Abstract
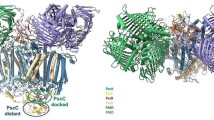
The TT1485 gene from Thermus thermophilus HB8 encodes a hypothetical protein of unknown function with about 20 sequence homologs of bacterial or archaeal origin. Together they form a family of uncharacterized proteins, the cluster of orthologous group COG3253. Using a combination of amino acid sequence analysis, three-dimensional structural studies and biochemical assays, we identified TT1485 as a novel heme-binding protein. The crystal structure reveals that this protein is a pentamer and each monomer exhibits a β-barrel fold. TT1485 is structurally similar to muconolactone isomerase, but this provided no functional clues. Amino acid sequence analysis revealed remote homology to a heme enzyme, chlorite dismutase. Strikingly, amino acid residues that are highly conserved in the homologous hypothetical proteins and chlorite dismutase cluster around a deep cavity on the surface of each monomer. Molecular modeling shows that the cavity can accommodate a heme group with a strictly conserved His as a heme ligand. TT1485 reconstituted with iron protoporphyrin IX chloride gave a low chlorite dismutase activity, indicating that TT1485 catalyzes a reaction other than chlorite degradation. The presence of a possible Fe–His–Asp triad in the heme proximal site suggests that TT1485 functions as a novel heme peroxidase to detoxify hydrogen peroxide within the cell.
Similar content being viewed by others
Abbreviations
- MAD:
-
multiple wavelength anomalous dispersion
- MLI:
-
muconolactone isomerase
- SeMet:
-
selenomethionine
References
S.J. Cordwell (1999) Arch. Microbiol. 172 269–279 Occurrence Handle10.1007/s002030050780 Occurrence Handle1:CAS:528:DyaK1MXnt1Gnurk%3D Occurrence Handle10550468
A. Osterman R. Overbeek (2003) Curr. Opin. Chem. Biol. 7 238–251 Occurrence Handle10.1016/S1367-5931(03)00027-9 Occurrence Handle1:CAS:528:DC%2BD3sXjtVKgsLY%3D Occurrence Handle12714058
C. Chothia A.M. Lesk (1986) EMBO J. 5 823–826 Occurrence Handle1:CAS:528:DyaL28XitVals7o%3D Occurrence Handle3709526
C.A. Orengo A.E. Todd J.M. Thornton (1999) Curr. Opin. Struct. Biol. 9 374–382 Occurrence Handle10.1016/S0959-440X(99)80051-7 Occurrence Handle1:CAS:528:DyaK1MXks1Wqs78%3D Occurrence Handle10361094
S.A. Teichmann A.G. Murzin C. Chothia (2001) Curr. Opin. Struct. Biol. 11 354–363 Occurrence Handle10.1016/S0959-440X(00)00215-3 Occurrence Handle1:CAS:528:DC%2BD3MXks1GqsLs%3D Occurrence Handle11406387
C. Zhang S.H. Kim (2003) Curr. Opin. Chem. Biol. 7 28–32 Occurrence Handle10.1016/S1367-5931(02)00015-7 Occurrence Handle1:CAS:528:DC%2BD3sXltlaqsA%3D%3D Occurrence Handle12547423
S. Yokoyama H. Hirota T. Kigawa T. Yabuki M. Shirouzu T. Terada Y. Ito Y. Matsuo Y. Kuroda Y. Nishimura Y. Kyogoku K. Miki R. Masui S. Kuramitsu (2000) Nat. Struct. Biol. 7 IssueIDSuppl 943–945 Occurrence Handle1:CAS:528:DC%2BD3cXnvVOgt7s%3D Occurrence Handle11103994
