Abstract
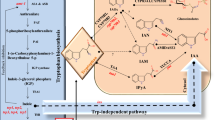
The plant auxin indole-3-acetic acid (IAA) plays critical roles in plant growth and development. There are two main strategies proposed for plant synthesis of IAA: the Trp-dependent (TD) and the Trp-independent (TI) pathways. Four TD pathways, namely the indole-3-acetamide pathway, the indole-3-pyruvic acid pathway, the tryptamine pathway and the indole-3-acetaldoxime pathway, have been postulated, identified and extensively studied. On the other hand, neither genes nor mutants involved in the TI pathway have been identified to date. Interestingly, some bacteria have auxin synthesis pathways that are similar to those in plants, indicating conserved biosynthetic mechanisms. Over the past few years, genetic, biochemical and molecular studies have greatly advanced our understanding of auxin biosynthesis. This review both summarizes recent advances in genetic and molecular knowledge and addresses the unsolved questions regarding auxin biosynthesis pathways in plants.


Similar content being viewed by others
References
Alarcon MV, Lloret PG, Salguero J (2014) Synergistic action of auxin and ethylene on root elongation inhibition is caused by a reduction of epidermal cell length. Plant Signal Behav 9:e28361
Bartel B (1997) Auxin biosynthesis. Annu Rev Plant Phys 48:49–64
Bartel B, Fink GR (1994) Differential regulation of an auxin-producing nitrilase gene family in Arabidopsis thaliana. Proc Natl Acad Sci USA 91(14):6649–6653
Bartling D, Seedorf M, Mithofer A, Weiler EW (1992) Cloning and expression of an Arabidopsis nitrilase which can convert indole-3-acetonitrile to the plant hormone, indole-3-acetic acid. Eur J Biochem 205(1):417–424
Brumos J, Alonso JM, Stepanova AN (2014) Genetic aspects of auxin biosynthesis and its regulation. Physiol Plant 151(1):3–12
Cai XT, Xu P, Zhao PX, Liu R, Yu LH, Xiang CB (2014) Arabidopsis ERF109 mediates cross-talk between jasmonic acid and auxin biosynthesis during lateral root formation. Nat Commun 5:5833
Casanova E, Trillas MI, Moysset L, Vainstein A (2005) Influence of rol genes in floriculture. Biotechnol Adv 23(1):3–39
Chandler JW (2009) Auxin as compere in plant hormone crosstalk. Planta 231(1):1–12
Chapman EJ, Greenham K, Castillejo C, Sartor R, Bialy A, Sun TP, Estelle M (2012) Hypocotyl transcriptome reveals auxin regulation of growth-promoting genes through GA-dependent and -independent pathways. PLoS ONE 7(5):e36210
Cheng Y, Dai X, Zhao Y (2006) Auxin biosynthesis by the YUCCA flavin monooxygenases controls the formation of floral organs and vascular tissues in Arabidopsis. Genes Dev 20(13):1790–1799
Cheng Y, Dai X, Zhao Y (2007) Auxin synthesized by the YUCCA flavin monooxygenases is essential for embryogenesis and leaf formation in Arabidopsis. Plant Cell 19(8):2430–2439
Choi YI, Noh EW, Kim HJ, Park WJ (2014) Differential regulation of cytokinin oxidase genes and cytokinin-induced auxin biosynthesis by cellular cytokinin level in transgenic poplars. Plant Cell Rep 33(10):1737–1744
Cooney TP, Nonhebel HM (1991) Biosynthesis of indole-3-acetic acid in tomato shoots: measurement, mass-spectral identification and incorporation of (−2)H from (−2)H2O into indole-3-acetic acid, d- and l-tryptophan, indole-3-pyruvate and tryptamine. Planta 184(3):368–376
Costacurta A, Keijers V, Vanderleyden J (1994) Molecular cloning and sequence analysis of an Azospirillum brasilense indole-3-pyruvate decarboxylase gene. Mol Gen Genet 243(4):463–472
