Abstract
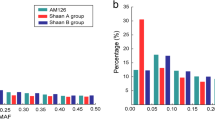
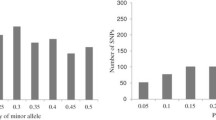
Advances in high-throughput SNP genotyping and genome sequencing technologies have enabled genome-wide association mapping in dissecting the genetic basis of complex quantitative traits. In this study, 82 SSRs and 884 SNPs with minor allele frequencies (MAF) over 0.20 were used to compare their ability to assess population structure, principal component analysis (PCA) and relative kinship in a maize association panel consisting of 154 inbred lines. Compared to SNPs, SSRs provided more information on genetic diversity. The expected heterozygosity (He) of SSRs and SNPs averaged 0.65 and 0.44, and the polymorphic information content of these two markers was 0.61 and 0.34 in this panel, respectively. Additionally, SSRs performed better at clustering all lines into groups using STRUCTURE and PCA approaches, and estimating relative kinship. For both marker systems, the same clusters were observed based on PCA and the first two eigenvectors accounted for similar percentage of genetic variations in this panel. The correlation coefficients of each eigenvector from SSRs and SNPs decreased sharply when the eigenvector varied from 1 to 3, but kept around 0 when the eigenvector were over 3. The kinship estimates based on SSRs and SNPs were moderately correlated (r 2 = 0.69). All these results suggest that SSR markers with moderate density are more informative than SNPs for assessing genetic relatedness in maize association mapping panels.






Similar content being viewed by others
References
Altshuler D, Daly MJ, Lander ES (2008) Genetic mapping in human disease. Science 322:881–888
Aranzana MJ, Kim S, Zhao KY, Bakker E, Horton M, Jakob K, Lister C, Molitor J, Shindo C, Tang CL, Toomajian C, Traw B, Zheng HG, Bergelson J, Dean C, Marjoram P, Nordborg M (2005) Genome-wide association mapping in Arabidopsis identifies previously known flowering time and pathogen resistance genes. PLoS Genet 1(5):e60
Beló A, Zheng P, Luck S, Shen B, Meyer DJ, Li BL, Tingey S, Rafalski A (2008) Whole genome scan detects an allelic variant of fad2 associated with increased oleic acid levels in maize. Mol Genet Genomics 279:1–10
Breseghello F, Sorrells ME (2006) Association analysis as a strategy for improvement of quantitative traits in plants. Crop Sci 46:1323–1330
Buckler ES, Gore M (2007) An Arabidopsis haplotype map takes root. Nat Genet 39(9):1056–1057
Chan EKF, Rowe HC, Kliebenstein DJ (2009) Understanding the evolution of defense metabolites in Arabidopsis thaliana using genome-wide association mapping. Genetics 109:108522
Clark RM, Schweikert G, Toomajian C, Ossowski S, Zeller G, Shinn P, Warthmann N, Hu TT, Fu G, Hinds DA, Chen H, Frazer KA, Huson DH, Schölkopf B, Nordborg M, Rätsch G, Ecker JR, Weige D (2007) Common sequence polymorphisms shaping genetic diversity in Arabidopsis thaliana. Science 317:338–342
Devlin B, Roeder K (1999) Genomic control for association studies. Biometrics 55:997–1004
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Fan JB, Gunderson KL, Bibikova M, Yeakley JM, Chen J, Garcia EW, Lebruska LL, Laurent M, Shen R, Barker D (2006) Illumina universal bead arrays. Methods Enzymol 410:57–73
Gore M, Chia JM, Elshire R, Ersoz E, Hurwitz B, Grills G, Ware D, Buckler ES (2009) A first generation haplotype map of the maize genome. In: The 51st maize genetics conference abstracts, p 39
Gupta PK, Rustgi S, Mir RR (2008) Array-based high-throughput DNA markers for crop improvement. Heredity 101:5–18
Hamblin MT, Warburton ML, Buckler ES (2007) Empirical comparison of simple sequence repeats and single nucleotide polymorphisms in assessment of maize diversity and relatedness. PLoS ONE 12:e1367
Hardy OJ, Vekemans X (2002) Spagedi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Harjes CE, Rocheford TR, Bai L, Brutnell TP, Kandianis CB, Sowinski SG, Stapleton AE, Vallabhaneni R, Williams M, Wurtzel ET, Yan JB, Buckler ES (2008) Natural genetic variation in lycopene epsilon cyclase tapped for maize biofortification. Science 319:330–333
Inghelandt DV, Melchinger AE, Lebreton C, Stich B (2010) Population structure and genetic diversity in a commercial maize breeding program assessed with SSR and SNP markers. Theor Appl Genet 120(7):1289–1299
Jones ES, Sullivan H, Bhattramakki D, Smith JSC (2007) A comparison of simple sequence repeat and single nucleotide polymorphism marker technologies for the genotypic analysis of maize (Zea mays L.). Theor Appl Genet 115:361–371
Kruglyak S, Durrett RT, Schug MD, Aquadro CF (1998) Equilibrium distributions of microsatellite repeat length resulting from a balance between slippage events and point mutations. Proc Natl Acad Sci USA 95:10774–10778
Li WH, Gojobori T, Nei M (1981) Pseudogenes as a paradigm of neutral evolution. Nature 292:237–239
Liu KJ, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21(9):2128–2129