R.L. Tatusov D.A. Natale I.V. Garkavtsev T.A. Tatusova U.T. Shankavaram B.S. Rao B. Kiryutin M.Y. Galperin N.D. Fedorova E.V. Koonin (2001) Nucleic Acids Res. 29 22–28 Occurrence Handle10.1093/nar/29.1.22 Occurrence Handle1:CAS:528:DC%2BD3MXjtlWnsLo%3D Occurrence Handle11125040
D.M. LeMaster F.M. Richards (1985) Biochemistry 24 7263–7268 Occurrence Handle1:CAS:528:DyaL28XitlOm Occurrence Handle3910099
T. Kumasaka M. Yamamoto E. Yamashita H. Moriyama T. Ueki (2002) Structure (Camb) 10 1205–1210 Occurrence Handle10.1016/S0969-2126(02)00830-4 Occurrence Handle1:CAS:528:DC%2BD38XntVaisrk%3D
Z. Otwinowski W. Minor (1997) Methods Enzymol. 276 307–326 Occurrence Handle1:CAS:528:DyaK2sXivFehsbw%3D
N. CollaborativeComputational Project (1994) Acta Crystallogr D 50 760–763
T.C. Terwilliger J. Berendzen (1999) Acta Crystallogr. D 55 849–861 Occurrence Handle10.1107/S0907444999000839 Occurrence Handle10089316
T.C. Terwilliger (2000) Acta Crystallogr. D 56 965–972 Occurrence Handle10.1107/S0907444900005072 Occurrence Handle10944333
D.E. McRee (1999) J. Struct. Biol. 125 156–165 Occurrence Handle10.1006/jsbi.1999.4094 Occurrence Handle1:CAS:528:DyaK1MXis1Krt7k%3D Occurrence Handle10222271
A.T. Brunger P.D. Adams G.M. Clore W.L. DeLano P. Gros R.W. Grosse-Kunstleve J.S. Jiang J. Kuszewski M. Nilges N.S. Pannu R.J. Read L.M. Rice T. Simonson G.L. Warren (1998) Acta Crystallogr. D 54 905–921 Occurrence Handle10.1107/S0907444998003254 Occurrence Handle9757107
P.J. Kraulis (1991) J. Appl. Crystallogr. 24 946–950 Occurrence Handle10.1107/S0021889891004399
E.A. Merritt D.J. Bacon (1997) Methods Enzymol 277 505–524 Occurrence Handle1:CAS:528:DyaK2sXntFals7s%3D
J.A. Christopher (1998) SPOCK: the structural properties observation and calculation kit (program manual). The Center for Macromolecular Design Texas A & M University College Station Texas
P.K. Smith R.I. Krohn G.T. Hermanson A.K. Mallia F.H. Gartner M.D. Provenzano E.K. Fujimoto N.M. Goeke B.J. Olson D.C. Klenk (1985) Anal. Biochem. 150 76–85 Occurrence Handle10.1016/0003-2697(85)90442-7 Occurrence Handle1:CAS:528:DyaL2MXlsFKksL0%3D Occurrence Handle3843705
J.W. Priest S.L. Hajduk (1992) J. Biol. Chem. 267 20188–20195 Occurrence Handle1:CAS:528:DyaK38XlsVGrurg%3D Occurrence Handle1328195
E.A. Berry B.L. Trumpower (1987) Anal. Biochem. 161 1–15 Occurrence Handle10.1016/0003-2697(87)90643-9 Occurrence Handle1:CAS:528:DyaL2sXhvFaitrg%3D Occurrence Handle3578775
S.L. Hazen F.F. Hsu J.P. Gaut J.R. Crowley J.W. Heinecke (1999) Methods Enzymol. 300 88–105 Occurrence Handle1:CAS:528:DyaK1MXltVKg Occurrence Handle9919513
R.F. Beers SuffixJr. I.W. Sizer (1952) J. Biol. Chem. 195 133–140 Occurrence Handle1:CAS:528:DyaG38Xjt12juw%3D%3D Occurrence Handle14938361
C. Zhang S.H. Kim (2000) J. Mol. Biol. 299 1075–1089 Occurrence Handle10.1006/jmbi.2000.3678 Occurrence Handle1:CAS:528:DC%2BD3cXjvVSrsrc%3D Occurrence Handle10843859