Cui D, Zhao J, Jing Y, Fan M, Liu J, Wang Z, Xin W, Hu Y (2013) The arabidopsis IDD14, IDD15, and IDD16 cooperatively regulate lateral organ morphogenesis and gravitropism by promoting auxin biosynthesis and transport. PLoS Genet 9(9):e1003759
Dai X, Mashiguchi K, Chen Q, Kasahara H, Kamiya Y, Ojha S, DuBois J, Ballou D, Zhao Y (2013) The biochemical mechanism of auxin biosynthesis by an arabidopsis YUCCA flavin-containing monooxygenase. J Biol Chem 288(3):1448–1457
De Luca V, Fernandez JA, Campbell D, Kurz WG (1988) Developmental regulation of enzymes of indole alkaloid biosynthesis in Catharanthus roseus. Plant Physiol 86(2):447–450
Delarue M, Prinsen E, Onckelen HV, Caboche M, Bellini C (1998) Sur2 mutations of Arabidopsis thaliana define a new locus involved in the control of auxin homeostasis. Plant J 14(5):603–611
Dello Ioio R, Nakamura K, Moubayidin L, Perilli S, Taniguchi M, Morita MT, Aoyama T, Costantino P, Sabatini S (2008) A genetic framework for the control of cell division and differentiation in the root meristem. Science 322(5906):1380–1384
Ehlert B, Schottler MA, Tischendorf G, Ludwig-Muller J, Bock R (2008) The paramutated SULFUREA locus of tomato is involved in auxin biosynthesis. J Exp Bot 59(13):3635–3647
Eklund DM, Staldal V, Valsecchi I, Cierlik I, Eriksson C, Hiratsu K, Ohme-Takagi M, Sundstrom JF, Thelander M, Ezcurra I, Sundberg E (2010) The Arabidopsis thaliana STYLISH1 protein acts as a transcriptional activator regulating auxin biosynthesis. Plant Cell 22(2):349–363
Enders TA, Strader LC (2015) Auxin activity: past, present, and future. Am J Bot 102(2):180–196
Epstein E, Cohen J, Slovin J (2002) The biosynthetic pathway for indole-3-acetic acid changes during tomato fruit development. Plant Growth Regul 38:15–20
Facchini PJ, Huber-Allanach KL, Tari LW (2000) Plant aromatic l-amino acid decarboxylases: evolution, biochemistry, regulation, and metabolic engineering applications. Phytochemistry 54(2):121–138
Franklin KA, Lee SH, Patel D, Kumar SV, Spartz AK, Gu C, Ye S, Yu P, Breen G, Cohen JD, Wigge PA, Gray WM (2011) Phytochrome-interacting factor 4 (PIF4) regulates auxin biosynthesis at high temperature. Proc Natl Acad Sci USA 108(50):20231–20235
Gaudin V, Jouanin L (1995) Expression of Agrobacterium rhizogenes auxin biosynthesis genes in transgenic tobacco plants. Plant Mol Biol 28(1):123–136
Gray WM, Ostin A, Sandberg G, Romano CP, Estelle M (1998) High temperature promotes auxin-mediated hypocotyl elongation in Arabidopsis. Proc Natl Acad Sci USA 95(12):7197–7202
Grubb CD, Zipp BJ, Ludwig-Muller J, Masuno MN, Molinski TF, Abel S (2004) Arabidopsis glucosyltransferase UGT74B1 functions in glucosinolate biosynthesis and auxin homeostasis. Plant J 40(6):893–908
Guillet G, Poupart J, Basurco J, De Luca V (2000) Expression of tryptophan decarboxylase and tyrosine decarboxylase genes in tobacco results in altered biochemical and physiological phenotypes. Plant Physiol 122(3):933–943
Harrison E, Burbidge A, Okyere JP, Thompson AJ, Taylor IB (2011) Identification of the tomato ABA-deficient mutant sitiens as a member of the ABA-aldehyde oxidase gene family using genetic and genomic analysis. Plant Growth Regul 64(3):301–309
He Y, Zhao Y (2015) A key link between jasmonic acid signaling and auxin biosynthesis. Sci Ch Life Sci 3:014
