Abstract
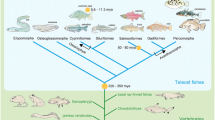
Whole-genome duplication (WGD) is believed to be one of the major evolutionary events that shaped the genome organization of vertebrates. Here, we review recent research on vertebrate genome evolution, specifically on WGD and its consequences for gene and genome evolution in teleost fishes. Recent genome analyses confirmed that all vertebrates experienced two rounds of WGD early in their evolution, and that teleosts experienced a subsequent additional third-round (3R)-WGD. The 3R-WGD was estimated to have occurred 320–400 million years ago in a teleost ancestor, but after its divergence from a common ancestor with living non-teleost actinopterygians (Bichir, Sturgeon, Bowfin, and Gar) based on the analyses of teleost-specific duplicate genes. This 3R-WGD was confirmed by synteny analysis and ancestral karyotype inference using the genome sequences of Tetraodon and medaka. Most of the tetrapods, on the other hand, have not experienced an additional WGD; however, they have experienced repeated chromosomal rearrangements throughout the whole genome. Therefore, different types of chromosomal events have characterized the genomes of teleosts and tetrapods, respectively. The 3R-WGD is useful to investigate the consequences of WGD because it is an evolutionarily recent WGD and thus teleost genomes retain many more WGD-derived duplicates and “traces” of their evolution. In addition, the remarkable morphological, physiological, and ecological diversity of teleosts may facilitate understanding of macrophenotypic evolution on the basis of genetic/genomic information. We highlight the teleosts with 3R-WGD as unique models for future studies on ecology and evolution taking advantage of emerging genomics technologies and systems biology environments.







Similar content being viewed by others
References
Abi-Rached L, Gilles A, Shiina T, Pontarotti P, Inoko H (2002) Evidence of en bloc duplication in vertebrate genomes. Nat Genet 31:100–105. doi:10.1038/ng855
Allendorf FW, Thorgaard GH (1984) Tetraploidy and the evolution of salmonid fishes. In: Turner BJ (ed) The evolutionary genetics of fishes. Plenum Press, New York, pp 1–53
Amemiya CT, Prohaska SJ, Hill-Force A, Cook A, Wasserscheid J, Ferrier DE, Pascual-Anaya J, Garcia-Fernàndez J, Dewar K, Stadler PF (2008) The amphioxus Hox cluster: characterization, comparative genomics, and evolution. J Exp Zoolog B Mol Dev Evol 310:465–477. doi:10.1002/jez.b.21213
Amores A, Force A, Yan YL, Joly L, Amemiya C, Fritz A, Ho RK, Langeland J, Prince V, Wang YL, Westerfield M, Ekker M, Postlethwait JH (1998) Zebrafish hox clusters and vertebrate genome evolution. Science 282:1711–1714. doi:10.1126/science.282.5394.1711
Amores A, Suzuki T, Yan YL, Pomeroy J, Singer A, Amemiya C, Postlethwait JH (2004) Developmental roles of pufferfish Hox clusters and genome evolution in ray-fin fish. Genome Res 14:1–10. doi:10.1101/gr.1717804
Aparicio S (2000) Vertebrate evolution: recent perspectives from fish. Trends Genet 16:54–56. doi:10.1016/S0168-9525(99)01934-4
Aparicio S, Hawker K, Cottage A, Mikawa Y, Zuo L, Venkatesh B, Chen E, Krumlauf R, Brenner S (1997) Organization of the Fugu rubripes Hox clusters: evidence for continuing evolution of vertebrate Hox complexes. Nat Genet 16:79–83. doi:10.1038/ng0597-79
Aparicio S, Chapman J, Stupka E, Putnam N, Chia JM, Dehal P, Christoffels A, Rash S, Hoon S, Smit A, Gelpke MD, Roach J, Oh T, Ho IY, Wong M, Detter C, Verhoef F, Predki P, Tay A, Lucas S, Richardson P, Smith SF, Clark MS, Edwards YJ, Doggett N, Zharkikh A, Tavtigian SV, Pruss D, Barnstead M, Evans C, Baden H, Powell J, Glusman G, Rowen L, Hood L, Tan YH, Elgar G, Hawkins T, Venkatesh B, Rokhsar D, Brenner S (2002) Whole-genome shotgun assembly and analysis of the genome of Fugu rubripes. Science 297:1301–1310. doi:10.1126/science.1072104