Liu KJ, Goodman M, Muse S, Smith JS, Buckler ES, Doebley J (2003) Genetic structure and diversity among maize inbred lines as inferred from DNA microsatellites. Genetics 165:2117–2128
Loiselle BA, Sork VL, Nason J, Graham C (1995) Spatial genetic structure of a tropical understory shrub, Psychotria officinalis (Rubiaceae). Am J Bot 82:1420–1425
Mackay TF (2001) The genetic architecture of quantitative traits. Annu Rev Genet 35:303–339
Malosetti M, van der Linden CG, Vosman B, van Eeuwijk FA (2007) A mixed-model approach to association mapping using pedigree information with an illustration of resistance to Phytophthora infestans in potato. Genetics 175:879–889
Martinez-Arias R, Calafell F, Mateu E, Comas D, Andrés A, Bertranpetit J (2001) Sequence variability of a human pseudogene. Genome Res 11:1071–1085
Murray SC, Rooney WL, Hamblin MT, Mitchell SE, Kresovich S (2009) Sweet sorghum genetic diversity and association mapping for brix and height. Plant Genome 2:48–62
Myles S, Peiffer J, Brown PJ, Ersoz ES, Zhang ZW, Costich DE, Buckler ED (2009) Association mapping: critical considerations shift from genotyping to experimental design. Plant Cell (http://www.plantcell.org/cgi/doi/10.1105/tpc.109.068437)
Nei M (1972) Genetic distance between populations. Am Nat 106:283–292
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38(8):904–909
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Reif JC, Warburton ML, Xia XC, Hoisington DA, Crossa J, Taba S, Muminovic J, Bohn M, Frisch M, Melchinger AE (2006) Grouping of accessions of Mexican races of maize revisited with SSR markers. Theor Appl Genet 113:177–185
Rohlf FJ (2000) NTSYS-pc. Numerical taxonomy and multivariate analysis system, version 2.1. Exeter Soft ware, New York
Smith JSC, Chin ECL, Shu H, Smith S, Wall SJ, Senior ML, Mitchell SE, Kresovich S, Ziegle J (1997) An evaluation of the utility of SSR loci as molecular markers in maize (Zea mays L.): comparisons with data from RFLPS and pedigree. Theor Appl Genet 95:163–173
Song TM, Chen SJ (2004) Long term selection for oil concentration in five maize populations. Maydica 49:9–14
Teng WT, Can JS, Chen YH, Liu XH, Jing XQ, Zhang FJ, Li JS (2004) Analysis of maize heterotic groups and patterns during past decade in China. Sci Agric Sin 37:1804–1811
Thomson MJ, Septiningsih EM, Suwardjo F, Santoso TJ, Silitonga TS, McCouch SR (2007) Genetic diversity analysis of traditional and improved Indonesian rice (Oryza sativa L.) germplasm using microsatellite markers. Theor Appl Genet 114:559–568
Vignal A, Milan D, Sancristobal M, Eggen A (2002) A review on SNP and other types of molecular markers and their use in animal genetics. Genet Sel Evol 34:275–305
Wang SC, Basten CJ, Zeng ZB (2005) Windows QTL cartographer 2.5 user manual. North Carolina State University, Raleigh
Yan JB, Yang XH, Hector S, Sánchez H, Li JS, Warburton M, Zhou Y, Crouch JH, Xu YB (2010) High-throughput SNP genotyping with the golden gate assay in maize. Mol Breed 25:441–451
Yang XH, Yan JB, Zheng YP, Yu JM, Li JS (2007) Reviews of association analysis for quantitative traits in plant. Acta Agron Sin 33:523–530
Yang XH, Yan JB, Shah T, Warburton ML, Li Q, Li L, Gao YF, Chai YC, Fu ZY, Zhou Y, Xu XT, Bai GH, Meng YJ, Zheng YP, Li JS (2010) Genetic analysis and characterization of a new maize association mapping panel for quantitative trait loci dissection. Theor Appl Genet 121:417–431
Yu JM, Buckler ES (2006) Genetic association mapping and genome organization of maize. Curr Opin Biotechnol 17:1–6
Yu JM, Pressoir G, Briggs WH, Bi IV, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Yu JM, Zhang ZW, Zhu CS, Tabanao DA, Pressoir G, Tuinstra MR, Kresovich S, Todhunter RJ, Buckler ES (2009) Simulation appraisal of the adequacy of number of background markers for relationship estimation in association mapping. Plant Genome 2:63–77
Zhao K, Aranzana MJ, Kim S, Lister C, Shindo C, Tang C, Toomajian C, Zheng H, Dean C, Marjoram P, Nordborg M (2007) An Arabidopsis example of association mapping in structured samples. PLoS Genet 3:e4
Zheng G, Freidlin B, Li ZH, Gastwirth JL (2005) Genomic control for association studies under various genetic models. Biometrics 61:186–192
Zhu CS, Yu JM (2009) Nonmetric multidimensional scaling corrects for population structure in association mapping with different sample types. Genetics 182:875–888
Zhu CS, Gore M, Buckler ES, Yu JM (2008) Status and prospects of association mapping in plants. Plant Genome 1:5–20
Acknowledgments
This research was supported by the National High Technology Research and Development Program of China.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yang, X., Xu, Y., Shah, T. et al. Comparison of SSRs and SNPs in assessment of genetic relatedness in maize. Genetica 139, 1045–1054 (2011). https://doi.org/10.1007/s10709-011-9606-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-011-9606-9