Abstract
Key message
CPPU-induced San Pedro type fig main crop parthenocarpy exhibited constantly increasing IAA content and more significantly enriched KEGG pathways in the receptacle than in female flowers.
Abstract
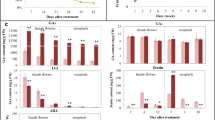
N-(2-chloro-4-pyridyl)-N-phenylurea (CPPU) was applied to San Pedro fig (Ficus carica L.) main crop to induce parthenocarpy; the optimal effect was obtained with 25 mg L−1 application to syconia when female flowers were at anthesis. To elucidate the key expression changes in parthenocarpy conversion, significant changes in phytohormone level and transcriptome of fig female flowers and receptacles were monitored. HPLC–MS revealed increased IAA content in female flowers and receptacle 2, 4 and 10 days after treatment (DAT), decreased zeatin level in the receptacle 2, 4 and 10 DAT, decreased GA3 content 2 and 4 DAT, and increased GA3 content 10 DAT. ABA level increased 2 and 4 DAT, and decreased 10 DAT. CPPU-treated syconia released more ethylene than the control except 2 DAT. RNA-Seq and bioinformatics analysis revealed notably more differentially expressed KEGG pathways in the receptacle than in female flowers. In the phytohormone gene network, GA-biosynthesis genes GA20ox and GA3ox were upregulated, along with GA signal-transduction genes GID1 and GID2, and IAA-signaling genes AUX/IAA and GH3. ABA-biosynthesis gene NCED and signaling genes PP2C and ABF were downregulated 10 DAT. One ACO gene showed consistent upregulation in both female flowers and receptacle after CPPU treatment, and more than a dozen of ERFs demonstrated opposing changes in expression. Our results revealed early-stage spatiotemporal phytohormone and transcriptomic responses in CPPU-induced San Pedro fig main crop parthenocarpy, which could be valuable for further understanding the nature of the parthenocarpy of different fig types.








Similar content being viewed by others
References
Ainalidou A, Tanou G, Belghazi M, Samiotaki M, Diamantidis G, Molassiotis A, Karamanoli K (2016) Integrated analysis of metabolites and proteins reveal aspects of the tissue-specific function of synthetic cytokinin in kiwifruit development and ripening. J Proteom 143:318–333. https://doi.org/10.1016/j.jprot.2016.02.013
Bangerth F, Bangerth F, Quinlan J (2000) Abscission and thinning of young fruit and their regulation by plant hormones and bioregulators. Plant Growth Regul 31(1–2):43–59. https://doi.org/10.1023/A:1006398513703
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate—a practical and powerful approach to multiple testing. J R Stat Soc 57(1):289–300
Boudsocq M, Laurière C (2005) Osmotic signaling in plants: multiple pathways mediated by emerging kinase families. Plant Physiol 138(3):1185–1194. https://doi.org/10.1104/pp.105.061275
Chai L, Wang Z, Chai P, Chen S, Ma H (2017) Transcriptome analysis of San Pedro-type fig (Ficus carica L.) parthenocarpic breba and non-parthenocarpic main crop reveals divergent phytohormone-related gene expression. Tree Genet Genomes 13(4):83. https://doi.org/10.1007/s11295-017-1166-4
Crane JC, Van Overbeek J (1965) Kinin-induced parthenocarpy in the fig, Ficus carica L. Science 147(3664):1468–1469. https://doi.org/10.1126/science.147.3664.1468
Ding J, Chen B, Xia X, Mao W, Shi K, Zhou Y (2013) Cytokinin-induced parthenocarpic fruit development in tomato is partly dependent on enhanced gibberellin and auxin biosynthesis. PLoS ONE 8(7):e70080. https://doi.org/10.1371/journal.pone.0070080
Du L, Bao C, Hu T, Zhu Q, Hu H, He Q, Mao W (2016) SmARF8, a transcription factor involved in parthenocarpy in eggplant. Mol Genet Genom 291(1):93–105. https://doi.org/10.1007/s00438-015-1088-5
Flaishman MA, Rodov V, Stover E (2008) The fig: botany, horticulture, and breeding. Hortic Rev 34:113–196