He W, Brumos J, Li H, Ji Y, Ke M, Gong X, Zeng Q, Li W, Zhang X, An F, Wen X, Li P, Chu J, Sun X, Yan C, Yan N, Xie DY, Raikhel N, Yang Z, Stepanova AN, Alonso JM, Guo H (2011) A small-molecule screen identifies L-kynurenine as a competitive inhibitor of TAA1/TAR activity in ethylene-directed auxin biosynthesis and root growth in Arabidopsis. Plant Cell 23(11):3944–3960
Hentrich M, Bottcher C, Duchting P, Cheng Y, Zhao Y, Berkowitz O, Masle J, Medina J, Pollmann S (2013) The jasmonic acid signaling pathway is linked to auxin homeostasis through the modulation of YUCCA8 and YUCCA9 gene expression. Plant J 74(4):626–637
Huang J, Yue J, Hu X (2014) Origin of plant auxin biosynthesis in charophyte algae: a reply to Wang et al. Trends Plant Sci 19(12):743
Hull AK, Celenza JL (2000) Bacterial expression and purification of the Arabidopsis NADPH-cytochrome P450 reductase ATR2. Protein Express Purif 18(3):310–315
Hull AK, Vij R, Celenza JL (2000) Arabidopsis cytochrome P450 s that catalyze the first step of tryptophan-dependent indole-3-acetic acid biosynthesis. Proc Natl Acad Sci USA 97(5):2379–2384
Ishida Y, Hayashi K, Soeno K, Asami T, Nakamura S, Suzuki M, Nakamura A, Shimada Y (2014) Analysis of a putative auxin biosynthesis inhibitor, indole-3-oxoethylphosphonic acid, in Arabidopsis. Biosci Biotechnol Biochem 78(1):67–70
Ismond KP, Dolferus R, de Pauw M, Dennis ES, Good AG (2003) Enhanced low oxygen survival in Arabidopsis through increased metabolic flux in the fermentative pathway. Plant Physiol 132(3):1292–1302
King JJ, Stimart DP, Fisher RH, Bleecker AB (1995) A mutation altering auxin homeostasis and plant morphology in Arabidopsis. Plant Cell 7(12):2023–2037
Klee H, Horsch R, Hinchee M, Hein M, Hoffmann N (1987) The effects of overproduction of two Agrobacterium tumefaciens T-DNA auxin biosynthetic gene products in transgenic petunia plants. Genes Dev 1(1):86–96
Korasick DA, Enders TA, Strader LC (2013) Auxin biosynthesis and storage forms. J Exp Bot 64(9):2541–2555
Kriechbaumer V, Wang P, Hawes C, Abell BM (2012) Alternative splicing of the auxin biosynthesis gene YUCCA4 determines its subcellular compartmentation. Plant J 70(2):292–302
Kürsteiner O, Dupuis I, Kuhlemeier C (2003) The pyruvate decarboxylase1 gene of Arabidopsis is required during anoxia but not other environmental stresses. Plant Physiol 132(2):968–978
Last RL, Bissinger PH, Mahoney DJ, Radwanski ER, Fink GR (1991) Tryptophan mutants in Arabidopsis: the consequences of duplicated tryptophan synthase beta genes. Plant Cell 3(4):345–358
Lehmann T, Hoffmann M, Hentrich M, Pollmann S (2010) Indole-3-acetamide-dependent auxin biosynthesis: a widely distributed way of indole-3-acetic acid production? Eur J Cell Biol 89(12):895–905
Lemcke K, Prinsen E, van Onckelen H, Schmulling T (2000) The ORF8 gene product of Agrobacterium rhizogenes TL-DNA has tryptophan 2-monooxygenase activity. Mol Plant Microbe Interact 13(7):787–790
Li J, Last R (1996) The Arabidopsis thaliana trp5 mutant has a feedback-resistant anthranilate synthase and elevated soluble tryptophan. Plant Physiol 110(1):51–59
Li J, Chen S, Zhu L, Last RL (1995) Isolation of cDNAs encoding the tryptophan pathway enzyme indole-3-glycerol phosphate synthase from Arabidopsis thaliana. Plant Physiol 108(2):877–878
Li L, Ljung K, Breton G, Schmitz RJ, Pruneda-Paz J, Cowing-Zitron C, Cole BJ, Ivans LJ, Pedmale UV, Jung HS, Ecker JR, Kay SA, Chory J (2012) Linking photoreceptor excitation to changes in plant architecture. Genes Dev 26(8):785–790
