Abstract
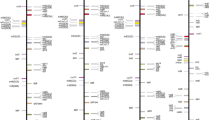
Plant mitochondrial genomes contain a large number of mitotype-specific sequences (MSS) which establish a mitochondrial genome structure distinct from other mitotypes. In rice, nine mitochondrial genomes have been sequenced, which provides us with the possibility of characterizing the MSS of rice and probing their relationship to cytoplasmic male sterility (CMS) in rice. We therefore analyzed the mitochondrial genomes of CW-CMS, LD-CMS, WA-CMS, N and Nipponbare lines, and found 57 MSS with sizes ranging from 102 to 5,745 bp, and with an aggregate length of 92.4 kb. The MSS account for more than 14.5 % of the rice mitochondrial genome and are a significant contributing factor in the variation of mitochondrial genome sizes. Of the MSS tested, 34 MSS exhibited polymorphism among rice lines, and 14 MSS were further confirmed as being specific to CMS. This includes nine MSS specific to sporophytic CMS, three specific to gametophytic CMS, and two shared by all types of CMS. Interestingly, except for CMS genes orf(H)79 and orf352 which are partly or fully overlapping with some MSS fragments, there are ten more open reading frames of unknown function that were detected in CMS-specific MSS, hinting at their possible roles in plant CMS. These novel findings provide us with potential new molecular tools to direct the breeding of CMS lines in hybrid rice breeding programs.




Similar content being viewed by others
References
Akagi H, Sakamoto M, Shinjyo C, Shimada H, Fujimura T (1994) A unique sequence located downstream from the rice mitochondrial apt6 may cause male sterility. Curr Genet 25:52–58
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Bentolila S, Stefanov S (2012) A reevaluation of rice mitochondrial evolution based on the complete sequence of male-fertile and male-sterile mitochondrial genomes. Plant Physiol 158:996–1017
Clifton SW, Minx P, Fauron CM, Gibson M, Allen JO, Sun H, Thompson M, Barbazuk WB, Kanuganti S, Tayloe C, Meyer L, Wilson RK, Newton KJ (2004) Sequence and comparative analysis of the maize NB mitochondrial genome. Plant Physiol 136:3486–3503
Fujii S, Kazama T, Yamada M, Toriyama K (2010) Discovery of global genomic re-organization based on comparison of two newly sequenced rice mitochondrial genomes with cytoplasmic male sterility-related genes. BMC Genomics 11:209
Fujii S, Toda T, Kikuchi S, Suzuki R, Yokoyama K, Tsuchida H, Yano K, Toriyama K (2011) Transcriptome map of plant mitochondria reveals islands of unexpected transcribed regions. BMC Genomics 12:279
Handa H (2003) The complete nucleotide sequence and RNA editing content of the mitochondrial genome of rapeseed (Brassica napus L.): comparative analysis of the mitochondrial genomes of rapeseed and Arabidopsis thaliana. Nucleic Acids Res 31:5907–5916
Hanson MR, Bentolila S (2004) Interactions of mitochondrial and nuclear genes that affect male gametophyte development. Plant Cell 16:S154–S169
Heazlewood JL, Howell KA, Whelan J, Millar AH (2003) Towards an analysis of the rice mitochondrial proteome. Plant Physiol 132:230–242
Igarashi K, Kazama T, Motomura K, Toriyama K (2013) Whole genomic sequencing of RT98 mitochondria derived from Oryza rufipogon and northern blot analysis to uncover a cytoplasmic male sterility-associated gene. Plant Cell Physiol 54:237–243
Itabashi E, Kazama T, Toriyama K (2009) Characterization of cytoplasmic male sterility of rice with Lead Rice cytoplasm in comparison with that with Chinsurah Boro II cytoplasm. Plant Cell Rep 28(2):233–239
Kim S, Lim H, Park S, Cho KH, Sung SK, Oh DG, Kim KT (2007) Identification of a novel mitochondrial genome type and development of molecular markers for cytoplasm classification in radish (Raphanus sativus L.). Theor Appl Genet 115:1137–1145
Kubo T, Mikami T (2007) Organization and variation of angiosperm mitochondrial genome. Physiol Plantarum 129:6–13
Li SQ, Tan YP, Wang K, Wan CX, Zhu YG (2008) Gametophytically alloplasmic CMS line of rice (Oryza sativa L.) with variant orfH79 haplotype corresponds to specific fertility restorer. Theor Appl Genet 117:1389–1397
