Abstract
Soil salinity negatively impacts rapeseed (Brassica napus) crop production. In particular, high soil salinity is known to hinder seedling growth and establishment. Identifying natural genetic variation for high salt tolerance in Brassica napus seedlings is an effective way to breed for improved productivity under salt stress. To identify genetic variants involved in differential response to salt stress, we evaluated a diverse association panel of 228 Brasica napus accessions for four seedling traits under salt stress to establish stress susceptibility index (SSI) and stress tolerance index (STI) values, and performed genome-wide association studies (GWAS) using 201,817 high-quality single nucleotide polymorphic (SNP) markers. Our GWAS identified 142 significant SNP markers strongly associated with salt tolerance distributed across all rapeseed chromosomes, with 78 SNPs in the C genome and 64 SNPs in the A genome, and our analyses subsequently pinpointed both favorable alleles and elite cultivars. We identified 117 possible candidate genes associated with these SNPs: 95/117 were orthologous with Arabidopsis thaliana genes encoding transcription factors, aquaporins, and binding proteins. The expression level of ten candidate genes was validated by quantitative real-time PCR (qRT-PCR), and these genes were found to be differentially expressed between salt-tolerant and salt-susceptible lines under salt stress conditions. Our results provide new genetic resources and information for improving salt tolerance in rapeseed genotypes at the seed germination and seedling stages via genomic or marker-assisted selection, and for future functional characterization of putative gene candidates.






Similar content being viewed by others

Abbreviations
- GWAS:
-
Genome-wide association studies
- SLAF- seq:
-
Specific length amplified fragment sequencing
- SNPs:
-
Single nucleotide polymorphisms
- GLM:
-
General linear model
- MLM:
-
Mixed linear model
- SSI:
-
Stress Susceptible Index
- STI:
-
Stress Tolerance Index
- GR:
-
Germination
- RL:
-
Root length
- SDW:
-
Shoot dry weight
- SVI:
-
Seed Vigor Index
References
Abdul-Baki AA, Anderson JD (1973) Vigor determination in soybean seed by multiple criteria 1. Crop sci 13:630–633
Alexander DH, Novembre J, Lange K (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19:1655–1664
Babitha KC, Ramu SV, Pruthvi V, Mahesh P, Nataraja KN, Udayakumar M (2013) Co-expression of AtbHLH17 and AtWRKY28 confers resistance to abiotic stress in Arabidopsis. Transgenic Res 22:327–341
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat SocSer B (Methodological) 57:289–300
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Bui EN (2013) Soil salinity: A neglected factor in plant ecology and biogeography. J Arid Environ 92:14–25
Chalhoub B, Denoeud F, Liu S, Parkin IA, Tang H, Wang X, Chiquet J, Belcram H, Tong C, Samans B, Correa M, Da Silva C, Just J, Falentin C, Koh CS, Le Clainche I, Bernard M, Bento P, Noel B, Labadie K, Alberti A, Charles M, Arnaud D, Guo H, Daviaud C, Alamery S, Jabbari K, Zhao M, Edger PP, Chelaifa H, Tack D, Lassalle G, Mestiri I, Schnel N, Le Paslier MC, Fan G, Renault V, Bayer PE, Golicz AA, Manoli S, Lee TH, Thi VH, Chalabi S, Hu Q, Fan C, Tollenaere R, Lu Y, Battail C, Shen J, Sidebottom CH, Wang X, Canaguier A, Chauveau A, Berard A, Deniot G, Guan M, Liu Z, Sun F, Lim YP, Lyons E, Town CD, Bancroft I, Wang X, Meng J, Ma J, Pires JC, King GJ, Brunel D, Delourme R, Renard M, Aury JM, Adams KL, Batley J, Snowdon RJ, Tost J, Edwards D, Zhou Y, Hua W, Sharpe AG, Paterson AH, Guan C, Wincker P (2014) Plant genetics. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345:950–953
Dai X, Xu Y, Ma Q, Xu W, Wang T, Xue Y, Chong K (2007) Overexpression of an R1R2R3 MYB gene, OsMYB3R-2, increases tolerance to freezing, drought, and salt stress in transgenic Arabidopsis. Plant Physiol 143:1739–1751
Duval M, Hsieh TF, Kim SY, Thomas TL (2002) Molecular characterization of AtNAM: a member of the Arabidopsis NAC domain superfamily. Plant MolBiol 50:237–248