Magie AR, Wilson EE, Kosuge T (1963) Indoleacetamide as an intermediate in the synthesis of indoleacetic acid in Pseudomonas savastanoi. Science 141(3587):1281–1282
Maharjan PM, Dilkes BP, Fujioka S, Pencik A, Ljung K, Burow M, Halkier BA, Choe S (2014) Arabidopsis gulliver1/SUPERROOT2-7 identifies a metabolic basis for auxin and brassinosteroid synergy. Plant J 80(5):797–808
Mano Y, Nemoto K (2012) The pathway of auxin biosynthesis in plants. J Exp Bot 63(8):2853–2872
Mano Y, Nemoto K, Suzuki M, Seki H, Fujii I, Muranaka T (2010) The AMI1 gene family: indole-3-acetamide hydrolase functions in auxin biosynthesis in plants. J Exp Bot 61(1):25–32
Mashiguchi K, Tanaka K, Sakai T, Sugawara S, Kawaide H, Natsume M, Hanada A, Yaeno T, Shirasu K, Yao H, McSteen P, Zhao Y, Hayashi K, Kamiya Y, Kasahara H (2011) The main auxin biosynthesis pathway in Arabidopsis. Proc Natl Acad Sci USA 108(45):18512–18517
Michalczuk L, Ribnicky DM, Cooke TJ, Cohen JD (1992) Regulation of indole-3-acetic acid biosynthetic pathways in carrot cell cultures. Plant Physiol 100(3):1346–1353
Mikkelsen MD, Hansen CH, Wittstock U, Halkier BA (2000) Cytochrome P450 CYP79B2 from Arabidopsis catalyzes the conversion of tryptophan to indole-3-acetaldoxime, a precursor of indole glucosinolates and indole-3-acetic acid. J Biol Chem 275(43):33712–33717
Nafisi M, Goregaoker S, Botanga CJ, Glawischnig E, Olsen CE, Halkier BA, Glazebrook J (2007) Arabidopsis cytochrome P450 monooxygenase 71A13 catalyzes the conversion of indole-3-acetaldoxime in camalexin synthesis. Plant Cell 19(6):2039–2052
Nemoto K, Hara M, Goto S, Kasai K, Seki H, Suzuki M, Oka A, Muranaka T, Mano Y (2009) The aux1 gene of the Ri plasmid is sufficient to confer auxin autotrophy in tobacco BY-2 cells. J Plant Physiol 166(7):729–738
Neu D, Lehmann T, Elleuche S, Pollmann S (2007) Arabidopsis amidase 1, a member of the amidase signature family. FEBS J 274(13):3440–3451
Nishimura T, Hayashi K, Suzuki H, Gyohda A, Takaoka C, Sakaguchi Y, Matsumoto S, Kasahara H, Sakai T, Kato J, Kamiya Y, Koshiba T (2014) Yucasin is a potent inhibitor of YUCCA, a key enzyme in auxin biosynthesis. Plant J 77(3):352–366
Niyogi KK, Last RL, Fink GR, Keith B (1993) Suppressors of trp1 fluorescence identify a new arabidopsis gene, TRP4, encoding the anthranilate synthase beta subunit. Plant Cell 5(9):1011–1027
Normanly J (1997) Auxin metabolism. Physiol Plant 100(3):431–442
Normanly J (2010) Approaching cellular and molecular resolution of auxin biosynthesis and metabolism. Cold Spring Harb Perspect Biol 2(1):a001594
Normanly J, Cohen JD, Fink GR (1993) Arabidopsis thaliana auxotrophs reveal a tryptophan-independent biosynthetic pathway for indole-3-acetic acid. Proc Natl Acad Sci USA 90(21):10355–10359
Ouyang J, Shao X, Li J (2000) Indole-3-glycerol phosphate, a branchpoint of indole-3-acetic acid biosynthesis from the tryptophan biosynthetic pathway in Arabidopsis thaliana. Plant J 24(3):327–333
Pacheco-Villalobos D, Sankar M, Ljung K, Hardtke CS (2013) Disturbed local auxin homeostasis enhances cellular anisotropy and reveals alternative wiring of auxin-ethylene crosstalk in Brachypodium distachyon seminal roots. PLoS Genet 9(6):e1003564
Patten CL, Glick BR (1996) Bacterial biosynthesis of indole-3-acetic acid. Can J Microbiol 42(3):207–220
Patten CL, Blakney AJ, Coulson TJ (2013) Activity, distribution and function of indole-3-acetic acid biosynthetic pathways in bacteria. Crit Rev Microbiol 39(4):395–415
