Abstract
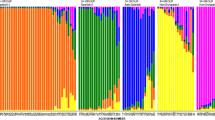
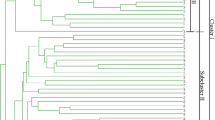
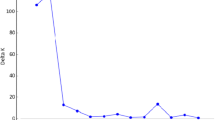
The genetic variability of 14 wild Prunus armeniaca populations was investigated using morphological analysis and inter-simple sequence repeat markers. 10 morphological characters revealed a high level variation, especially Fruit number, Fruit weight, Seed weight and Tree height. Totally, 15 selected primers generated 155 loci, with an average of 10.3 bands per primer. Nei’s gene diversity (H e ) and Shannon’s index of diversity (I) were fairly high at the species level (H e = 0.2741, I = 0.4220). High molecular and morphological variability indicated that wild apricots in the Ili Valley still maintained a relatively high level of diversity. The G ST of 0.2275 revealed a low level of genetic differentiation among populations, and genetic variation mainly resided within populations (81.51 %), which was identified with the moderate gene flow value (N m = 1.6974). The relatively high intraspecific genetic diversity and low inter-population genetic differentiation was largely attributed to long-distance dispersal of pollen, continuous distribution of populations and the self-incompatible breeding system.


Similar content being viewed by others
References
Asma BM, Ozturk K (2005) Analysis of morphological, pomological and yield characteristics of some apricot germplasm in Turkey. Genet Resour Crop Evol 52(3):305–313
Ballester J, de Vicente MC (1998) Determination of F-1 hybrid seed purity in pepper using PCR-based markers. Euphytica 103(2):223–226. doi:10.1023/a:1018372523343
Eriksson L, Johansson E, Kettaneh-Wold N, Wold S (1999) Introduction to multi- and megavariate data analysis using projection methods (PCA & PLS). Umetrics AB, Umea
Excoffier L, Smouse PE, Quattro JM (1992) Aanlysis of molecular variance inferred from metric distances among DNA haplotypes—application to human mitochondrial-DNA restriction data. Genetics 131(2):479–491
Ganopoulos IV, Kazantzis K, Chatzicharisis I, Karayiannis I, Tsaftaris AS (2011) Genetic diversity, structure and fruit trait associations in Greek sweet cherry cultivars using microsatellite based (SSR/ISSR) and morpho-physiological markers. Euphytica 181(2):237–251. doi:10.1007/s10681-011-0416-z
Godoy JA, Jordano P (2001) Seed dispersal by animals: exact identification of source trees with endocarp DNA microsatellites. Mol Ecol 10(9):2275–2283. doi:10.1046/j.0962-1083.2001.01342.x
He TM, Chen XS, Xu Z, Gao JS, Lin PJ, Liu W, Liang Q, Wu Y (2007) Using SSR markers to determine the population genetic structure of wild apricot (Prunus armeniaca L.) in the Ily Valley of West China. Genet Resour Crop Evol 54(3):563–572. doi:10.1007/s10722-006-0013-5
Hormaza JI (2002) Molecular characterization and similarity relationships among apricot (Prunus armeniaca L.) genotypes using simple sequence repeats. Theor Appl Genet 104(2–3):321–328. doi:10.1007/s001220100684
Hou B, Xu Z (2005) Relationship of the occurences and evolutions of Wild-Fruit forests with climatic factors in the Tianshan Mountain. Acta Botanica Boreali-Occidentalla Sinica 25(11):2266 (in Chinese)
Hurtado MA, Romero C, Vilanova S, Abbott AG, Llacer G, Badenes ML (2002) Genetic linkage maps of two apricot cultivars (Prunus armeniaca L.), and mapping of PPV (sharka) resistance. Theor Appl Genet 105(2-3):182–191. doi:10.1007/s00122-002-0936-y
Li MM, Cai YL, Qian ZQ, Zhao GF (2009) Genetic diversity and differentiation in Chinese sour cherry Prunus pseudocerasus Lindl., and its implications for conservation. Genet Resour Crop Evol 56(4):455–464. doi:10.1007/s10722-008-9378-y
Liedloff A (1999) Mantel V2.0, nonparametric test calculator. Queensland University of Technology, Australia
Liu W, Liu D, Zhang A, Feng C, Yang J, Yoon J, Li S (2007) Genetic diversity and phylogenetic relationships among plum germplasm resources in China assessed with inter-simple sequence repeat markers. J Am Soc Hortic Sci 132(5):619–628
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27(2):209–220
Martin C, Herrero M, Hormaza JI (2011) Molecular characterization of apricot germplasm from an old stone collection. PLoS ONE 6(8):e23979. doi:10.1371/journal.pone.0023979