Fernandez G (1992) Effective selection criteria for assessing stress tolerance. IN Kuo, CG. In: proceedings of the international symposium on adaptation of vegetables and other food crops in temperature and water stress, pp 115–121
Fischer R, Maurer R (1978) Drought resistance in spring wheat cultivars. I. Grain yield responses. Aust J Agric Res 29:897–912
Ginestet C (2011) ggplot2: elegant graphics for data analysis. J R Stat SocSer A (Statistics in Society) 174:245–246
Guan H, Liu X, Niu F, Zhao Q, Fan N, Cao D, Meng D, He W, Guo B, Wei Y, Fu Y (2019) OoNAC72, a NAC-type oxytropisochrocephala transcription factor, conferring enhanced drought and salt stress tolerance in Arabidopsis. Front Plant Sci 10:890
Guo L, Wang ZY, Lin H, Cui WE, Chen J, Liu M, Chen ZL, Qu LJ, Gu H (2006) Expression and functional analysis of the rice plasma-membrane intrinsic protein gene family. Cell Res 16:277–286
Guo Y, Gan S (2006) AtNAP, a NAC family transcription factor, has an important role in leaf senescence. The Plant J 46:601–612
Gupta B, Huang B (2014) Mechanism of salinity tolerance in plants: physiological, biochemical, and molecular characterization. Int J Genomics 2014:701596
Hachez C, Veljanovski V, Reinhardt H, Guillaumot D, Vanhee C, Chaumont F, Batoko H (2014) The Arabidopsis abiotic stress-induced TSPO-related protein reduces cell-surface expression of the aquaporin PIP2; 7 through protein-protein interactions and autophagic degradation. Plant Cell 26:4974–4990
Han B, Huang X (2013) Sequencing-based genome-wide association study in rice. CurrOpin Plant Biol 16:133–138
Hao YJ, Wei W, Song QX, Chen HW, Zhang YQ, Wang F, Zou HF, Lei G, Tian AG, Zhang WK, Ma B, Zhang JS, Chen SY (2011) Soybean NAC transcription factors promote abiotic stress tolerance and lateral root formation in transgenic plants. The Plant J 68:302–313
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. MolEcol Notes 2:618–620
He YJ, Wu D, You J, Qian W (2017) Genome-wide association analysis of salt tolerance related traits in Brassica napus and candidate gene prediction. ScieAgri Sin 50:189–1201
Hu R, Qi G, Kong Y, Kong D, Gao Q, Zhou G (2010) Comprehensive analysis of NAC domain transcription factor gene family in Populustrichocarpa. BMC Plant Biol 10:145
Hussain S, Shaukat M, Ashraf M, Zhu C, Jin Q, Zhang J (2019) Salinity stress in arid and semi-arid climates: effects and management in field crops. In: Climate Change And Agriculture. London: IntechOpen
Jeong JS, Kim YS, Baek KH, Jung H, Ha SH, Do Choi Y, Kim M, Reuzeau C, Kim JK (2010) Root-specific expression of OsNAC10 improves drought tolerance and grain yield in rice under field drought conditions. Plant Physiol 153:185–197
Kan G, Zhang W, Yang W, Ma D, Zhang D, Hao D, Hu Z, Yu D (2015) Association mapping of soybean seed germination under salt stress. Mol Genet Genomics 290:2147–2162
Kawaura K, Mochida K, Ogihara Y (2008) Genome-wide analysis for identification of salt-responsive genes in common wheat. FunctIntegr Genomics 8:277–286
Keerio AA, Shen C, Nie Y, Ahmed MM, Zhang X, Lin Z (2018) QTL mapping for fiber quality and yield traits based on introgression lines derived from Gossypium hirsutum× G. tomentosum. Int J MolSci 19:243
Kumar V, Singh A, Mithra SV, Krishnamurthy SL, Parida SK, Jain S, Tiwari KK, Kumar P, Rao AR, Sharma SK, Khurana JP, Singh NK, Mohapatra T (2015) Genome-wide association mapping of salinity tolerance in rice (Oryza sativa). DNA Res 22:133–145
Lang L, Xu A, Ding J, Zhang Y, Zhao N, Tian Z, Liu Y, Wang Y, Liu X, Liang F, Zhang B, Qin M, Dalelhan J, Huang Z (2017) Quantitative trait locus mapping of salt tolerance and identification of salt-tolerant genes in Brassica napus L. Front Plant Sci 8:1000
Li J, Yu G, Sun X, Liu Y, Liu J, Zhang X, Jia C, Pan H (2015) AcPIP2, a plasma membrane intrinsic protein from halophyte Atriplexcanescens, enhances plant growth rate and abiotic stress tolerance when overexpressed in Arabidopsis thaliana. Plant cell Rep 34:1401–1415