Phillips KA, Skirpan AL, Liu X, Christensen A, Slewinski TL, Hudson C, Barazesh S, Cohen JD, Malcomber S, McSteen P (2011) Vanishing tassel2 encodes a grass-specific tryptophan aminotransferase required for vegetative and reproductive development in maize. Plant Cell 23(2):550–566
Pinon V, Prasad K, Grigg SP, Sanchez-Perez GF, Scheres B (2013) Local auxin biosynthesis regulation by PLETHORA transcription factors controls phyllotaxis in Arabidopsis. Proc Natl Acad Sci USA 110(3):1107–1112
Piotrowski M, Schonfelder S, Weiler EW (2001) The Arabidopsis thaliana isogene NIT4 and its orthologs in tobacco encode beta-cyano-l-alanine hydratase/nitrilase. J Biol Chem 276(4):2616–2621
Pollmann S, Muller A, Piotrowski M, Weiler EW (2002) Occurrence and formation of indole-3-acetamide in Arabidopsis thaliana. Planta 216(1):155–161
Pollmann S, Neu D, Weiler EW (2003) Molecular cloning and characterization of an amidase from Arabidopsis thaliana capable of converting indole-3-acetamide into the plant growth hormone, indole-3-acetic acid. Phytochemistry 62(3):293–300
Pollmann S, Neu D, Lehmann T, Berkowitz O, Schafer T, Weiler EW (2006) Subcellular localization and tissue specific expression of amidase 1 from Arabidopsis thaliana. Planta 224(6):1241–1253
Quittenden LJ, Davies NW, Smith JA, Molesworth PP, Tivendale ND, Ross JJ (2009) Auxin biosynthesis in pea: characterization of the tryptamine pathway. Plant Physiol 151(3):1130–1138
Radwanski ER, Last RL (1995) Tryptophan biosynthesis and metabolism: biochemical and molecular genetics. Plant Cell 7(7):921–934
Radwanski ER, Barczak AJ, Last RL (1996) Characterization of tryptophan synthase alpha subunit mutants of Arabidopsis thaliana. Mol Gen Genet MGG 253(3):353–361
Rapparini FCJ, Slovin JP (1999) Indole-3-acetic acid biosynthesis in Lemnagibba studied using stable isotope labeled anthranilate and tryptophan. Plant Growth Regul 27:139–144
Reverberi M, Fanelli C, Zjalic S, Briganti S, Picardo M, Ricelli A, Fabbri AA (2005) Relationship among lipoperoxides, jasmonates and indole-3-acetic acid formation in potato tuber after wounding. Free Radic Res 39(6):637–647
Ribnicky DM, Ilic N, Cohen JD, Cooke TJ (1996) The effects of exogenous auxins on endogenous indole-3-acetic acid metabolism (the implications for carrot somatic embryogenesis). Plant Physiol 112(2):549–558
Ribnicky DM, Cohen JD, Hu WS, Cooke TJ (2002) An auxin surge following fertilization in carrots: a mechanism for regulating plant totipotency. Planta 214(4):505–509
Rizzardi K, Landberg K, Nilsson L, Ljung K, Sundas-Larsson A (2011) TFL2/LHP1 is involved in auxin biosynthesis through positive regulation of YUCCA genes. Plant J 65(6):897–906
Ross JJ, Tivendale ND, Davidson SE, Reid JB, Davies NW, Quittenden LJ, Smith JA (2012) A mutation affecting the synthesis of 4-chloroindole-3-acetic acid. Plant Signal Behav 7(12):1533–1536
Roumeliotis E, Visser RG, Bachem CW (2012) A crosstalk of auxin and GA during tuber development. Plant Signal Behav 7(10):1360–1363
Schneider EA, Gibson RA, Wightman F (1972) Biosynthesis and metabolism of indol-3yl-acetic acid I.The native indoles of barley and tomato shoots. J Exp Bot 23(1):152–170
Seo M, Akaba S, Oritani T, Delarue M, Bellini C, Caboche M, Koshiba T (1998) Higher activity of an aldehyde oxidase in the auxin-overproducing superroot1 mutant of Arabidopsis thaliana. Plant Physiol 116(2):687–693
