Abstract
Freezing temperature/frosts can cause significant damage to plants by rupturing plant cells. Rapeseed/canola (Brassica napus L.) is susceptible to freezing temperature at early seedling stage. The degree of cell rupture or seedling damage can be evaluated through the measurement of electrolyte leakage. Here, we measured the electrolyte leakage of a diversity panel of B. napus germplasm accessions under simulated freezing conditions. Preliminary data for electrolyte leakage measurement indicated that cold acclimation of two-week-old seedlings for 7 days at 4 °C followed by freezing treatment at − 12 °C for 2 h provided a reasonable diversity in response. With this protocol for electrolyte leakage, a genome-wide association study was conducted on 157 winter, semi-winter, and spring types of B. napus accessions that originated from 17 countries. A total of 37,454 single-nucleotide polymorphism (SNP) markers based upon genotyping-by-sequencing were used for the analysis. Ten QTL were identified as associated with electrolyte leakage of canola seedlings, which together explained 43% phenotypic variation. Five of the QTL were located on A-genome. We identified at least 33 orthologs of the functional candidate genes. Although no well-characterized cold regulatory genes were identified, there were some indications that genes involved in membrane structure, developmental processes, and extracellular transport may be involved in altering the electrolyte leakage following the short-term hard freeze and rapid defrosting suffered by the plants in our protocol.



Similar content being viewed by others
References
Anchordoguy TJ, Rudolph AS, Carpenter JF, Crowe JH (1987) Modes of interaction of cryoprotectants with membrane phospholipids during freezing. Cryobiology 24:324–331
Bajji M, Kinet JM, Lutts S (2002) Osmotic and ionic effects of NaCl on germination, early seedling growth and ion content of Atriplex halimus (Chenopodiaceae). Can J Bot 80:297–304
Blum A, Ebercon A (1981) Cell membrane stability as a measure of drought and heat tolerance in wheat. Crop Sci 21:43–47
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23(19):2633–2635. https://doi.org/10.1093/bioinformatics/btm308
Chalhoub B, Denoeud F, Liu S, Parkin IAP, Tang H, Wang X, Chiquet J, Belcram H, Tong C, Samans B, Correa M, da Silva C, Just J, Falentin C, Koh CS, le Clainche I, Bernard M, Bento P, Noel B, Labadie K, Alberti A, Charles M, Arnaud D, Guo H, Daviaud C, Alamery S, Jabbari K, Zhao M, Edger PP, Chelaifa H, Tack D, Lassalle G, Mestiri I, Schnel N, le Paslier MC, Fan G, Renault V, Bayer PE, Golicz AA, Manoli S, Lee TH, Thi VHD, Chalabi S, Hu Q, Fan C, Tollenaere R, Lu Y, Battail C, Shen J, Sidebottom CHD, Wang X, Canaguier A, Chauveau A, Berard A, Deniot G, Guan M, Liu Z, Sun F, Lim YP, Lyons E, Town CD, Bancroft I, Wang X, Meng J, Ma J, Pires JC, King GJ, Brunel D, Delourme R, Renard M, Aury JM, Adams KL, Batley J, Snowdon RJ, Tost J, Edwards D, Zhou Y, Hua W, Sharpe AG, Paterson AH, Guan C, Wincker P (2014) Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345(6199):950–953
Chinnusamy V, Zhu J, Zhu JK (2007) Cold stress regulation of gene expression in plants. Trends Plant Sci 12(10):444–451
Ehsan H, Ray WK, Phinney B, Wang X, Huber SC, Clouse SD (2005) Interaction of Arabidopsis BRASSINOSTEROID-INSENSITIVE 1 receptor kinase with a homolog of mammalian TGF-beta receptor interacting protein. Plant J 43:251–261
Elshire RJ, Glaubitz JC, Poland JA, Kawamoto K, Buckler ES, Mitchell SE (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS One 6(5):e19379