Long NV, Dolstra O, Malosetti M, Kilian B, Graner A, Visser RG, van der Linden CG (2013a) Association mapping of salt tolerance in barley (Hordeum vulgare L.). Theor Appl Genet 126:2335–2351
Long WH, Pu HM, Zhang JF, Qi CK, Zhang XK (2013b) Screening of Brassica napus for salinity tolerance at germination stage. China J Oil Crop Sci 35:271–275
Lu HY, Liu XF, Wei SP, Zhang YM (2011) Epistatic association mapping in homozygous crop cultivars. PLoS ONE 6:e17773
Misra N, Dwivedi U (2004) Genotypic difference in salinity tolerance of green gram cultivars. Plant Sci 166:1135–1142
Munns R, Gilliham M (2015) Salinity tolerance of crops–what is the cost? New Phytol 208:668–673
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Nie L, Wang R, Xia Y, Li G (2015) CDPK1, an Arabidopsis thaliana calcium-dependent protein kinase, is involved in plant defense response. Russian J Plant Physiol 62:866–874
Nongpiur RC, LataSingla-Pareek S, Pareek A (2016) Genomics approaches for improving salinity stress tolerance in crop plants. Curr Genomics 17:343–357
Nuruzzaman M, Sharoni AM, Satoh K, Moumeni A, Venuprasad R, Serraj R, Kumar A, Leung H, Attia K, Kikuchi S (2012) Comprehensive gene expression analysis of the NAC gene family under normal growth conditions, hormone treatment, and drought stress conditions in rice using near-isogenic lines (NILs) generated from crossing Aday Selection (drought tolerant) and IR64. Mol Genet Genomics 287:389–410
Pandey M, Penna S (2017) Time course of physiological, biochemical, and gene expression changes under short-term salt stress in Brassica juncea L. Crop J 5:219–230
Patishtan J, Hartley TN, Fonseca de Carvalho R, Maathuis FJM (2018) Genome-wide association studies to identify rice salt-tolerance markers. Plant Cell Environ 41:970–982
Puppala N, Fowler JL, Poindexter L, Bhardwaj HL (1999) Evaluation of salinity tolerance of canola germination. Perspectives on new crops and new uses. ASHS Press, Alexardria, pp 251–253
Rahman M, Mamidi S, del Rio L, Ross A, Kadir MM, Rahaman MM, Arifuzzaman M (2016) Association mapping in Brassica napus (L) accessions identifies a major QTL for blackleg disease resistance on chromosome A01. Mol Breed 36:90
Raman H, Raman R, Coombes N, Song J, Prangnell R, Bandaranayake C, Tahira R, Sundaramoorthi V, Killian A, Meng J, Dennis ES, Balasubramanian S (2016) Genome-wide association analyses reveal complex genetic architecture underlying natural variation for flowering time in canola. Plant Cell Environ 39:1228–1239
Ranty B, Aldon D, Galaud J-P (2006) Plant calmodulins and calmodulin-related proteins: multifaceted relays to decode calcium signals. Plant Signal Behav 1:96–104
Romeis T, Piedras P, Jones JD (2000) Resistance gene-dependent activation of a calcium-dependent protein kinase in the plant defense response. Plant Cell 12:803–816
Roy SJ, Negrao S, Tester M (2014) Salt resistant crop plants. CurrOpin Biotech 26:115–124
Rushton DL, Tripathi P, Rabara RC, Lin J, Ringler P, Boken AK, Langum TJ, Smidt L, Boomsma DD, Emme NJ, Chen X, Finer JJ, Shen QJ, Rushton PJ (2012) WRKY transcription factors: key components in abscisic acid signalling. Plant Biotechnol J 10:2–11
Saijo Y, Hata S, Kyozuka J, Shimamoto K, Izui K (2000) Over-expression of a single Ca2+-dependent protein kinase confers both cold and salt/drought tolerance on rice plants. The Plant J 23:319–327
Scarpeci TE, Zanor MI, Carrillo N, Mueller-Roeber B, Valle EM (2008) Generation of superoxide anion in chloroplasts of Arabidopsis thaliana during active photosynthesis: a focus on rapidly induced genes. Plant MolBiol 66:361–378
Shabani A, Sepaskhah AR, Kamgar-Haghighi AA (2015) A model to predict the dry matter and yield of rapeseed under salinity and deficit irrigation. Arch Agron Soil Sci 61:525–542
Shi H, Xiong L, Stevenson B, Lu T, Zhu JK (2002) The Arabidopsis salt overly sensitive 4 mutants uncover a critical role for vitamin B6 in plant salt tolerance. Plant Cell 14:575–588
