Abstract
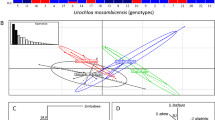
Dendrocalamus giganteus Munro is a high-value woody bamboo widely grown in Southeast Asia and China’s Yunnan Province. We investigated its genetic diversity in Yunnan as a prelude to considering effective breeding programs and the protection of germplasm resources. Inter-simple sequence repeat (ISSR) markers were used to assess the genetic structure and differentiation of seven populations. Seven ISSR primers generated 140 bands, of which 124 were polymorphic (88.57%). Genetic diversity within populations was relatively low, averaging 11.33% polymorphic bands (PPB), while diversity was considerably higher among populations, with PPB = 88.57%. Greater genetic differentiation was detected among populations (G ST = 0.8474). We grouped these seven populations into two clusters within an UPGMA dendrogram—one comprised the Xinping and Shiping populations from central Yunnan, the other included the remaining five populations. Mantel tests indicated no significant correlation between genetic and geographic distances among populations. Breeding system characteristics, genetic drift, and limited gene flow (N m = 0.0901) might be important factors for explaining this differentiation. Based on the overall high genetic diversity and differentiation among D. giganteus populations in Yunnan, we suggest the implementation of in situ conservation measures for all populations and sufficient sampling for ex situ conservation collections.



Similar content being viewed by others
References
Akciz S, Burchfiel BC, Crowley JL, Yin JY, Chen LZ (2008) Geometry, kinematics, and regional significance of the Chong Shan shear zone, Eastern Himalayan Syntaxis, Yunnan, China. Geosphere 4:292–314
Barkley NA, Newman ML, Wang ML, Hotchkiss MW, Pederson GA (2005) Assessment of the genetic diversity and phylogenetic relationships of a temperate bamboo collection by using transferred EST-SSR markers. Genome 48:731–737
Das M, Bhattacharya S, Pal A (2005) Generation and characterization of SCARs by cloning and sequencing of RAPD products: a strategy for species-specific marker development in bamboo. Ann Bot (Lond) 95:835–841
Doyle JJ, Doyle JL (1991) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Dransfield S, Widjaja EA (1995) Plant resources of South-East Asia, (No. 7): bamboos. Backhuys Publishers, Leiden, pp 85–87
Du F, Xue JR, Yang YM, Hui CM, Wang J (2000) Study on flowering phenomenon and its type of bamboo in Yunnan in past fifteen years. Sci Silvae Sin 36(6):57–68
Franklin DC (2004) Synchrony and asynchrony: observations and hypotheses for the flowering wave in a long-lived semelparous bamboo. J Biogeogr 31:773–786
Hamrick JL, Godt MJ (1990) Allozyme diversity in plant species. In: Brown AHD, Clegg MT, Kahler AL, Weir BS (eds) Plant population genetics, breeding, and genetic resources. Sinauer, Sunderland, pp 43–63
Holttum RE (1958) The bamboos of the Malay Peninsula. Gard Bull Singapore 16:1–135
Janzen DH (1976) Why do bamboos wait so long to flower? Annu Rev Ecol Syst 7:347–391
Ji SC, Wang Q, Sun SS, Xu ZQ, Li HB (2008) Continental extrusion and seismicity in China. Acta Geol Sin 82:1643–1667
Keng PC, Wang CP (1996) Flora reipublicae popularis sinicae, vol 9(1): Bambusoideae. Science Press, Beijing, pp 155–157
Leu A (1999) The flowering of one selection of the Indonesian type of Dendrocalamus asper known as Bambu Betung. Am Bamb Soc Newsl 20(5):7–9
Lewontin RC (1972) The apportionment of human diversity. Evol Biol 6:381–398
Li DZ, Hsueh CJ (1988) A study on the genus Dendrocalamus Nees from China. J Bamb Res 7(4):12–13
Lin XC, Ruan XS, Lou YF, Guo XQ, Fang W (2009) Genetic similarity among cultivars of Phyllostachys pubescens. Plant Syst Evol 277:67–73
McClure FA (1966) The bamboos, a fresh perspective. Harvard University Press, Cambridge