Lilly JW, Harvey MJ (2001) Small, repetitive DNAs contribute significantly to the expanded mitochondrial genome of cucumber. Genetics 159:317–328
Liu HT, Cui P, Zhan KH, Lin Q, Zhuo GY, Guo XL, Ding F, Yang WL, Liu DC, Hu SN (2011) Comparative analysis of mitochondrial genomes between a wheat K-type cytoplasmic male sterility (CMS) line and its maintainer line. BMC Genomics 12:163
Luo DP, Xu H, Liu ZL, Guo JX, Li HY, Chen LT, Fang C, Zhang QY, Bai M, Yao N, Wu H, Wu H, Ji CH, Zheng HQ, Chen YL, Ye S, Li XY, Zhao XZ, Li RQ, Liu YG (2013) A detrimental mitochondrial-nuclear interaction causes cytoplasmic male sterility in rice. Nat Genet 45:573–577
Ngangkham U, Parida SK, De S, Raj Kumar KA, Singh AK, Singh NK, Mohapatra T (2010) Genic markers for wild abortive (WA) cytoplasm based male sterility and its fertility restoration in rice. Mol Breed 26:275–292
Notsu Y, Masood S, Nishikawa T, Kubo N, Akiduki G, Nakazono M, Hirai A, Kadowaki K (2002) The complete sequence of the rice (Oryza sativa L.) mitochondrial genome: frequent DNA sequence acquisition and loss during the evolution of flowering plants. Mol Genet Genomics 268:434–445
Palmer JD, Herbon LA (1988) Plant mitochondrial DNA evolves rapidly in structure, but slowly in sequence. J Mol Evol 28:87–97
Peng XJ, Wang K, Hu CF, Zhu YL, Wang T, Yang J, Tong JP, Li SQ, Zhu YG (2010) The mitochondrial gene orfH79 plays a critical role in impairing both male gametophyte development and root growth in CMS-Honglian rice. BMC Plant Biol 10:125–135
Rajendran N, Gandhimani R, Singh S, Palchamy K (2007) Development of a DNA marker for distinguishing CMS lines from fertile lines in rice (Oryza sativa L.). Euphytica 156:129–139
Rogers SO, Bendich AJ (1985) Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol Biol 5(2):69–76
Rohlf FJ (1992) NTSYS-pc: numerical taxonomy and multivariate analysis system, version 1.8. Exter Software, New York
Satoh M, Kubo T, Mikami T (2006) The Owen mitochondrial genome in sugar beet (Beta vulgaris L.): possible mechanisms of extensive rearrangements and the origin of the mitotype-unique regions. Theor Appl Genet 113:477–484
Tian XJ, Zheng J, Hu SN, Yu J (2006) The rice mitochondrial genomes and their variations. Plant Physiol 140:401–410
Wang ZH, Zou YJ, Li XY, Zhang QY, Chen L, Wu H, Su DH, Chen YL, Guo JX, Luo D, Long YM, Zhong Y, Liu YG (2006) Cytoplasmic male sterility of rice with boro II cytoplasm is caused by a cytotoxic peptide and is restored by two related PPR motif genes via distinct modes of mRNA silencing. Plant Cell 18:676–687
Wang K, Peng XJ, Ji YX, Yang PF, Zhu YG, Li SQ (2013) Gene, protein and network of male sterility in rice. Front Plant Sci 4:92
Wise RP, Fliss AE, Pring DR, Gengenbach BG (1987) URF13-T of T-cytoplasm maize mitochondria encodes a 13 KD polypeptide. Plant Mol Biol 9:121–126
Yi P, Wang L, Sun QP, Zhu YG (2002) Discovery of mitochondrial chimeric gene associated with cytoplasmic male sterility of HL-rice. Chin Sci Bull 47:744–747
Young EG, Hanson MR (1987) A fused mitochondrial gene associated with cytoplasmic male sterility is developmentally regulated. Cell 50:41–49
Zhang TW, Hu S, Zhang GY, Pan LL, Zhang XW, Almssallem IS, Yu J (2012) The organelle genomes of Hassawi rice (Oryza sativa L.) and its hybrid in Saudi Arabia: genome variation, rearrangement, and origins. PLoS ONE 7(7):e42041
Zhao HX, Li ZJ, Hu SW, Sun GL, Chang JJ, Zhang ZH (2010) Identification of cytoplasm types in rapeseed (Brassica napus L.) accessions by a multiplex PCR assay. Theor Appl Genet 121:643–650
Acknowledgments
This work was supported by the National Natural Science Foundation (31071391) and the 973 Program (2011CB100102) of China.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xie, H., Wang, J., Qian, M. et al. Mitotype-specific sequences related to cytoplasmic male sterility in Oryza species. Mol Breeding 33, 803–811 (2014). https://doi.org/10.1007/s11032-013-9993-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-013-9993-y