Abstract
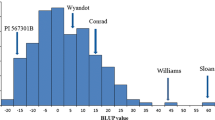
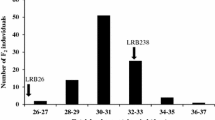
‘Conrad’, a soybean cultivar tolerant to Phytophthora root rot (PRR), and ‘OX760-6-1’, a breeding line with low tolerance to PRR, were crossed. F2 derived recombinant inbred lines were advanced to F6 to generate a population through single-seed descent. This population was used to identify quantitative trait loci (QTLs) influencing PRR tolerance in ‘Conrad’. A total of 99 simple sequence repeat (SSR), or microsatellite, markers that were polymorphic and clearly segregated in the F6 mapping population were used for QTL detection. Based on the data of PRR in the field at two planting locations, Woodslee and Weaver, for the years 2000 and 2001, one putative QTL, designated as Qsatt414-596, was detected using MapMaker/QTL. Qsatt414-596 was flanked by two SSR markers from the linkage group MLG J, Satt414 and Satt596. Satt414 and Satt596 were also detected to be significantly (P < 0.005) associated with PRR using the SAS GLM procedure and were estimated to explain 13.7% and 21.5% of the total phenotypic variance, respectively.

Similar content being viewed by others
References
Abney TS, Melgar JC, Richards TL, Scott DH, Grogan J, Young J (1997) New races of Phytophthora sojae with Rps1d virulence. Plant Dis 81:653–655.
Anderson TR, Buzzell RI (1992) Diversity and frequency of races of Phytophthora sojae f. sp. glycinea in soybean field in Essex County, Ontario, 1980–1989. Plant Dis 76:587–589.
Arahana VS, Graef GL, Specht JE, Steadman JR, Eskridge KM (2001) Identification of QTLs for resistance to Scleritinia sclerotiorum in soybean. Crop Sci 41:180–188.
Bernard RL, Smith PE, Kaufmann MJ, Schmitthenner AF (1957) Inheritance of resistance to Phytophthora root rot and stem rot in the soybean. Agron J 49:391.
Bianchi-Hall CM, Carter TE, Bailey JMA, Main MAR, Rufty TW, Ashley DA, Boerma HR, Arellano C, Hussey RS, Parrott WA (2000) Aluminum tolerance associated with quantitative trait loci derived from soybean PI 416937 in hydroponics. Crop Sci 40:538–545.
Burnham KD, Dorrance AE, VanToai TT, St. Martin SK (2003) Quantitative trait loci for partial resistance to Phytophthora sojae in soybean. Crop Sci 43:1610–1617.
Buzzell RI, Anderson TR (1982) Plant loss response of soybean cultivars to Phytophthora megasperma f. sp. glycinea under field conditions. Plant Dis 66:1146–1148
Cregan PB, Jarvik T, Bush AL, Shoemaker RC, Lark KG, Kathler AL, Kaya N, VanToai TT, Lohnes DG, Chung J, Specht JE (1999) An integrated genetic linkage map of the soybean genome. Crop Sci 39:1464–1490
Dorrance AE, McClure SA, deSilva A (2003a) Pathogenic diversity of Phytophthora sojae in Ohio soybean fields. Plant Dis 87:139–146
Dorrance AE, McClure SA, St. Martin SK (2003b) Effect of partial resistance on Phytophthora stem rot incidence and yield of soybean in Ohio. Plant Dis 87:308–312
Drenth A, Whisson SC, Maclean DJ, Irwin JAG, Obst NR, Ryley MJ (1996) The evolution of races of Phytophthora sojae in Australia. Genetics 86:163–169
Faris MA, Sabo FE (1989) The systematics of Phytophthora sojae and P. megasperma. Can J Bot 67:1442
Fehr WR, Cavines CE, Burmood DT, Pennington JS (1971) Stages of development descriptions for soybeans, Glycine max (L.) Merrill. Crop Sci 11:929–931
Fehr WR, Cianzio SR, Voss BK, Schultz SP (1989) Registration of ‘Conrad’ soybean. Crop Sci 29:830
Forster H, Tyler BM, Coffey MD (1994). Phytophthora sojae races have arisen by clonal evolution and by rare outcrosses. Mol Plant-Microbe Interact 7:780–791
Glover KD, Scott RA (1998) Heritability and phenotypic variation of tolerance to Phytophthora root rot of soybean. Crop Sci 38:1495–1500
Hildebrand AA (1959) A root and stalk rot of soybean caused by Phytophthora megasperma var. sojae var. nov. Can J Bot 37:927–957