Avise JC, Kitto GB (1973) Phosphoglucose isomerase gene duplication in the bony fishes: an evolutionary history. Biochem Genet 8:113–132. doi:10.1007/BF00485540
Azuma Y, Kumazawa Y, Miya M, Mabuchi K, Nishida M (2008) Mitogenomic evaluation of the historical biogeography of cichlids toward reliable dating of teleostean divergences. BMC Evol Biol 8:215. doi:10.1186/1471-2148-8-215
Benton MJ (1993) The fossil record volume 2. Chapman & Hall, London
Benton MJ, Donoghue PC (2007) Paleontological evidence to date the tree of life. Mol Biol Evol 24:26–53. doi:10.1093/molbev/msl150
Braasch I, Salzburger W, Meyer A (2006) Asymmetric evolution in two fish-specifically duplicated receptor tyrosine kinase paralogons involved in teleost coloration. Mol Biol Evol 23:1192–1202. doi:10.1093/molbev/msk003
Braasch I, Schartl M, Volff JN (2007) Evolution of pigment synthesis pathways by gene and genome duplication in fish. BMC Evol Biol 7:74. doi:10.1186/1471-2148-7-74
Braasch I, Brunet F, Volff JN, Schartl M (2009) Pigmentation pathway evolution after whole genome duplication in fish. Genome Biol Evol, in press. doi:10.1093/gbe/evp050
Chambers KE, McDaniell R, Raincrow JD, Deshmukh M, Stadler PF, Chiu CH (2009) Hox cluster duplication in the basal teleost Hiodon alosoides (Osteoglossomorpha). Theory Biosci 128:109–120. doi:10.1007/s12064-009-0056-1
Cheng CH, Chen L (1999) Evolution of an antifreeze glycoprotein. Nature 401:443–444. doi:10.1038/46721
Chiu CH, Amemiya C, Dewar K, Kim CB, Ruddle FH, Wagner GP (2002) Molecular evolution of the HoxA cluster in the three major gnathostome lineages. Proc Natl Acad Sci U S A 99:5492–5497. doi:10.1073/pnas.052709899
Chiu CH, Dewar K, Wagner GP, Takahashi K, Ruddle F, Ledje C, Bartsch P, Scemama JL, Stellwag E, Fried C, Prohaska SJ, Stadler PF, Amemiya CT (2004) Bichir HoxA cluster sequence reveals surprising trends in ray-finned fish genomic evolution. Genome Res 14:11–17. doi:10.1101/gr.1712904
Christoffels A, Koh EG, Chia JM, Brenner S, Aparicio S, Venkatesh B (2004) Fugu genome analysis provides evidence for a whole-genome duplication early during the evolution of ray-finned fishes. Mol Biol Evol 21:1146–1151. doi:10.1093/molbev/msh114
Christoffels A, Brenner S, Venkatesh B (2006) Tetraodon genome analysis provides further evidence for whole-genome duplication in the ray-finned fish. Comp Biochem Physiol Part D Genomics Proteomics 1:13–19. doi:10.1016/j.cbd.2005.06.001
Conant GC, Wolfe KH (2008) Turning a hobby into a job: how duplicated genes find new functions. Nat Rev Genet 9:938–950. doi:10.1038/nrg2482
Crow KD, Wagner GP (2006) What is the role of genome duplication in the evolution of complexity and diversity? Mol Biol Evol 23:887–892. doi:10.1093/molbev/msj083
Crow KD, Stadler PF, Lynch VJ, Amemiya C, Wagner GP (2006) The “fish-specific” Hox cluster duplication is coincident with the origin of teleosts. Mol Biol Evol 23:121–136. doi:10.1093/molbev/msj020
de Boer JG, Yazawa R, Davidson WS, Koop BF (2007) Bursts and horizontal evolution of DNA transposons in the speciation of pseudotetraploid salmonids. BMC Genomics 8:422
Dehal P, Boore JL (2005) Two rounds of whole genome duplication in the ancestral vertebrate. PLoS Biol 3:e314. doi:10.1371/journal.pbio.0030314
Donoghue PC, Purnell MA (2005) Genome duplication, extinction and vertebrate evolution. Trends Ecol Evol 20:312–319. doi:10.1016/j.tree.2005.04.008
Douard V, Brunet F, Boussau B, Ahrens I, Vlaeminck-Guillem V, Haendler B, Laudet V, Guiguen Y (2008) The fate of the duplicated androgen receptor in fishes: a late neofunctionalization event? BMC Evol Biol 8:336. doi:10.1186/1471-2148-8-336