J.S. Richardson (1981) Adv. Protein Chem. 34 167–339 Occurrence Handle1:CAS:528:DyaL3MXlsVGgs7c%3D Occurrence Handle7020376
L. Holm C. Sander (1998) Nucleic Acids Res. 26 316–319 Occurrence Handle10.1093/nar/26.1.316 Occurrence Handle1:CAS:528:DyaK1cXovVChsQ%3D%3D Occurrence Handle9399863
A.G. Murzin S.E. Brenner T. Hubbard C. Chothia (1995) J. Mol. Biol. 247 536–540 Occurrence Handle10.1006/jmbi.1995.0159 Occurrence Handle1:CAS:528:DyaK2MXltVGgsr4%3D Occurrence Handle7723011
S.K. Katti B.A. Katz H.W. Wyckoff (1989) J. Mol. Biol. 205 557–571 Occurrence Handle10.1016/0022-2836(89)90226-X Occurrence Handle1:CAS:528:DyaL1MXhtlWgsLs%3D Occurrence Handle2926818
S.F. Altschul T.L. Madden A.A. Schaffer J. Zhang Z. Zhang W. Miller D.J. Lipman (1997) Nucleic Acids Res. 25 3389–3402 Occurrence Handle1:CAS:528:DyaK2sXlvFyhu7w%3D Occurrence Handle9254694
J.D. Thompson D.G. Higgins T.J. Gibson (1994) Nucleic Acids Res. 22 4673–4680 Occurrence Handle1:CAS:528:DyaK2MXitlSgu74%3D Occurrence Handle7984417
K.B. Nicholas SuffixJr. N.H.B. D.W.I. Deerfield (1997) EMBNEW. NEWS 4 14
C.G. Ginkel Particlevan G.B. Rikken A.G. Kroon S.W. Kengen (1996) Arch. Microbiol. 166 321–326 Occurrence Handle10.1007/s002030050390 Occurrence Handle8929278
K. Stenklo H.D. Thorell H. Bergius R. Aasa T. Nilsson (2001) J. Biol. Inorg. Chem. 6 601–607 Occurrence Handle10.1007/s007750100237 Occurrence Handle1:CAS:528:DC%2BD3MXlsFOntL8%3D Occurrence Handle11472023
V. Kery D. Elleder J.P. Kraus (1995) Arch. Biochem. Biophys. 316 24–29 Occurrence Handle10.1006/abbi.1995.1005 Occurrence Handle1:CAS:528:DyaK2MXjsV2iu7w%3D Occurrence Handle7840623
J. Switala P.C. Loewen (2002) Arch. Biochem. Biophys. 401 145–154 Occurrence Handle10.1016/S0003-9861(02)00049-8 Occurrence Handle1:CAS:528:DC%2BD38Xks1Ggtro%3D Occurrence Handle12054464
A.E. Todd C.A. Orengo J.M. Thornton (2001) J. Mol. Biol. 307 1113–1143 Occurrence Handle10.1006/jmbi.2001.4513 Occurrence Handle1:CAS:528:DC%2BD3MXitlOju7Y%3D Occurrence Handle11286560
D.B. Goodin D.E. McRee (1993) Biochemistry 32 3313–3324 Occurrence Handle10.1021/bi00064a014 Occurrence Handle1:CAS:528:DyaK3sXhs1CitL0%3D Occurrence Handle8384877
G. Michal (Eds) (1999) Biochemical Pathways: An Atlas of Biochemistry and Molecular Biology Wiley and Spektrum New York, NY
K.G. Welinder (1992) Curr. Opin. Struct. Biol. 2 388–393 Occurrence Handle10.1016/0959-440X(92)90230-5 Occurrence Handle1:CAS:528:DyaK38XltVKrsLY%3D
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ebihara, A., Okamoto, A., Kousumi, Y. et al. Structure-based functional identification of a novel heme-binding protein from Thermus thermophilus HB8. J Struct Funct Genomics 6, 21–32 (2005). https://doi.org/10.1007/s10969-005-1103-x
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s10969-005-1103-x