Freiman ZE, Rosianskey Y, Dasmohapatra R, Kamara I, Flaishman MA (2015) The ambiguous ripening nature of the fig (Ficus carica L.) fruit: a gene-expression study of potential ripening regulators and ethylene-related genes. J Exp Bot 66(11):3309. https://doi.org/10.1093/jxb/erv140
Fu FQ, Mao WH, Shi K, Zhou YH, Yu JQ (2010) Spatio-temporal changes in cell division, endoreduplication and expression of cell cycle-related genes in pollinated and plant growth substances-treated ovaries of cucumber. Plant Biol (Stuttg) 12(1):98–107. https://doi.org/10.1111/j.1438-8677.2009.00203.x
Goetz M, Hooper LC, Johnson SD, Rodrigues JC, Viviansmith A, Koltunow AM (2007) Expression of aberrant forms of AUXIN RESPONSE FACTOR8 stimulates parthenocarpy in arabidopsis and tomato. Plant Physiol 145(2):351–366. https://doi.org/10.1104/pp.107.104174
Gorguet B, Van Heusden AW, Lindhout P (2005) Parthenocarpic fruit development in tomato. Plant Biol 7(02):131–139. https://doi.org/10.1055/s-2005-837494
Greenboim-Wainberg Y, Weiss D (2005) Cross talk between gibberellin and cytokinin: the Arabidopsis GA response inhibitor SPINDLY plays a positive role in cytokinin signaling. Plant Cell 17(1):92–102. https://doi.org/10.1105/tpc.104.028472
Hayata Y, Niimi Y, Iwasaki N (1995) Synthetic cytokinin-1-(2= chloro= 4= pyridyl)-3-phenylurea (CPPU)-promotes fruit set and induces parthenocarpy in watermelon. J Am Soc Hortic Sci 120(6):997–1000
Hossain MA, Lee Y, Cho J, Ahn C, Lee S, Jeon J, Kang H, Lee C, An G, Park PB (2010) The bZIP transcription factor osABF1 is an ABA responsive element binding factor that enhances abiotic stress signaling in rice. Plant Mol Biol 72(4–5):557–566. https://doi.org/10.1007/s11103-009-9592-9
Jones B, Gunnerås SA, Petersson SV, Tarkowski P, Graham N, May S, Dolezal K, Sandberg G, Ljung K (2010) Cytokinin regulation of auxin synthesis in Arabidopsis involves a homeostatic feedback loop regulated via auxin and cytokinin signal transduction. Plant Cell 22(9):2956–2969. https://doi.org/10.1105/tpc.110.074856
Jong MD, Woltersarts M, Feron R, Mariani C, Vriezen WH (2009) The Solanum lycopersicum auxin response factor 7 (SIARF7) regulates auxin signaling during tomato fruit set and development. Plant J 57(1):160–170. https://doi.org/10.1111/j.1365-313X.2008.03671.x
Koiwai M, Okuyama F, Tanaka KI, Yamazaki A, Honma H, Ikeda K, Taira S (2012) Induction of parthenocarpy by GA and CPPU treatments aimed for seedless berry production in female vines of Japanese wild grape, Vitis coignetiae Pulliat. Hortic Res 11:87–95
Kondo S, Kawai M (1998) Relationship between free and conjugated ABA levels in seeded and gibberellin-treated seedless, maturing ‘Pione’ grape berries. J Am Soc Hortic Sci 123(5):750–754
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9(4):357–359. https://doi.org/10.1038/nmeth.1923
Li Y, Yu JQ, Ye QJ, Zhu ZJ, Guo ZJ (2003) Expression of CycD3 is transiently increased by pollination and N-(2-chloro-4-pyridyl)-N′-phenylurea in ovaries of Lagenaria leucantha. J Exp Bot 54(385):1245–1251
Li J, Wu Z, Cui L, Zhang T, Guo Q, Xu J, Jia L, Luo Q, Huang S, Li Z, Chen J (2014) Transcriptome comparison of global distinctive features between pollination and parthenocarpic fruit set reveals transcriptional phytohormone cross-talk in cucumber (Cucumis sativus L.). Plant Cell Physiol 55(7):1325–1342. https://doi.org/10.1093/pcp/pcu051
Lin W, Zhao F, Li R, Xiao H (2015) Cross-talk modulation between ABA and ethylene by transcription factor SIZFP2 during fruit development and ripening in tomato. Plant Signal Behav 10(12):e1107691. https://doi.org/10.1080/15592324.2015.1107691
Liu M, Gomes BL, Mila I, Purgatto E, Peres LE, Frasse P, Maza E, Zouine M, Roustan JP, Bouzayen M, Pirrello J (2016) Comprehensive profiling of ethylene response factors expression identifies ripening-associated ERF genes and their link to key regulators of fruit ripening in tomato (Solanum lycopersicum). Plant Physiol 170(3):1732–1744. https://doi.org/10.1104/pp.15.01859