Shi Y, Gao L, Wu Z, Zhang X, Wang M, Zhang C, Zhang F, Zhou Y, Li Z (2017) Genome-wide association study of salt tolerance at the seed germination stage in rice. BMC Plant Biol 17:92
Singh H, Kumar P, Kumar A, Kyriacou MC, Colla G, Rouphael Y (2020) Grafting tomato as a tool to improve salt tolerance. Agronomy 10:263
Song J, Wang B (2015) Using euhalophytes to understand salt tolerance and to develop saline agriculture: Suaeda salsa as a promising model. Ann Bot 115:541–553
Sun X, Liu D, Zhang X, Li W, Liu H, Hong W, Jiang C, Guan N, Ma C, Zeng H (2013) SLAF-seq: an efficient method of large-scale de novo SNP discovery and genotyping using high-throughput sequencing. PLoS ONE 8:e58700
Turner SD (2014) qqman: an R package for visualizing GWAS results using QQ and manhattan plots. Biorxiv:005165
Tuteja N (2007) Mechanisms of high salinity tolerance in plants. Methods in enzymology. Elsevier, Amsterdam, pp 419–438
Valenzuela CE, Acevedo-Acevedo O, Miranda GS, Vergara-Barros P, Holuigue L, Figueroa CR, Figueroa PM (2016) Salt stress response triggers activation of the jasmonate signaling pathway leading to inhibition of cell elongation in Arabidopsis primary root. J Exp Bot 67:4209–4220
Vallejo AJ, Yanovsky MJ, Botto JF (2010) Germination variation in Arabidopsis thaliana accessions under moderate osmotic and salt stresses. Ann Bot 106:833–842
Wan H, Chen L, Guo J, Li Q, Wen J, Yi B, Ma C, Tu J, Fu T, Shen J (2017) Genome-wide association study reveals the genetic architecture underlying salt tolerance-related traits in rapeseed (Brassica napus L). Front Plant Sci 8:593
Wan H, Wei Y, Qian J, Gao Y, Wen J, Yi B, Ma C, Tu J, Fu T, Shen J (2018) Association mapping of salt tolerance traits at germination stage of rapeseed (Brassica napus L). Euphytica 214:190
Wang L, Li Q, Lei Q, Feng C, Gao Y, Zheng X, Zhao Y, Wang Z, Kong J (2015) MzPIP2; 1: an aquaporin involved in radial water movement in both water uptake and transportation, altered the drought and salt tolerance of transgenic Arabidopsis. PLoS ONE 10:e0142446
Xe W, Basnayake BVS, Zhang H, Li G, Li W, Virk N, Mengiste T, Song F (2009) The Arabidopsis ATAF1, a NAC transcription factor, is a negative regulator of defense responses against necrotrophic fungal and bacterial pathogens. Mol Plant Microbe Interact 22:1227–1238
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer, New York
Wu H, Guo J, Wang C, Li K, Zhang X, Yang Z, Li M, Wang B (2019) An effective screening method and a reliable screening trait for salt tolerance of Brassica napus at the germination stage. Front Plant Sci 10:530
Yong HY, Wang C, Bancroft I, Li F, Wu X, Kitashiba H, Nishio T (2015) Identification of a gene controlling variation in the salt tolerance of rapeseed (Brassica napus L.). Planta 242:313–326
Yuan F, Guo J, Shabala S, Wang B (2019) Reproductive physiology of halophytes: current standing. Front Plant Sci 9:1954
Zhang R, Deng W, Yang L, Wang Y, Xiao F, He J, Lu K (2017) Genome-wide association study of root length and hypocotyl length at germination stage under saline conditions in Brassica napus. ScieAgri Sin 50:15–27
Zhang X, Ju HW, Chung MS, Huang P, Ahn SJ, Kim CS (2010) The RR-type MYB-like transcription factor, AtMYBL, is involved in promoting leaf senescence and modulates an abiotic stress response in Arabidopsis. Plant Cell Physiol 52:138–148
Zhang Y, Xu A, Lang L, Wang Y, Liu X, Liang F, Zhang B, Qin M, Dalelhan J, Huang Z (2018) Genetic mapping of a lobed-leaf gene associated with salt tolerance in Brassica napus L. Plant Sci 269:75–84
Zhou Q, Han D, Mason AS, Zhou C, Zheng W, Li Y, Wu C, Fu D, Huang Y (2017a) Earliness traits in rapeseed (Brassica napus): SNP loci and candidate genes identified by genome-wide association analysis. DNA Res 25:229–244
Zhou Q, Zhou C, Zheng W, Mason AS, Fan S, Wu C, Fu D, Huang Y (2017b) Genome-wide SNP markers based on SLAF-Seq uncover breeding traces in rapeseed (Brassica napus L.). Front Plant Sci 8:648
Acknowledgments
This research was funded by the National Natural Science Foundation of China, grant number 31860417.