Miller MP (1997) Tools for population genetic analysis (TFPGA), version 1.3. Department of Biological Science, Northern Arizona University, Flagstaff
Muller J (1996) Cultivated Gigantochloa: escape from “death by flowering”. Am Bamb Soc Newsl 17(1):4–7
Nei M (1972) Genetic distance between populations. Am Natural 106:283–292
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nelson BW (1994) Natural forest disturbance and change in the Brazilian Amazon. Rem Sens Rev 10:105–125
Nybom H, Bartish IV (2000) Effects of life history traits and sampling strategies on genetic diversity estimates obtained with RAPD markers in plants. Pers Plant Ecol Evol Syst 3:93–114
Ohrnberger D, Goerrings J (1986) The bamboos of the world. Odenthal, Germany
Ramanayake SMSD, Yakandawala K (1998) Incidence of flowering death and phenology of development in the giant bamboo (Dendrocalamus giganteus Wall. ex Munro). Ann Bot (Lond) 82:779–785
Ramanayake SMSD, Meemaduma VN, Weerawardene TE (2007) Genetic diversity and relationships between nine species of bamboo in Sri Lanka, using random amplified polymorphic DNA. Plant Syst Evol 269:55–61
Rohlf FJ (2000) NTSYSpc, numerical taxonomy and multivariate analysis system version 2.1. Department of Ecology and Evolution, State University of New York, Stony Brook
Sharma ML (1994) The flowering of bamboo: fallacies and facts. In: Proceedings of the bamboo in Asia and the Pacific, Chiangmai, Thailand, pp 68–70
Sharma RK, Gupta P, Sharma V, Sood A, Mohapatra T, Ahuja PS (2008) Evaluation of rice and sugarcane SSR markers for phylogenetic and genetic diversity analyses in bamboo. Genome 51:91–103
Slatkin M, Barton NH (1989) A comparison of three indirect methods for estimating average levels of gene flow. Evolution 43:1349–1368
Tapponnier P, Lacassin R, Leloup PH, Schärer U, Zhong DL, Wu HW, Liu XH, Ji SC, Zhang LH, Zhong JY (1990) The Ailao Shan/Red River metamorphic belt: tertiary left-lateral shear between Indochina and South China. Nature 343:431–437
Ueda K (1960) Studies on the Physiology of bamboo, with references to practical application. Prime Minister’s Office, Tokyo
Williams JGK, Kubelik AR, Livak KJ, Rafalski JA, Tingey SV (1990) DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 18:6531–6535
Xiao LQ, Ge XJ, Gong X, Hao G, Zheng S (2004) ISSR variation in the endemic and endangered plant Cycas guizhouensis (Cycadaceae). Ann Bot (Lond) 94:133–138
Yeh FC, Yang RC, Boyle T (1999) POPGENE. Microsoft Windows-based freeware for population genetic analysis. Release 1.31. University of Alberta, Edmonton
Zhang LJ, Dai SL (2010) Genetic variation within and among populations of Orychophragmus violaceus (Cruciferae) in China as detected by ISSR analysis. Genet Resour Crop Evol 57:55–64
Zhang WY, Ma NX (1990) Vitality of bamboo pollens and natural pollination in bamboo plants. For Res 3:250–255
Zhang WY, Ma NX, Chen HX (1989) The shape of bamboo pollens and their germination test. For Res 2:67–69
Acknowledgments
We thank two anonymous reviewers for their constructive comments and suggestions. We are also grateful to Dr. Long-Qian Xiao for his kind help in data analysis. This work was funded through projects of the National Natural Science Foundation of China (31070593), Applied Basic Research Program of Yunnan Province, China (2010CD141), the State Forestry Administration of China (2008-4-30), and International Centre for Bamboo and Rattan (1632009014).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tian, B., Yang, HQ., Wong, KM. et al. ISSR analysis shows low genetic diversity versus high genetic differentiation for giant bamboo, Dendrocalamus giganteus (Poaceae: Bambusoideae), in China populations. Genet Resour Crop Evol 59, 901–908 (2012). https://doi.org/10.1007/s10722-011-9732-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-011-9732-3