Ferris SD, Portnoy SL, Whitt GS (1979) The roles of speciation and divergence time in the loss of duplicate gene expression. Theor Popul Biol 15:114–139. doi:10.1016/0040-5809(79)90030-3
Flicek P, Aken BL, Beal K, Ballester B, Caccamo M, Chen Y, Clarke L, Coates G, Cunningham F, Cutts T, Down T, Dyer SC, Eyre T, Fitzgerald S, Fernandez-Banet J, Gräf S, Haider S, Hammond M, Holland R, Howe KL, Howe K, Johnson N, Jenkinson A, Kähäri A, Keefe D, Kokocinski F, Kulesha E, Lawson D, Longden I, Megy K, Meidl P, Overduin B, Parker A, Pritchard B, Prlic A, Rice S, Rios D, Schuster M, Sealy I, Slater G, Smedley D, Spudich G, Trevanion S, Vilella AJ, Vogel J, White S, Wood M, Birney E, Cox T, Curwen V, Durbin R, Fernandez-Suarez XM, Herrero J, Hubbard TJ, Kasprzyk A, Proctor G, Smith J, Ureta-Vidal A, Searle S (2008) Ensembl 2008. Nucleic Acids Res 36:D707–D714. doi:10.1093/nar/gkm988
Force A, Lynch M, Pickett FB, Amores A, Yan YL, Postlethwait J (1999) Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151:1531–1545
Furlong RF, Holland PW (2002) Were vertebrates octoploid? Philos Trans R Soc Lond B Biol Sci 357:531–544. doi:10.1098/rstb.2001.1035
Gehring WJ (1998) The homeobox story. Yale University Press, New Haven
Gibson TJ, Spring J (2000) Evidence in favour of ancient octaploidy in the vertebrate genome. Biochem Soc Trans 28:259–264
Gregory TR (2005) The evolution of the genome. Elsevier, San Diego
Hashiguchi Y, Nishida M (2007) Evolution of trace amine associated receptor (TAAR) gene family in vertebrates: lineage-specific expansions and degradations of a second class of vertebrate chemosensory receptors expressed in the olfactory epithelium. Mol Biol Evol 24:2099–2107. doi:10.1093/molbev/msm140
He X, Zhang J (2005) Rapid subfunctionalization accompanied by prolonged and substantial neofunctionalization in duplicate gene evolution. Genetics 169:1157–1164. doi:10.1534/genetics.104.037051
Hedges SB, Kumar S (2003) Genomic clocks and evolutionary timescales. Trends Genet 19:200–206. doi:10.1016/S0168-9525(03)00053-2
Hoegg S, Meyer A (2005) Hox clusters as models for vertebrate genome evolution. Trends Genet 21:421–424. doi:10.1016/j.tig.2005.06.004
Hoegg S, Meyer A (2007) Phylogenomic analyses of KCNA clusters in vertebrates: why do some clusters stay intact? BMC Evol Biol 7:139. doi:10.1186/1471-2148-7-139
Hoegg S, Brinkmann H, Taylor JS, Meyer A (2004) Phylogenetic timing of the fish-specific genome duplication correlates with the diversification of teleost fish. J Mol Evol 59:190–203. doi:10.1007/s00239-004-2613-z
Holland PW, Garcia-Fernàndez J, Williams NA, Sidow A (1994) Gene duplications and the origins of vertebrate development. Dev Suppl 1994:125–133
Horton AC, Mahadevan NR, Ruvinsky I, Gibson-Brown JJ (2003) Phylogenetic analyses alone are insufficient to determine whether genome duplication(s) occurred during early vertebrate evolution. J Exp Zoolog B Mol Dev Evol 299:41–53. doi:10.1002/jez.b.40
Hubbs CL (1955) Hybridization between fish species in nature. Syst Zool 4:1–20
Hufton AL, Groth D, Vingron M, Lehrach H, Poustka AJ, Panopoulou G (2008) Early vertebrate whole genome duplications were predated by a period of intense genome rearrangement. Genome Res 18:1582–1591. doi:10.1101/gr.080119.108
Hughes MK, Hughes AL (1993) Evolution of duplicate genes in a tetraploid animal, Xenopus laevis. Mol Biol Evol 10:1360–1369
Hughes AL, da Silva J, Friedman R (2001) Ancient genome duplications did not structure the human Hox-bearing chromosomes. Genome Res 11:771–780. doi:10.1101/gr.GR-1600R