Author information
Authors and Affiliations
Contributions
GMW and HK performed the experiments, analyses the data, and wrote the paper. QZ provides seed material. AAK, SK, AMS, and MF provide technical support in data analysis, critical reading, and suggestions regarding the manuscript. DF and HH conceived the project and supervised this study. All of the authors discussed the results and commented on the manuscript. ASM, revised the whole manuscript for the language.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
438_2020_1749_MOESM1_ESM.tif
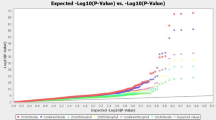
Supplementary Fig. S1 Quantile-quantile plot for the GWAS using a generalized linear model (GLM) and mixed linear model (MLM) based on salt stress susceptibility and salt stress tolerance indices in four seedling traits in Brassica napus. (A) Germination % GLM and MLM by stress susceptibility index (SSI) (B) Germination % GLM and MLM by stress tolerance index (STI) (C) root length of GLM and MLM by SSI (D) root length of GLM and MLM by STI (E) shoot dry weight of GLM and MLM by SSI (F) shoot dry weight of GLM and MLM by STI (G) seed vigor index of GLM and MLM by SSI and (H) seed vigor index of GLM and MLM by STI (TIF 16627 KB)
438_2020_1749_MOESM2_ESM.tif
Supplementary Fig. S2 Manhattan plots of genome-wide association studies (GWAS) using a generalized linear model (GLM) for four seedling traits related to salt stress of Brassica napus based on stress susceptibility index (SSI) and stress tolerance index (STI). The log 10 (P-value) is a measure of the significance with which a SNP is associated with a trait. The blue horizontal dotted lines represent the genome wide significance threshold of p < 0.05 after FDR correction. Each chromosome is represented by a different color. (A) germination % using SSI (B) germination % using STI (C) root length in SSI (D) root length in STI (E) shoot dry weight in SSI (F) shoot dry weight in STI (G) seed vigor index in SSI and (H) seed vigor index in STI (TIF 16077 KB)
438_2020_1749_MOESM3_ESM.tif
Supplementary Fig. S3 Manhattan plots of genome-wide association studies (GWAS) using a mixed linear model (MLM) for four seedling traits related to salt stress of Brassica napus based on stress susceptibility index (SSI) and stress tolerance index (STI). The log 10 (P-value) is a measure of the degree to which a SNP is significantly associated with a trait. The blue horizontal dotted lines represent the genome-wide significance threshold. Each chromosome is represented by a different color. (A) germination % using stress susceptibility index (SSI) (B) germination % using stress tolerance index (STI) (C) root length based on SSI (D) root length based on STI (E) shoot dry weight based on SSI (F) shoot dry weight based on STI (G) seed vigor index based on SSI (H) seed vigor index based on STI (TIF 16080 KB)
438_2020_1749_MOESM4_ESM.xlsx
Supplementary Table S1 Detailed information about the 228 lines along with their subgroup of Brassica napus (XLSX 21 KB)
438_2020_1749_MOESM5_ESM.xlsx
Supplementary Table S2 Phenotypic variations among the lines in pilot experiment under various salt treatments (XLSX 11 KB)
438_2020_1749_MOESM6_ESM.docx
Supplementary Table S3 Correlation coefficients (r) between germination and seedling traits under control/non-stress conditions (CK) above the diagonal in blue, while under salt stress below diagonal with pink color in rapeseed natural population (DOCX 14 KB)
Supplementary Table S4 SNPs associated with germination by GLM and MLM in Brassica napus (DOC 81 KB)
Supplementary Table S8 SNPs associated with RL and SVI by GLM and MLM in Brassica napus (DOCX 19 KB)
438_2020_1749_MOESM15_ESM.xlsx
Supplementary Table S12 Candidate genes highly associated with seedling traits of salt tolerance in Brassica napus (XLSX 51 KB)
438_2020_1749_MOESM16_ESM.xlsx
Supplementary Table S13 Salt related genes tagged by the associated SNPs in Brassica napus and homologous with Arabidopsis thaliana genes (XLSX 21 KB)
Rights and permissions
About this article
Cite this article
Wassan, G.M., Khanzada, H., Zhou, Q. et al. Identification of genetic variation for salt tolerance in Brassica napus using genome-wide association mapping. Mol Genet Genomics 296, 391–408 (2021). https://doi.org/10.1007/s00438-020-01749-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-020-01749-8