Seo M, Aoki H, Koiwai H, Kamiya Y, Nambara E, Koshiba T (2004) Comparative studies on the Arabidopsis aldehyde oxidase (AAO) gene family revealed a major role of AAO3 in ABA biosynthesis in seeds. Plant Cell Physiol 45(11):1694–1703
Shi H, Chen L, Ye T, Liu X, Ding K, Chan Z (2014) Modulation of auxin content in Arabidopsis confers improved drought stress resistance. Plant Physio Biochem 82:209–217
Simon S, Petrasek J (2011) Why plants need more than one type of auxin. Plant Sci 180(3):454–460
Sitbon F, Astot C, Edlund A, Crozier A, Sandberg G (2000) The relative importance of tryptophan-dependent and tryptophan-independent biosynthesis of indole-3-acetic acid in tobacco during vegetative growth. Planta 211(5):715–721
Smith JK, Schloss JV, Mazur BJ (1989) Functional expression of plant acetolactate synthase genes in Escherichia coli. Proc Natl Acad Sci USA 86(11):4179–4183
Soeno K, Goda H, Ishii T, Ogura T, Tachikawa T, Sasaki E, Yoshida S, Fujioka S, Asami T, Shimada Y (2010) Auxin biosynthesis inhibitors, identified by a genomics-based approach, provide insights into auxin biosynthesis. Plant Cell Physiol 51(4):524–536
Songstad DD, De Luca V, Brisson N, Kurz WG, Nessler CL (1990) High levels of tryptamine accumulation in transgenic tobacco expressing tryptophan decarboxylase. Plant Physiol 94(3):1410–1413
Spaepen S, Versees W, Gocke D, Pohl M, Steyaert J, Vanderleyden J (2007) Characterization of phenylpyruvate decarboxylase, involved in auxin production of Azospirillum brasilense. J Bacteriol 189(21):7626–7633
Staldal V, Cierlik I, Chen S, Landberg K, Baylis T, Myrenas M, Sundstrom JF, Eklund DM, Ljung K, Sundberg E (2012) The Arabidopsis thaliana transcriptional activator STYLISH1 regulates genes affecting stamen development, cell expansion and timing of flowering. Plant Mol Biol 78(6):545–559
Stepanova AN, Hoyt JM, Hamilton AA, Alonso JM (2005) A Link between ethylene and auxin uncovered by the characterization of two root-specific ethylene-insensitive mutants in Arabidopsis. Plant Cell 17(8):2230–2242
Stepanova AN, Robertson-Hoyt J, Yun J, Benavente LM, Xie DY, Dolezal K, Schlereth A, Jurgens G, Alonso JM (2008) TAA1-mediated auxin biosynthesis is essential for hormone crosstalk and plant development. Cell 133(1):177–191
Stepanova AN, Yun J, Robles LM, Novak O, He W, Guo H, Ljung K, Alonso JM (2011) The Arabidopsis YUCCA1 flavin monooxygenase functions in the indole-3-pyruvic acid branch of auxin biosynthesis. Plant Cell 23(11):3961–3973
Stirk WA, Novák O, Hradecká V (2009) Endogenous cytokinins, auxins and abscisic acid in Ulvafasciata (Chlorophyta) and Dictyotahumifusa (Phaeophyta): towards understanding their biosynthesis and homoeostasis. Eur J Phycol 44(2):231–240
Stone SL, Braybrook SA, Paula SL, Kwong LW, Meuser J, Pelletier J, Hsieh TF, Fischer RL, Goldberg RB, Harada JJ (2008) Arabidopsis LEAFY COTYLEDON2 induces maturation traits and auxin activity: implications for somatic embryogenesis. Proc Natl Acad Sci USA 105(8):3151–3156
Su YH, Liu YB, Zhang XS (2011) Auxin-cytokinin interaction regulates meristem development. Mol Plant 4(4):616–625
Sugawara S, Hishiyama S, Jikumaru Y, Hanada A, Nishimura T, Koshiba T, Zhao Y, Kamiya Y, Kasahara H (2009) Biochemical analyses of indole-3-acetaldoxime-dependent auxin biosynthesis in Arabidopsis. Proc Natl Acad Sci USA 106(13):5430–5435