Imai S, Sasaki T, Shimizu A, Asakawa S, Hori H, Shimizu N (2007) The genome size evolution of medaka (Oryzias latipes) and fugu (Takifugu rubripes). Genes Genet Syst 82:135–144. doi:10.1266/ggs.82.135
Inoue JG, Miya M, Tsukamoto K, Nishida M (2003) Basal actinopterygian relationships: a mitogenomic perspective on the phylogeny of the “ancient fish”. Mol Phylogenet Evol 26:110–120. doi:10.1016/S1055-7903(02)00331-7
Inoue JG, Miya M, Venkatesh B, Nishida M (2005) The mitochondrial genome of Indonesian coelacanth Latimeria menadoensis (Sarcopterygii: Coelacanthiformes) and divergence time estimation between the two coelacanths. Gene 349:227–235. doi:10.1016/j.gene.2005.01.008
International Human Genome Sequencing Consortium (2004) Finishing the euchromatic sequence of the human genome. Nature 431:931–945. doi:10.1038/nature03001
Jaillon O, Aury JM, Brunet F, Petit JL, Stange-Thomann N, Mauceli E, Bouneau L, Fischer C, Ozouf-Costaz C, Bernot A, Nicaud S, Jaffe D, Fisher S, Lutfalla G, Dossat C, Segurens B, Dasilva C, Salanoubat M, Levy M, Boudet N, Castellano S, Anthouard V, Jubin C, Castelli V, Katinka M, Vacherie B, Biémont C, Skalli Z, Cattolico L, Poulain J, De Berardinis V, Cruaud C, Duprat S, Brottier P, Coutanceau JP, Gouzy J, Parra G, Lardier G, Chapple C, McKernan KJ, McEwan P, Bosak S, Kellis M, Volff JN, Guigó R, Zody MC, Mesirov J, Lindblad-Toh K, Birren B, Nusbaum C, Kahn D, Robinson-Rechavi M, Laudet V, Schachter V, Quétier F, Saurin W, Scarpelli C, Wincker P, Lander ES, Weissenbach J, Roest Crollius H (2004) Genome duplication in the teleost fish Tetraodon nigroviridis reveals the early vertebrate proto-karyotype. Nature 431:946–957. doi:10.1038/nature03025
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32(Database issue):D277-D280. doi: 10.1093/nar/gkh063
Kao HW, Lee SC (2002) Phosphoglucose isomerases of hagfish, zebrafish, gray mullet, toad, and snake, with reference to the evolution of the genes in vertebrates. Mol Biol Evol 19:367–374
Karp PD, Ouzounis CA, Moore-Kochlacs C, Goldovsky L, Kaipa P, Ahrén D, Tsoka S, Darzentas N, Kunin V, López-Bigas N (2005) Expansion of the BioCyc collection of pathway/genome databases to 160 genomes. Nucleic Acids Res 33:6083–6089. doi:10.1093/nar/gki892
Kasahara M, Nakaya J, Satta Y, Takahata N (1997) Chromosomal duplication and the emergence of the adaptive immune system. Trends Genet 13:90–92. doi:10.1016/S0168-9525(97)01065-2
Kasahara M, Naruse K, Sasaki S, Nakatani Y, Qu W, Ahsan B, Yamada T, Nagayasu Y, Doi K, Kasai Y, Jindo T, Kobayashi D, Shimada A, Toyoda A, Kuroki Y, Fujiyama A, Sasaki T, Shimizu A, Asakawa S, Shimizu N, Hashimoto S, Yang J, Lee Y, Matsushima K, Sugano S, Sakaizumi M, Narita T, Ohishi K, Haga S, Ohta F, Nomoto H, Nogata K, Morishita T, Endo T, Shin-I T, Takeda H, Morishita S, Kohara Y (2007) The medaka draft genome and insights into vertebrate genome evolution. Nature 447:714–719. doi:10.1038/nature05846
Kawahara R, Miya M, Mabuchi K, Lavoué S, Inoue JG, Satoh TP, Kawaguchi A, Nishida M (2008) Interrelationships of the 11 gasterosteiform families (sticklebacks, pipefishes, and their relatives): a new perspective based on whole mitogenome sequences from 75 higher teleosts. Mol Phylogenet Evol 46:224–236. doi:10.1016/j.ympev.2007.07.009
Kawahara R, Miya M, Mabuchi K, Near TJ, Nishida M (2009) Stickleback phylogenies resolved: evidence from mitochondrial genomes and 11 nuclear genes. Mol Phylogenet Evol 50:401–404. doi:10.1016/j.ympev.2008.10.014
Kellis M, Birren BW, Lander ES (2004) Proof and evolutionary analysis of ancient genome duplication in the yeast Saccharomyces cerevisiae. Nature 428:617–624. doi:10.1038/nature02424