Fiebelkorn D, Rahman M (2016) Development of a protocol for frost-tolerance evaluation in rapeseed/canola (Brassica napus L.). Crop J 4:147–152
Graham D, Patterson BD (1982) Response of plants to low, non-freezing temperatures: proteins, metabolism, and acclimation. Annu Rev Plant Physiol 84:872–878
Gurung S, Mamidi S, Bonman JM, Xiong M, Brown-Guedira G, Adhikari TB (2014) Genome-wide association study reveals novel quantitative trait loci associated with resistance to multiple leaf spot diseases of spring wheat. PLoS One 9(9):e108179
Gusta LV, Wisniewski M (2013) Understanding plant cold hardiness: an opinion. Physiol Plant 147:4–14
Gusta LV, Wisniewski M, Nesbitt NT, Gusta ML (2004) The effect of water, sugars, and proteins on the pattern of ice nucleation and propagation in acclimated and nonacclimated canola leaves. Plant Physiol 135:1642–1653
Hansen M, Kraft T, Ganestam S, Säll T, Nilsson NO (2001) Linkage disequilibrium mapping of the bolting gene in sea beet using AFLP markers. Genet Res 77(1):61–66
Hetherington SE, He J, Smillie RM (1989) Photoinhibition at low temperature in chilling-sensitive and -resistant plants. Plant Physiol 90:1609–1615
Hong EP, Park JW (2012) Sample size and statistical power calculation in genetic association studies. Genome Inform 10:117–122
Horton MW, Willems G, Sasaki E, Koornneef M, Nordborg M (2016) The genetic architecture of freezing tolerance varies across the range of Arabidopsis thaliana. Plant Cell Environ 39(11):2570–2579
Huang DQ, Wu WR, Abrams SR, Cutler AJ (2008) The relationship of drought-related gene expression in Arabidopsis thaliana to hormonal and environmental factors. J Exp Bot 59:2991–3007
Kaldis A, Tsementzi D, Tanriverdi O, Vlachonasios KE (2010) Arabidopsis thaliana transcriptional co-activators ADA2b and SGF29a are implicated in salt stress responses. Planta 233:749–762
Kieffer M, Stern Y, Cook H, Clerici E, Maulbetsch C, Laux T, Davies B (2006) Analysis of the transcription factor WUSCHEL and its functional homologue in Anthirrhinum reveals a potential mechanism for their roles in meristem maintenance. Plant Cell 18:560–573
Kindel PK, Liao SY, Liske MR, Olien CR (1989) Arabinoxylans from rye and wheat seed that interact with ice. Carbohydr Res 187:173–185
Koboldt DC, Zhang Q, Larson DE, Shen D, McLellan MD et al (2012) VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res 22:568–576
Kraakman ATW, Martínez F, Mussiraliev B, van Eeuwijk FA, Niks RE (2006) Linkage disequilibrium mapping of morphological, resistance, and other agronomically relevant traits in modern spring barley cultivars. Mol Breed 17(1):41–58
Lagercrantz U (1998) Comparative mapping between Arabidopsis thaliana and Brassica nigra indicates that Brassica genomes have evolved through extensive genome replication accompanied by chromosome fusions and frequent rearrangements. Genetics 150:1217–1228
Lagercrantz U, Lydiate DJ (1996) Comparative genome mapping in Brassica. Genetics 144:1903–1910
Lamberto I, Percudani R, Gatti R, Folli C, Petrucco S (2010) Conserved alternative splicing of Arabidopsis transthyretin-like determines protein localization and S-allantoin synthesis in peroxisomes. Plant Cell 22:1564–1574
Lee BH, Zhu JK (2010) Phenotypic analysis of Arabidopsis mutants: electrolyte leakage after freezing stress. Cold Spring Harb Protoc. https://doi.org/10.1101/pdb.prot4970