Sun J, Qi L, Li Y, Chu J, Li C (2012) PIF4-mediated activation of YUCCA8 expression integrates temperature into the auxin pathway in regulating arabidopsis hypocotyl growth. PLoS Genet 8(3):e1002594
Suzuki H, Yokokura J, Ito T, Arai R, Yokoyama C, Toshima H, Nagata S, Asami T, Suzuki Y (2014) Biosynthetic pathway of the phytohormone auxin in insects and screening of its inhibitors. Insect Biochem Mol Biol 53:66–72
Sztein AE, Ilic N, Cohen JD, Cooke TJ (2002) Indole-3-acetic acid biosynthesis in isolated axes from germinating bean seeds: the effect of wounding on the biosynthetic pathway. Plant Growth Regul 36(3):201–207
Tao Y, Ferrer JL, Ljung K, Pojer F, Hong F, Long JA, Li L, Moreno JE, Bowman ME, Ivans LJ, Cheng Y, Lim J, Zhao Y, Ballare CL, Sandberg G, Noel JP, Chory J (2008) Rapid synthesis of auxin via a new tryptophan-dependent pathway is required for shade avoidance in plants. Cell 133(1):164–176. doi:10.1016/j.cell.2008.01.049
Teale WD, Paponov IA, Palme K (2006) Auxin in action: signalling, transport and the control of plant growth and development. Nat Rev Mol Cell Biol 7(11):847–859
Tivendale ND, Davies NW, Molesworth PP, Davidson SE, Smith JA, Lowe EK, Reid JB, Ross JJ (2010) Reassessing the role of N-hydroxytryptamine in auxin biosynthesis. Plant Physiol 154(4):1957–1965
Tivendale ND, Ross JJ, Cohen JD (2014) The shifting paradigms of auxin biosynthesis. Trends Plant Sci 19(1):44–51
Torrens-Spence MP, Liu P, Ding H, Harich K, Gillaspy G, Li J (2013) Biochemical evaluation of the decarboxylation and decarboxylation-deamination activities of plant aromatic amino acid decarboxylases. J Biol Chem 288(4):2376–2387
Trigueros M, Navarrete-Gomez M, Sato S, Christensen SK, Pelaz S, Weigel D, Yanofsky MF, Ferrandiz C (2009) The NGATHA genes direct style development in the Arabidopsis gynoecium. Plant Cell 21(5):1394–1409
Ursache R, Miyashima S, Chen Q, Vaten A, Nakajima K, Carlsbecker A, Zhao Y, Helariutta Y, Dettmer J (2014) Tryptophan-dependent auxin biosynthesis is required for HD-ZIP III-mediated xylem patterning. Development 141(6):1250–1259
Vandenbussche F, Callebert P, Zadnikova P, Benkova E, Van Der Straeten D (2013) Brassinosteroid control of shoot gravitropism interacts with ethylene and depends on auxin signaling components. Am J Bot 100(1):215–225
Vorwerk S, Biernacki S, Hillebrand H, Janzik I, Muller A, Weiler EW, Piotrowski M (2001) Enzymatic characterization of the recombinant Arabidopsis thaliana nitrilase subfamily encoded by the NIT2/NIT1/NIT3-gene cluster. Planta 212(4):508–516
Wang C, Liu Y, Li SS, Han GZ (2014) Origin of plant auxin biosynthesis in charophyte algae. Trends Plant Sci 19(12):741–743
Wang B, Chu J, Yu T, Xu Q, Sun X, Yuan J, Xiong G, Wang G, Wang Y, Li J (2015) Tryptophan-independent auxin biosynthesis contributes to early embryogenesis in Arabidopsis. Proc Natl Acad Sci USA 112(15):4821–4826
Winter A (1966) A hypothetical route for the biogenisis of IAA. Planta 71(3):229–239. doi:10.1007/BF00384885
Wojcikowska B, Jaskola K, Gasiorek P, Meus M, Nowak K, Gaj MD (2013) LEAFY COTYLEDON2 (LEC2) promotes embryogenic induction in somatic tissues of Arabidopsis, via YUCCA-mediated auxin biosynthesis. Planta 238(3):425–440
Won C, Shen X, Mashiguchi K, Zheng Z, Dai X, Cheng Y, Kasahara H, Kamiya Y, Chory J, Zhao Y (2011) Conversion of tryptophan to indole-3-acetic acid by TRYPTOPHAN AMINOTRANSFERASES OF ARABIDOPSIS and YUCCAs in Arabidopsis. Proc Natl Acad Sci USA 108(45):18518–18523