Kim CB, Amemiya C, Bailey W, Kawasaki K, Mezey J, Miller W, Minoshima S, Shimizu N, Wagner G, Ruddle F (2000) Hox cluster genomics in the horn shark, Heterodontus francisci. Proc Natl Acad Sci U S A 97:1655–1660. doi:10.1073/pnas.030539697
Koh EG, Lam K, Christoffels A, Erdmann MV, Brenner S, Venkatesh B (2003) Hox gene clusters in the Indonesian coelacanth, Latimeria menadoensis. Proc Natl Acad Sci U S A 100:1084–1088. doi:10.1073/pnas.0237317100
Kumar S, Hedges SB (1998) A molecular timescale for vertebrate evolution. Nature 392:917–920. doi:10.1038/31927
Kuraku S, Meyer A, Kuratani S (2009) Timing of genome duplications relative to the origin of the vertebrates: did cyclostomes diverge before or after? Mol Biol Evol 26:47–59. doi:10.1093/molbev/msn222
Kurosawa G, Takamatsu N, Takahashi M, Sumitomo M, Sanaka E, Yamada K, Nishii K, Matsuda M, Asakawa S, Ishiguro H, Miura K, Kurosawa Y, Shimizu N, Kohara Y, Hori H (2006) Organization and structure of hox gene loci in medaka genome and comparison with those of pufferfish and zebrafish genomes. Gene 370:75–82. doi:10.1016/j.gene.2005.11.015
Lander ES, International Human Genome Sequencing Consortium et al (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Lavoué S, Miya M, Inoue JG, Saitoh K, Ishiguro NB, Nishida M (2005) Molecular systematics of the gonorynchiform fishes (Teleostei) based on whole mitogenome sequences: implications for higher-level relationships within the Otocephala. Mol Phylogenet Evol 37:165–177. doi:10.1016/j.ympev.2005.03.024
Lavoué S, Miya M, Poulsen JY, Møller PR, Nishida M (2008) Monophyly, phylogenetic position and inter-familial relationships of the Alepocephaliformes (Teleostei) based on whole mitogenome sequences. Mol Phylogenet Evol 47:1111–1121. doi:10.1016/j.ympev.2007.12.002
Ledje C, Kim CB, Ruddle FH (2002) Characterization of Hox genes in the bichir, Polypterus palmas. J Exp Zool 294:107–111. doi:10.1002/jez.10152
Leggatt RA, Iwama GK (2003) Occurrence of polyploidy in the fishes. Rev Fish Biol Fish 13:237–246. doi:10.1023/B:RFBF.0000033049.00668.fe
Lemons D, McGinnis W (2006) Genomic evolution of Hox gene clusters. Science 313:1918–1922. doi:10.1126/science.1132040
Lister JA, Close J, Raible DW (2001) Duplicate mitf genes in zebrafish: complementary expression and conservation of melanogenic potential. Dev Biol 237:333–344. doi:10.1006/dbio.2001.0379
Longhurst TJ, Joss JM (1999) Homeobox genes in the australian lungfish Neoceratodus forsteri. J Exp Zool 285:140–145. doi:10.1002/(SICI)1097-010X(19990815)285:2<140::AID-JEZ6>3.0.CO;2-V
Lundin LG (1993) Evolution of the vertebrate genome as reflected in paralogous chromosomal regions in man and the house mouse. Genomics 16:1–19. doi:doi:10.1006/geno.1993.1133
Lundin LG, Larhammar D, Hallböök F (2003) Numerous groups of chromosomal regional paralogies strongly indicate two genome doublings at the root of the vertebrates. J Struct Funct Genomics 3:53–63. doi:10.1023/A:1022600813840
Lynch M (2002) Gene duplication and evolution. Science 297:945–947. doi:10.1126/science.1075472
Lynch M (2007) Genomic expansion by gene duplication. In: Lynch M (ed) The origins of genome architecture. Sinauer, Sunderland, pp 193–235
Lynch M, Force AG (2000) The origin of interspecific genomic incompatibility via gene duplication. Am Nat 156:590–605. doi:10.1086/316992
Mak HC, Daly M, Gruebel B, Ideker T (2007) CellCircuits: a database of protein network models. Nucleic Acids Res 35:D538–D545. doi:10.1093/nar/gkl937
Mank JE, Avise JC (2006a) Phylogenetic conservation of chromosome numbers in Actinopterygiian fishes. Genetica 127:321–327. doi:10.1007/s10709-005-5248-0
Mank JE, Avise JC (2006b) The evolution of reproductive and genomic diversity in ray-finned fishes: insights from phylogeny and comparative analysis. J Fish Biol 69:1–27. doi:10.1111/j.1095-8649.2006.01132.x