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70(12):3321–3323. doi:10.1073/pnas.70.12.3321
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89(3):583–590
Reddy MP, Sarla N, Siddiq EA (2002) Inter simple sequence repeat (ISSR) polymorphism and its application in plant breeding. Euphytica 128(1):9–17
Rohlf FJ (2000) NTSYS-PC, numerical taxonomy system for the PC ExeterSoftware, Version 2.1. Applied Biostatistics Inc Setauket, USA
Rubio-Moraga A, Candel-Perez D, Lucas-Borja ME, Tiscar PA, Viñegla B, Linares JC, Gómez-Gómez L, Ahrazem O (2012) Genetic diversity of Pinus nigra Arn. populations in southern Spain and northern Morocco revealed by inter-simple sequence repeat profiles. Int J Mol Sci 13(5):5645–5658. doi:10.3390/ijms13055645
Schaal BA, Hayworth DA, Olsen KM, Rauscher JT, Smith WA (1998) Phylogeographic studies in plants: problems and prospects. Mol Ecol 7:465–474. doi:10.1046/j.1365-294x.1998.00318.x
Shahi-Gharahlar A, Zamani Z, Fatahi R, Bouzari N (2011) Estimation of genetic diversity in some Iranian wild Prunus subgenus Cerasus accessions using inter-simple sequence repeat (ISSR) markers. Biochem Syst Ecol 39(4–6):826–833. doi:10.1016/j.bse.2011.07.018
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana
Shiran B, Amirbakhtiar N, Kiani S, Mohammadi S, Sayed-Tabatabaei BE, Moradi H (2007) Molecular characterization and genetic relationship among almond cultivars assessed by RAPD and SSR markers. Sci Hortic 111(3):280–292. doi:10.1016/j.scienta.2006.10.024
Slatkin M (1985) Gene flow in natural populations. Annu Rev Ecol Syst 16:393–430. doi:10.1146/annurev.ecolsys.16.1.393
Slatkin M (1987) Gene flow and the geographic structure of natural populations. Science 236(4803):787–792. doi:10.1126/science.3576198
Sorkheh K, Shiran B, Gradzeil TM, Epperson P, Martinez-Gómez P, Asadi E (2007) Amplified fragment length polymorphism as a tool for molecular characterization of almond germplasm: genetic diversity among genotypes and related wild species of almond, and its relationships with agronomic traits. Euphytica 156:327–344
SPSS Rel. 16.0.0 (2007) SPSS Inc., Chicago, USA
Wang YZ, Zhang JH, Sun HY, Ning N, Yang L (2011) Construction and evaluation of a primary core collection of apricot germplasm in China. Sci Hortic 128(3):311–319. doi:10.1016/j.scienta.2011.01.025
Yeh FC, Yang RC, Boyle TJB (1999) Popgene, Version 1.31. Available from http://www.ualberta.ca/~fyeh/
Yilmaz KU, Paydas-Kargi S, Dogan Y, Kafkas S (2012) Genetic diversity analysis based on ISSR, RAPD and SSR among Turkish apricot germplasms in Iran Caucasian eco-geographical group. Sci Hortic 138:138–143. doi:10.1016/j.scienta.2012.02.017
Yılmaz KU, Ercişli S, Asma BM, Doğan Y, Kafkas S (2009) Genetic relatedness in Prunus genus revealed by inter-simple sequence repeat markers. HortScience 44(2):293–297
Zhang XS (1973) Some issues about the ecological and geographical characteristics and the communities of wild fruit forests in the Yili region. J Bot Sin 15(2):239–252 (in Chinese)
Zhang F, Ge S (2002) Data analysis in population genetics. I. Analysis of RAPD data with AMOVA. Chin Biodivers 10(4):438 (in Chinese)
Zhebentyayeva TN, Reighard GL, Gorina VM, Abbott AG (2003) Simple sequence repeat (SSR) analysis for assessment of genetic variability in apricot germplasm. Theor Appl Genet 106(3):435–444. doi:10.1007/s00122-002-1069-z
Zhebentyayeva TN, Ledbetter C, Burgos L, Llácer G (2012) Apricot. In: Badenes ML, Byrne DH (eds) Fruit breeding. Handbook of plant breeding, vol 8. Springer, New York, pp 415–457
Acknowledgments
This work was supported by grants from the Research Achievement Transformation Program of the Chinese Ministry of Science and Technology (No. 2009GB23600515) and Special Research Program for Public-welfare Forestry of the Chinese State Forestry Administration (No. 200904020).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, M., Zhao, Z. & Miao, X.J. Genetic variability of wild apricot (Prunus armeniaca L.) populations in the Ili Valley as revealed by ISSR markers. Genet Resour Crop Evol 60, 2293–2302 (2013). https://doi.org/10.1007/s10722-013-9996-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-013-9996-x