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2–∆∆Ct method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Martínez C, Manzano S, Megías Z, Garrido D, Picó B, Jamilena M (2013) Involvement of ethylene biosynthesis and signalling in fruit set and early fruit development in zucchini squash (Cucurbita pepo L.). BMC Plant Biol 13(1):139. https://doi.org/10.1186/1471-2229-13-139
Matsuo S, Kikuchi K, Fukuda M, Honda I, Imanishi S (2012) Roles and regulation of cytokinins in tomato fruit development. J Exp Bot 63(15):5569–5579. https://doi.org/10.1093/jxb/ers207
Müller D, Leyser O (2011) Auxin, cytokinin and the control of shoot branching. Ann Bot 107(7):1203–1212. https://doi.org/10.1093/aob/mcr069
Nishitani C, Yamaguchi-Nakamura A, Hosaka F, Terakami S, Shimizu T, Yano K, Itai A, Saito T, Yamamoto T (2012) Parthenocarpic genetic resources and gene expression related to parthenocarpy among four species in pear (Pyrus, spp.). Sci Hortic 136(2):101–109. https://doi.org/10.1016/j.scienta.2011.12.029
Niu Q, Wang T, Li J, Yang Q, Qian M, Teng Y (2015) Effects of exogenous application of GA4+7, and N-(2-chloro-4-pyridyl)-N′-phenylurea on induced parthenocarpy and fruit quality in Pyrus pyrifolia, ‘Cuiguan’. Plant Growth Regul 76(3):251–258. https://doi.org/10.1007/s10725-014-9995-8
Pan X, Welti R, Wang X (2010) Quantitative analysis of major plant hormones in crude plant extracts by high-performance liquid chromatography–mass spectrometry. Nat Protoc 5(6):986–992. https://doi.org/10.1038/nprot.2010.37
Picken AJF (2015) A review of pollination and fruit set in the tomato (Lycopersicon esculentum Mill.). J Hortic Sci 59(1):1–13. https://doi.org/10.1080/00221589.1984.11515163
Qian C, Ren N, Wang J et al (2017) Effects of exogenous application of CPPU, NAA and GA 4 + 7 on parthenocarpy and fruit quality in cucumber (Cucumis sativus L.). Food Chem 243:410–413. https://doi.org/10.1016/j.foodchem.2017.09.150
Reid KE, Olsson N, Schlosser J, Peng F, Lund ST (2006) An optimized grapevine RNA isolation procedure and statistical determination of reference genes for real-time RT-PCR during berry development. BMC Plant Biol 6(1):27. https://doi.org/10.1186/1471-2229-6-27
Robinson MD, Mccarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1):139–140. https://doi.org/10.1093/bioinformatics/btp616
Roxrud I, Lid SE, Fletcher JC, Schmidt ED, Opsahl-Sorteberg HG (2007) GASA4, one of the 14-member Arabidopsis GASA family of small polypeptides, regulates flowering and seed development. Plant Cell Physiol 48(3):471–483. https://doi.org/10.1093/pcp/pcm016
Serrani JC, Carrera E, Ruiz-Rivero O, Gallego-Giraldo L, Peres LE, García-Martínez JL (2010) Inhibition of auxin transport from the ovary or from the apical shoot induces parthenocarpic fruit-set in tomato mediated by gibberellins. Plant Physiol 153(2):851–862. https://doi.org/10.1104/pp.110.155424
Shinozaki Y, Hao S, Kojima M, Sakakibara H, Ozeki-Iida Y, Zheng Y, Fei Z, Zhong S, Giovannoni JJ, Rose JK, Okabe Y, Heta Y, Ezura H, Ariizumi T (2015) Ethylene suppresses tomato (Solanum lycopersicum) fruit set through modification of gibberellin metabolism. Plant J 83(2):237–251. https://doi.org/10.1111/tpj.12882
Sotelo-Silveira M, Marsch-Martínez N, de Folter S (2014) Unraveling the signal scenario of fruit set. Planta 239(6):1147–1158. https://doi.org/10.1007/s00425-014-2057-7
Tang N, Deng W, Hu G, Hu N, Li Z (2015) Transcriptome profiling reveals the regulatory mechanism underlying pollination dependent and parthenocarpic fruit set mainly mediated by auxin and gibberellin. PLoS ONE 10(4):e0125355. https://doi.org/10.1371/journal.pone.0125355
Uchiumi T, Okamoto T (2010) Rice fruit development is associated with an increased IAA content in pollinated ovaries. Planta 232(3):579–592. https://doi.org/10.1007/s00425-010-1197-7
Upton G (1992) Fisher’s exact test. J R Stat Soc 155:395–402. https://doi.org/10.2307/2982890