Martin A (2001) Is tetralogy true? Lack of support for the “one-to-four rule”. Mol Biol Evol 18:89–93
Mayden RL, Chen WJ, Bart HL, Doosey MH, Simons AM, Tang KL, Wood RM, Agnew MK, Yang L, Hirt MV, Clements MD, Saitoh K, Sado T, Miya M, Nishida M (2009) Reconstructing the phylogenetic relationships of the earth’s most diverse clade of freshwater fishes–order Cypriniformes (Actinopterygii: Ostariophysi): a case study using multiple nuclear loci and the mitochondrial genome. Mol Phylogenet Evol 51:500–514. doi:10.1016/j.ympev.2008.12.015
Merritt TJS, Quattro JM (2001) Evidence for a period of directional selection following gene duplication in a neurally expressed locus of triosephosphate isomerase. Genetics 159:689–697
Meyer A, Málaga-Trillo E (1999) Vertebrate genomics: More fishy tales about Hox genes. Curr Biol 9:R210–R213. doi:10.1016/S0960-9822(99)80131-6
Meyer A, Schartl M (1999) Gene and genome duplications in vertebrates: the one-to-four (-to-eight in fish) rule and the evolution of novel gene functions. Curr Opin Cell Biol 11:699–704. doi:10.1016/S0955-0674(99)00039-3
Miya M, Takeshima H, Endo H, Ishiguro NB, Inoue JG, Mukai T, Satoh TP, Yamaguchi M, Kawaguchi A, Mabuchi K, Shirai SM, Nishida M (2003) Major patterns of higher teleostean phylogenies: a new perspective based on 100 complete mitochondrial DNA sequences. Mol Phylogenet Evol 26:121–138. doi:10.1016/S1055-7903(02)00332-9
Miya M, Satoh TP, Nishida M (2005) The phylogenetic position of toadfishes (order Batrachoidiformes) in the higher ray-finned fish as inferred from partitioned Bayesian analysis of 102 whole mitochondrial genome sequences. Biol J Linn Soc Lond 85:289–306. doi:10.1111/j.1095-8312.2005.00483.x
Moghadam HK, Ferguson MM, Danzmann RG (2005) Evolution of Hox clusters in Salmonidae: a comparative analysis between Atlantic salmon (Salmo salar) and rainbow trout (Oncorhynchus mykiss). J Mol Evol 61:636–649. doi:10.1007/s00239-004-0338-7
Mouse Genome Sequencing Consortium (2002) Initial sequencing and comparative analysis of the mouse genome. Nature 420:520–562. doi:10.1038/nature01262
Mulley JF, Chiu CH, Holland PW (2006) Breakup of a homeobox cluster after genome duplication in teleosts. Proc Natl Acad Sci U S A 103:10369–10372. doi:10.1073/pnas.0600341103
Murphy WJ, Pevzner PA, O’Brien SJ (2004) Mammalian phylogenomics comes of age. Trends Genet 20:631–639. doi:10.1016/j.tig.2004.09.005
Nagasaki M, Doi A, Matsuno H, Miyano S (2003) Genomic object net: I. A platform for modelling and simulating biopathways. Appl Bioinformatics 2:181–184
Nagasaki M, Saito A, Doi A, Matsuno H, Miyano S (2009) Foundations of systems biology: using Cell Illustrator and pathway databases. Springer-Verlag, New York
Nakatani Y, Takeda H, Kohara Y, Morishita S (2007) Reconstruction of the vertebrate ancestral genome reveals dynamic genome reorganization in early vertebrates. Genome Res 17:1254–1265. doi:10.1101/gr.6316407
Naruse K, Fukamachi S, Mitani H, Kondo M, Matsuoka T, Kondo S, Hanamura N, Morita Y, Hasegawa K, Nishigaki R, Shimada A, Wada H, Kusakabe T, Suzuki N, Kinoshita M, Kanamori A, Terado T, Kimura H, Nonaka M, Shima A (2000) A detailed linkage map of medaka, Oryzias latipes: comparative genomics and genome evolution. Genetics 154:1773–1784
Naruse K, Tanaka M, Mita K, Shima A, Postlethwait J, Mitani H (2004) A medaka gene map: the trace of ancestral vertebrate proto-chromosomes revealed by comparative gene mapping. Genome Res 14:820–828. doi:10.1101/gr.2004004
Nelson JS (2006) Fishes of the world, 4th edn. Wiley, New Jersey
Ohno S (1970) Evolution by gene duplication. Springer-Verlag, New York