Kassem MA, Shultz J, Meksem K, Cho Y, Wood AJ, Iqbal MJ, Lightfood DA (2006) An updated ‘Essex’ by ‘Forrest’ linkage map and first composite interval map of QTL underlying six soybean traits. Theor Appl Genet 113:1015–1026
Kaufmann MJ, Gerdemann JW (1958) Root and stem rot of soybean caused by Phytophthora sojae n. sp. Phytopathology 48:201–208
Keeling BL (1982) Four new physiologic races of Phytophthora sojae n. sp. Glycinea. Plant Dis 66:334–335
Kim HS, Diers BW (2000) Inheritance of partial resistance to Sclerotinia stem rot in soybean. Crop Sci 40:55–61
Kyle DE, Nickell CD (1998) Genetic analysis of tolerance to Phytophthora sojae in the soybean cultivar, Jack. Soybean Genetics Newsletter 25:124–125
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MapMaker: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Mansur LM, Orf JH, Chase K, Jarvik T, Cregan PB, Lark KG (1996) Genetic mapping of agronomic traits using recombinant inbred lines of soybean. Crop Sci 36:1327–1336
Meksem K, Pantazopoulos P, Njiti VN, Hyten LD, Arelli PR, Lightfoot DA (2001) ‘Forrest’ resistance to the soybean cyst nematode is bigenic: saturation mapping of the Rhg1 and Rhg4 loci. Theor Appl Genet 103:710–717
Schafer JF (1971) Tolerance to plant disease. Annu Rev Phytopathol 9:235–252
Schmitthenner AF (1985) Problems and progress in control of Phytophthora root rot of soybean. Plant Dis 69:362–368
Schmitthenner AF, Hobe M, Bhat RG (1994) Phytophthora sojae races in Ohio over a 10-year interval. Plant Dis 78:269–276
Schuster I, Abdelnoor RV, Marin SRR, Carvalho VP, Kiihl RAS, Silva JFV, Sediyama CS, Barros EG, Moreira MA (2001) Identification of a new major QTL associated with resistance to soybean cyst nematode (Heterodera glycines). Theor Appl Genet 102:91–96
Shoemaker RC, Olson TC (1993) Molecular linkage map of soybean (Glycine max L. Merr.). In: O’Brien SJ (ed), Genetic maps: locus maps of complex genomes. Cold Spring Harbor Laboratory Press, New York, p 6.131–6.138
Song QJ, Marek LF, Shoemaker RC, Lark KG, Concibido VC, Delannay X, Specht JE, Cregan PB (2004) A new integrated genetic linkage map of the soybean. Theor Appl Genet 109:122–128
St Martin SK, Scott DR, Schmittchenner AF, McBlain BA (1991) Relationship between tolerance to Phytophthora rot and soybean yield. Plant Breed 113:331–334
Thomison PR, Thomas CA, Kenworthy WJ, Mcintosh MS (1988) Evidence of pathogen specificity in tolerance of soybean cultivars to Phytophthora rot. Crop Sci 28:714–715
Tooley PW, Grau CR (1984a) Field characterisation of rate-reducing resistance to Phytophthora megasperma f. sp. Glycinea in soybean. Phytopathology 74:1201–1208
Tooley PW, Grau CR (1984b) The relationship between rate-reducing resistance to Phytophthora megasperma f. sp. Glycinea and yield of soybean. Phytopathology 74:1209–1216
Walker AK, Schmitthenner AF (1984) Heritability of tolerance to Phytophthora rot in soybean. Crop Sci 24:490–491
Yu K, Park SJ, Poysa V (1999) Abundance and variation of microsatellite DNA sequences in beans (Phaseolus and Vigna). Genome 42:27–34
Yue P, Sleper DA, Arelli P (2001) Mapping resistance to multiple races of Heterodera glycines in soybean PI 89772. Crop Sci 41:1589–1595
Yue P, Arelli P, Sleper DA (2001) Molecular characterization of resistance to Heterodera glycines in soybean PI 438489B. Theor Appl Genet 102:921–928
Acknowledgements
The authors wish to thank Chuck Meharg and Elaine Lepp for technical assistance and the Ontario Soybean Growers for financial assistance for this project.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Weng, C., Yu, K., Anderson, T.R. et al. A quantitative trait locus influencing tolerance to Phytophthora root rot in the soybean cultivar ‘Conrad’. Euphytica 158, 81–86 (2007). https://doi.org/10.1007/s10681-007-9428-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-007-9428-0