Vivian-Smith A, Koltunow AM (1999) Genetic analysis of growth regulator-induced parthenocarpy in arabidopsis. Plant Physiol 121:437–451
Vriezen WH, Feron R, Maretto F, Keijman J, Mariani C (2008) Changes in tomato ovary transcriptome demonstrate complex hormonal regulation of fruit set. New Phytol 177(1):60–76. https://doi.org/10.1111/j.1469-8137.2007.02254.x
Wang H, Schauer N, Usadel B, Frasse P, Zouine M, Hernould M, Latché A, Pech JC, Fernie AR, Bouzayen M (2009) Regulatory features underlying pollination-dependent and -independent tomato fruit set revealed by transcript and primary metabolite profiling. Plant Cell 21(21):1428–1452. https://doi.org/10.1105/tpc.108.060830
Wang Y, Li L, Ye T, Zhao S, Liu Z, Feng YQ, Wu Y (2011) Cytokinin antagonizes ABA suppression to seed germination of Arabidopsis by downregulating ABI5 expression. Plant J Cell Mol Biol 68(2):249–261. https://doi.org/10.1111/j.1365-313X.2011.04683.x
Weiblen GD (2002) How to be a fig wasp. Ann Rev Entomol 47(1):299–330. https://doi.org/10.1146/annurev.ento.47.091201.145213
Werner T, Motyka V, Laucou V, Smets R, Van Onckelen H, Schmülling T (2003) Cytokinin-deficient transgenic Arabidopsis plants show multiple developmental alterations indicating opposite functions of cytokinins in the regulation of shoot and root meristem activity. Plant Cell 15(11):2532–2550. https://doi.org/10.1105/tpc.014928
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, Kong L, Gao G, Li CY, Wei L (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39:316–322. https://doi.org/10.1093/nar/gkr483
Ye X, Fu M, Liu Y, An D, Zheng X, Tan B, Li J, Cheng J, Wang W, Feng J (2018) Expression of grape ACS1 in tomato decreases ethylene and alters the balance between auxin and ethylene during shoot and root formation. J Plant Physiol 226:154–162. https://doi.org/10.1016/j.jplph.2018.04.015
Zhang J, Li X, He Z, Zhao X, Wang Q, Zhou B, Yu D, Huang X, Tang D, Guo X, Liu X (2013) Molecular character of a phosphatase 2C (PP2C) gene relation to stress tolerance in Arabidopsis thaliana. Mol Biol Rep 40(3):2633–2644. https://doi.org/10.1007/s11033-012-2350-0
Acknowledgements
This work was supported by National Natural Science Foundation of China project NSFC [31372007].
Author information
Authors and Affiliations
Contributions
HM and PC designed the experiments. PC and LC conducted the experiments and analyzed the results. SD carried out the CPPU treatment again and performed the ethylene assay in 2018. PC, SD, LC, HM, SC and MF prepared the manuscript. All authors have read and approved the manuscript for publication.
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11103_2019_820_MOESM1_ESM.docx
Supplementary material 1 Supplementary Table 1. Primer sequences of genes used for validation of RNA-Seq results by quantitative real-time PCR. Supplementary Table 2. Differential expression of genes related to plant hormone synthesis and signal transduction (|log2FC| > 1) in six pairwise comparison groups. Supplementary Table 3. Significantly enriched KEGG pathways in female flowers vs. receptacle during the San Pedro fig main crop fruit set-determination phase. Supplementary Fig. 1 Correlation of fold changes in gene expression between RNA-Seq and qRT-PCR. Equation for linear regression and correlation coefficient (R2) are shown. DAT, day after treatment. Supplementary Fig. 2 Comparison of the major phytohormone levels between female flowers and receptacle and the effect of CPPU treatment (DOCX 615 KB)
Rights and permissions
About this article
Cite this article
Chai, P., Dong, S., Chai, L. et al. Cytokinin-induced parthenocarpy of San Pedro type fig (Ficus carica L.) main crop: explained by phytohormone assay and transcriptomic network comparison. Plant Mol Biol 99, 329–346 (2019). https://doi.org/10.1007/s11103-019-00820-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-019-00820-2