Ohno S (1999) Gene duplication and the uniqueness of vertebrate genomes circa 1970–1999. Semin Cell Dev Biol 10:517–522. doi:10.1006/scdb.1999.0332
Pakchung AAH, Simpson PJL, Codd R (2006) Life on earth. Extremophiles continue to move the goal posts Environ Chem 3:77–93. doi:10.1071/EN05093
Panopoulou G, Poustka AJ (2005) Timing and mechanism of ancient vertebrate genome duplications—the adventure of a hypothesis. Trends Genet 21:559–567. doi:10.1016/j.tig.2005.08.004
Panopoulou G, Hennig S, Groth D, Krause A, Poustka AJ, Herwig R, Vingron M, Lehrach H (2003) New evidence for genome-wide duplications at the origin of vertebrates using an amphioxus gene set and completed animal genomes. Genome Res 13:1056–1066. doi:10.1101/gr.874803
Postlethwait JH, Yan YL, Gates MA, Horne S, Amores A, Brownlie A, Donovan A, Egan ES, Force A, Gong Z, Goutel C, Fritz A, Kelsh R, Knapik E, Liao E, Paw B, Ransom D, Singer A, Thomson M, Abduljabbar TS, Yelick P, Beier D, Joly JS, Larhammar D, Rosa F, Westerfield M, Zon LI, Johnson SL, Talbot WS (1998) Vertebrate genome evolution and the zebrafish gene map. Nat Genet 18:345–349. doi:10.1038/ng0498-345
Postlethwait JH, Woods IG, Ngo-Hazelett P, Yan YL, Kelly PD, Chu F, Huang H, Hill-Force A, Talbot WS (2000) Zebrafish comparative genomics and the origins of vertebrate chromosomes. Genome Res 10:1890–1902
Putnam NH, Butts T, Ferrier DE, Furlong RF, Hellsten U, Kawashima T, Robinson-Rechavi M, Shoguchi E, Terry A, Yu JK, Benito-Gutiérrez EL, Dubchak I, Garcia-Fernàndez J, Gibson-Brown JJ, Grigoriev IV, Horton AC, de Jong PJ, Jurka J, Kapitonov VV, Kohara Y, Kuroki Y, Lindquist E, Lucas S, Osoegawa K, Pennacchio LA, Salamov AA, Satou Y, Sauka-Spengler T, Schmutz J, Shin-I T, Toyoda A, Bronner-Fraser M, Fujiyama A, Holland LZ, Holland PW, Satoh N, Rokhsar DS (2008) The amphioxus genome and the evolution of the chordate karyotype. Nature 453:1064–1071. doi:10.1038/nature06967
Rastogi S, Liberles DA (2005) Subfunctionalization of duplicated genes as a transition state to neofunctionalization. BMC Evol Biol 5:28. doi:10.1186/1471-2148-5-28
Ravi V, Venkatesh B (2008) Rapidly evolving fish genomes and teleost diversity. Curr Opin Genet Dev 18:544–550. doi:10.1016/j.gde.2008.11.001
Robinson-Rechavi M, Marchand O, Escriva H, Laudet V (2001a) An ancestral whole-genome duplication may not have been responsible for the abundance of duplicated fish genes. Curr Biol 11:R458–R459. doi:10.1016/S0960-9822(01)00280-9
Robinson-Rechavi M, Marchand O, Escriva H, Bardet PL, Zelus D, Hughes S, Laudet V (2001b) Euteleost fish genomes are characterized by expansion of gene families. Genome Res 11:781–788
Robinson-Rechavi M, Boussau B, Laudet V (2004) Phylogenetic dating and characterization of gene duplications in vertebrates: the cartilaginous fish reference. Mol Biol Evol 21:580–586. doi:10.1093/molbev/msh046
Robinson-Rechavi M, Alibés A, Godzik A (2006) Contribution of electrostatic interactions, compactness and quaternary structure to protein thermostability: lessons from structural genomics of Thermotoga maritima. J Mol Biol 356:547–557. doi:10.1016/j.jmb.2005.11.065
Ruddle FH, Amemiya CT, Carr JL, Kim CB, Ledje C, Shashikant CS, Wagner GP (1999) Evolution of chordate hox gene clusters. Ann N Y Acad Sci 870:238–248
Salzburger W (2009) The interaction of sexually and naturally selected traits in the adaptive radiations of cichlid fishes. Mol Ecol 18:169–185. doi:10.1111/j.1365-294X.2008.03981.x
San Mauro D, Vences M, Alcobendas M, Zardoya R, Meyer A (2005) Initial diversification of living amphibians predated the breakup of Pangea. Am Nat 165:590–599. doi:10.1086/429523