Woodward AW, Bartel B (2005) Auxin: regulation, action, and interaction. Ann Bot 95(5):707–735
Wright AD, Sampson MB, Neuffer MG, Michalczuk L, Slovin JP, Cohen JD (1991) Indole-3-acetic acid biosynthesis in the mutant maize orange pericarp, a tryptophan auxotroph. Science 254(5034):998–1000
Yamada M, Greenham K, Prigge MJ, Jensen PJ, Estelle M (2009) The TRANSPORT INHIBITOR RESPONSE2 gene is required for auxin synthesis and diverse aspects of plant development. Plant Physiol 151(1):168–179
Yang ZB, Geng X, He C, Zhang F, Wang R, Horst WJ, Ding Z (2014) TAA1-regulated local auxin biosynthesis in the root-apex transition zone mediates the aluminum-induced inhibition of root growth in Arabidopsis. Plant Cell 26(7):2889–2904
Yoshimitsu Y, Tanaka K, Fukuda W, Asami T, Yoshida S, Hayashi K, Kamiya Y, Jikumaru Y, Shigeta T, Nakamura Y, Matsuo T, Okamoto S (2011) Transcription of DWARF4 plays a crucial role in auxin-regulated root elongation in addition to brassinosteroid homeostasis in Arabidopsis thaliana. PLoS ONE 6(8):e23851
Yu P, Hegeman AD, Cohen JD (2014) A facile means for the identification of indolic compounds from plant tissues. Plant J 79(6):1065–1075
Yue J, Hu X, Huang J (2014) Origin of plant auxin biosynthesis. Trends Plant Sci 19:764–770
Zdunek-Zastocka E (2008) Molecular cloning, characterization and expression analysis of three aldehyde oxidase genes from Pisum sativum L. Plant Physiol Biochem PPB/Societe francaise de physiologie vegetale 46(1):19–28
Zhang R, Wang B, Ouyang J, Li J, Wang Y (2008) Arabidopsis indole synthase, a homolog of tryptophan synthase alpha, is an enzyme involved in the Trp-independent indole-containing metabolite biosynthesis. J Integr Plant Biol 50(9):1070–1077
Zhao Y (2010) Auxin biosynthesis and its role in plant development. Ann Rev Plant Biol 61:49–64
Zhao Y (2014) Auxin biosynthesis. Arabidopsis B 12:e0173. doi:10.1199/tab.0173
Zhao Y, Christensen SK, Fankhauser C, Cashman JR, Cohen JD, Weigel D, Chory J (2001) A role for flavin monooxygenase-like enzymes in auxin biosynthesis. Science 291(5502):306–309
Zhao Y, Hull AK, Gupta NR, Goss KA, Alonso J, Ecker JR, Normanly J, Chory J, Celenza JL (2002) Trp-dependent auxin biosynthesis in Arabidopsis: involvement of cytochrome P450 s CYP79B2 and CYP79B3. Genes Dev 16(23):3100–3112
Zheng Z, Guo Y, Novak O, Dai X, Zhao Y, Ljung K, Noel JP, Chory J (2013) Coordination of auxin and ethylene biosynthesis by the aminotransferase VAS1. Nat Chem Biol 9(4):244–246
Zhou ZY, Zhang CG, Wu L, Chai J, Wang M, Jha A, Jia PF, Cui SJ, Yang M, Chen R, Guo GQ (2011) Functional characterization of the CKRC1/TAA1 gene and dissection of hormonal actions in the Arabidopsis root. Plant J 66(3):516–527
Zhu J, Zhang KX, Wang WS, Gong W, Liu WC, Chen HG, Xu HH, Lu YT (2015) Low temperature inhibits root growth by reducing auxin accumulation via ARR1/12. Plant Cell Physiol 56(4):727–736
Ziegler DM (1990) Flavin-containing monooxygenases: enzymes adapted for multisubstrate specificity. Trends Pharmacol Sci 11(8):321–324
Acknowledgments
This work was supported by Grants from the Chinese National Science Foundation (31030045 and 31371431).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Di, DW., Zhang, C., Luo, P. et al. The biosynthesis of auxin: how many paths truly lead to IAA?. Plant Growth Regul 78, 275–285 (2016). https://doi.org/10.1007/s10725-015-0103-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-015-0103-5