Lefebvre V, North H, Frey A, Sotta B, Seo M, Okamoto M, Nambara E, Marrion-Poll A (2006) Functional analysis of Arabidopsis NCED6 and NCED9 genes indicates that ABA synthesized in the endosperm is involved in the induction of seed dormancy. Plant J 45:309–319
Leopold AC, Musgrave ME, Williams KM (1981) Solute leakage resulting from leaf desiccation. Plant Physiol 68:1222–1225
Li H (2013) Aligning sequence reads, clone sequences, and assembly contigs with BWA-MEM. arXiv:1303.3997 http://arxiv.org/abs/1303.3997
Liang Y, Wang X, Hong S, Li Y, Zuo J (2012) Deletion of the initial 45 residues of ARR18 induces cytokinin response in Arabidopsis. J Genet Genomics 39:37–46
Liu J, Wang W, Mei D, Wang H, Fu L et al (2016) Characterizing variation of branch angle and genome-wide association mapping in rapeseed (Brassica napus L.). Front Plant Sci 7:21
Madakadze IC, Stewart KA, Madakadze RM, Smith DL (2003) Base temperatures for seedling growth and their correlation with chilling sensitivity for warm-season grasses. Crop Sci 43:874–878
Mamidi S, Chikara S, Goos RJ, Hyten DL, Annam D, Moghaddam SM, Lee RK, Cregan PB, McClean PE (2011) Genome-wide association analysis identifies candidate genes associated with iron deficiency chlorosis in soybean. Plant Genome 4:154–164
Mamidi S, Lee RK, Goos RJ, McClean PE (2014) Genome-wide association studies identifies seven major regions responsible for iron deficiency chlorosis in soybean (Glycine max). PLoS One 9(9):e107469
McCollum TG, McDonald RE (1991) Electrolyte leakage, respiration, and ethylene production as indices of chilling injury in grapefruit. Hortic Sci 26:1191–1192
Murray AJS, Blackwell RD, Lea PJ (1989) Metabolism of hydroxypyruvate in a mutant of barley lacking NADH-dependent hydroxypyruvate reductase, an important photorespiratory enzyme activity. Plant Physiol 91:395–400
Narusaka Y, Narusaka M, Seki M, Umezawa T, Ishida J et al (2004) Crosstalk in the responses to abiotic and biotic stresses in Arabidopsis: analysis of gene expression in cytochrome P450 gene superfamily by cDNA microarray. Plant Mol Biol 55:327–342
Olien CR (1974) Energies of freezing and frost desiccation. Plant Physiol 53:764–767
Olien CR, Smith MN (1977) Ice adhesions in relation to freeze stress. Plant Physiol 60:499–503
Osterhaut WJV (1922) Injury, recovery, and death, in relation to conductivity and permeability. JB Lippincott Company, Philadelphia and London, p 259
Poland JA, Rife TW (2012) Genotyping-by-sequencing for plant breeding and genetics. Plant Genome 5(3):92–102
Rahman M, Mamidi S, del Rio L, Ross A, Kadir MM, Rahaman MM, Arifuzzaman M (2016) Association mapping in Brassica napus (L.) accessions identifies a major QTL for blackleg disease resistance on chromosome A01. Mol Breed 36:90
Rakow G (2007) Rapeseed genetics and breeding research for sustainable oilseed production. Proceedings of the 12th International Rapeseed Congress 1:207–210
Rezaeizad A, Wittkop B, Snowdon R, Hasan M, Mohammadi V, Zali A, Friedt W (2011) Identification of QTLs for phenolic compounds in oilseed rape (Brassica napus L.) by association mapping using SSR markers. Euphytica 177:335–342
Salgado JP, Rife CL (1996) Selection for cold hardiness in oilseed rape (Brassica napus). Cruciferae Newsl 18:92–93
Scheet P, Stephens M (2006) A fast and flexible statistical model for large-scale population genotype data: applications to inferring missing genotypes and haplotypic phase. Am J Hum Genet 78:629–644
Schläppi MR, Jackson AK, Eizenga GC, Wang A, Chu C, Shi Y, Shimoyama N, Boykin DL (2017) Assessment of five chilling tolerance traits and GWAS mapping in rice using the USDA mini-core collection. Front Plant Sci 8:957
