). GO biological process functional categories for 1,250 genes were identified with the Singular Enrichment Analysis (SEA) of agriGO. The first pair of numerals represents the number of genes in the input list associated with the GO term and the number of genes in the input list. The second pair of numerals represents the number of genes associated with the GO term in the Arabidopsis database, the total number of Arabidopsis genes with GO annotations, and the p-value in parentheses. The color of the arrows indicates the relationship among the GO terms. Black stands for ‘is a’, which means that the whole of genes in upper GO term is included in the lower GO term on hierarchical tree. Orange stands for ‘part of’, which means that the part of genes in upper GO term is included in lower GO term on hierarchical tree. Red stands for ‘positive regulate’. Purple stands for ‘regulate’. Green indicates ‘negative regulate’. Long dashed lines indicate ‘two significant nodes’, and short dashed lines stands for ‘one significant node’. Long dashed, short dashed, and solid lines represent two-, one- and zero- enriched terms between connected boxes, respectively. Box colors indicate levels of statistical significance: yellow < 0.01; orange < 0.001. White boxes indicate non-significant terms. Supplementary file8 (TIF 2000 kb)
11103_2020_1033_MOESM9_ESM.tif
Supplementary Figure 9. GO functional categories of down-regulated DEGs in both the cortex and parenchyma (Figure 6b). GO biological process functional categories for 1,235 genes were identified with the Singular Enrichment Analysis (SEA) of agriGO. The first pair of numerals represents the number of genes in the input list associated with the GO term and the number of genes in the input list. The second pair of numerals represents the number of genes associated with the GO term in the Arabidopsis database, the total number of Arabidopsis genes with GO annotations, and the p-value in parentheses. The color of the arrows indicates the relationship among the GO terms. Black stands for ‘is a’, which means that the whole of genes in upper GO term is included in the lower GO term on hierarchical tree. Orange stands for ‘part of’, which means that the part of genes in upper GO term is included in lower GO term on hierarchical tree. Red stands for ‘positive regulate’. Purple stands for ‘regulate’. Green indicates ‘negative regulate’. Long dashed lines indicate ‘two significant nodes’, and short dashed lines stands for ‘one significant node’. Long dashed, short dashed, and solid lines represent two-, one- and zero- enriched terms between connected boxes, respectively. Box colors indicate levels of statistical significance: yellow < 0.01; orange < 0.001. White boxes indicate non-significant terms. Supplementary file9 (TIF 1582 kb)
11103_2020_1033_MOESM10_ESM.tif
Supplementary Figure 10. GO functional categories of down-regulated DEGs in the parenchyma (Figure 6b). GO biological process functional categories for 2,059 genes were identified with the Singular Enrichment Analysis (SEA) of agriGO. The first pair of numerals represents the number of genes in the input list associated with the GO term and the number of genes in the input list. The second pair of numerals represents the number of genes associated with the GO term in the Arabidopsis database, the total number of Arabidopsis genes with GO annotations, and the p-value in parentheses. The color of the arrows indicates the relationship among the GO terms. Black stands for ‘is a’, which means that the whole of genes in upper GO term is included in the lower GO term on hierarchical tree. Orange stands for ‘part of’, which means that the part of genes in upper GO term is included in lower GO term on hierarchical tree. Red stands for ‘positive regulate’. Purple stands for ‘regulate’. Green indicates ‘negative regulate’. Long dashed lines indicate ‘two significant nodes’, and short dashed lines stands for ‘one significant node’. Long dashed, short dashed, and solid lines represent two-, one- and zero- enriched terms between connected boxes, respectively. Box colors indicate levels of statistical significance: yellow < 0.01; orange < 0.001. White boxes indicate non-significant terms. Supplementary file10 (TIF 1293 kb)
11103_2020_1033_MOESM11_ESM.tif
Supplementary Figure 11. Heat map presenting relative expression values (log2) of DEGs identified by one-way ANOVA with the BH method (FDR < 5e-10) from whole data sets. Red and blue represent high and low gene expression levels, respectively, relative to the levels in the F4 root or L4 leaf samples. Supplementary file11 (TIF 2306 kb)
11103_2020_1033_MOESM12_ESM.tif
Supplementary Figure 12. Correlation between each BEL and ARF gene and the genes involved in phytohormone signaling. Positive and negative correlations are marked by red and blue lines, respectively. Supplementary file12 (TIF 6846 kb)
Rights and permissions
About this article
Cite this article
Utsumi, Y., Tanaka, M., Utsumi, C. et al. Integrative omics approaches revealed a crosstalk among phytohormones during tuberous root development in cassava. Plant Mol Biol 109, 249–269 (2022). https://doi.org/10.1007/s11103-020-01033-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-020-01033-8