Schoof H, Lenhard M, Haecker A, Mayer KFX, Jürgens G, Laux T (2000) The stem cell population of Arabidopsis shoot meristems is maintained by a regulatory loop between the CLAVATA and WUSCHEL genes. Cell 100:635–644
Seki M, Narusaka M, Abe H, Kasuga M, Yamaguchi-Shinozaki K et al (2001) Monitoring the expression pattern of 1300 Arabidopsis genes under drought and cold stresses by using a full-length cDNA microarray. Plant Cell 13:61–72
Singh JB, Johnson-Flanagan AM (1987) Membrane alterations in winter rye and Brassica napus cells during lethal freezing and plasmolysis. Plant Cell Environ 10:163–168
Steponkus PL, Lynch DV (1989) Freeze/thaw-induced destabilization of the plasma membrane and the effects of cold acclimation. J Bioenerg Biomembr 21:21–41
Stevanovic B, Sinzar J, Glisic O (1997) Electrolyte leakage differences between polikilohydrous and homoiohydrous species of Gesneriaceae. Biol Plant 40:299–303
Sun J, Guo N, Lei J, Li L, Hu G, Xing H (2014) Association mapping for partial resistance to Phytophthora sojae in soybean (Glycine max L.). J Genet 2:355–363
Takizawa PA, Malhotra V (1993) Coatamers and SNARES in promoting membrane traffic. Cell 75:593–596
U N (1935) Genome analysis in Brassica with special reference to the experimental formation of Brassica napus and peculiar mode of fertilization. Jpn J Bot 7:389–452
Veerabagu M, Kirchler T, Elgass K, Stadelhofer B, Stahl M, Harter K, Mira-Rodado V, Chaban C (2014) The interaction of the Arabidopsis response regulator ARR18 with bZIP63 mediates the regulation of PROLINE DEHYDROGENASE expression. Mol Plant 7:1560–1577
Yamada T, Kuroda K, Jitsuyama Y, Takezawa K, Arakawa K, Fujikawa S (2002) Roles of the plasma membrane and the cell wall in the responses of plant cells to freezing. Planta 215:770–778
Yoshimoto N, Takahashi H, Smith F, Yamaya T, Saito K (2002) Two distinct high affinity sulfate transporters with different inducibilities mediate uptake of sulfate in Arabidopsis roots. Plant J 29:465–473
Yu JM, Buckler ES (2006) Genetic association mapping and genome organization of maize. Curr Opin Biotechnol 17:155–160
Yu J, Pressoir G, Briggs WH, Bi IV, Yamasaki M et al (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Zhao K, Aranzana MJ, Kim S, Lister C, Shindo C, Tang C, Toomajian C, Zheng H, Dean C, Marjoram P, Nordborg M (2007) An Arabidopsis example of association mapping in structured samples. PLoS Genet 3:e4
Acknowledgments
The authors gratefully acknowledge the partial financial support for this project from the Northern Canola Growers Association (Project# NCGA-2014) and the USDA-NIFA CRIS project (Project# ND01581).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Fig. S1
The electrolyte leakage (%) of nine genotypes tested at − 4, − 8, − 12, and − 16 °C. The genotypes are as follows: (a) DKL 70-07, (b) NDSU 15-1000, (c) Sprinter, (d) Pioneer 45H26, (e) Hi-Q, (f) Kanada, (g) Fashion, (h) ARC 2180-1, and (i) Galileo. (PDF 242 kb)
Fig. S2
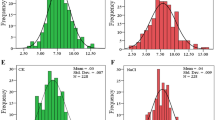
Histogram for electrolyte leakage (%) tested at − 12 °C. The x-axis is the median, and the y-axis, the number of genotypes that fit each median. (GIF 4 kb)
Table S1
(XLSX 15 kb)
Table S2
(DOCX 29 kb)
Rights and permissions
About this article
Cite this article
Fiebelkorn, D., Horvath, D. & Rahman, M. Genome-wide association study for electrolyte leakage in rapeseed/canola (Brassica napus L.). Mol Breeding 38, 129 (2018). https://doi.org/10.1007/s11032-018-0892-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-018-0892-0