Sasaki T, Nishihara H, Hirakawa M, Fujimura K, Tanaka M, Kokubo N, Kimura-Yoshida C, Matsuo I, Sumiyama K, Saitou N, Shimogori T, Okada N (2008) Possible involvement of SINEs in mammalian-specific brain formation. Proc Natl Acad Sci U S A 105:4220–4225. doi:10.1073/pnas.0709398105
Sato Y, Nishida M (2007) Post-duplication charge evolution of phosphoglucose isomerases in teleost fishes through weak selection on many amino acid sites. BMC Evol Biol 7:204. doi:10.1186/1471-2148-7-204
Sato Y, Nishida M (2009) Electric charge divergence in proteins: insights into the evolution of their three-dimensional properties. Gene 441:3–11. doi:10.1016/j.gene.2008.06.026
Sato Y, Hashiguchi Y, Nishida M (2009a) Temporal pattern of loss/persistence of duplicate genes involved in signal transduction and metabolic pathways after teleost-specific genome duplication. BMC Evol Biol 9:127. doi:10.1186/1471-2148-9-127
Sato Y, Hashiguchi Y, Nishida M (2009b) Evolution of multiple phosphodiesterase isoforms in stickleback involved in cAMP signal transduction pathway. BMC Syst Biol 3:23. doi:10.1186/1752-0509-3-23
Sauro HM, Hucka M, Finney A, Wellock C, Bolouri H, Doyle J, Kitano H (2003) Next generation simulation tools: the systems biology workbench and BioSPICE integration. OMICS 7:355–372. doi:10.1089/153623103322637670
Schwartz FJ (1972) World literature to fish hybrids with an analysis by family, species, and hybrid. Publication no. 3, Gulf Coast Research Laboratory and Museum, Ocean Springs, Mississippi
Seehausen O, Terai Y, Magalhaes IS, Carleton KL, Mrosso HD, Miyagi R, van der Sluijs I, Schneider MV, Maan ME, Tachida H, Imai H, Okada N (2008) Speciation through sensory drive in cichlid fish. Nature 455:620–626. doi:10.1038/nature07285
Sémon M, Wolfe KH (2007a) Reciprocal gene loss between Tetraodon and zebrafish after whole genome duplication in their ancestor. Trends Genet 23:108–112. doi:10.1016/j.tig.2007.01.003
Sémon M, Wolfe KH (2007b) Rearrangement rate following the whole-genome duplication in teleosts. Mol Biol Evol 24:860–867. doi:10.1093/molbev/msm003
Setiamarga DH, Miya M, Yamanoue Y, Mabuchi K, Satoh TP, Inoue JG, Nishida M (2008) Interrelationships of Atherinomorpha (medakas, flyingfishes, killifishes, silversides, and their relatives): the first evidence based on whole mitogenome sequences. Mol Phylogenet Evol 49:598–605. doi:10.1016/j.ympev.2008.08.008
Setiamarga DH, Miya M, Yamanoue Y, Azuma Y, Inoue JG, Ishiguro NB, Mabuchi K, Nishida M (2009) Divergence time of the two regional medaka populations in Japan as a new time scale for comparative genomics of vertebrates. Biol Lett 5:812–816. doi:10.1098/rsbl.2009.0419
Sidow A (1996) Gen(om)e duplications in the evolution of early vertebrates. Curr Opin Genet Dev 6:715–722. doi:10.1016/S0959-437X(96)80026-8
Siegel N, Hoegg S, Salzburger W, Braasch I, Meyer A (2007) Comparative genomics of ParaHox clusters of teleost fishes: gene cluster breakup and the retention of gene sets following whole genome duplications. BMC Genomics 8:312. doi:10.1186/1471-2164-8-312
Spring J (1997) Vertebrate evolution by interspecific hybridisation—are we polyploid? FEBS Lett 40:2–8. doi:10.1016/S0014-5793(96)01351-8
Stadler PF, Fried C, Prohaska SJ, Bailey WJ, Misof BY, Ruddle FH, Wagner GP (2004) Evidence for independent Hox gene duplications in the hagfish lineage: a PCR-based gene inventory of Eptatretus stoutii. Mol Phylogenet Evol 32:686–694. doi:10.1016/j.ympev.2004.03.015
Steinke D, Hoegg S, Brinkmann H, Meyer A (2006) Three rounds (1R/2R/3R) of genome duplications and the evolution of the glycolytic pathway in vertebrates. BMC Biol 4:16. doi:10.1186/1741-7007-4-16
Takehana Y, Nagai N, Matsuda M, Tsuchiya K, Sakaizumi M (2003) Geographic variation and diversity of the cytochrome b gene in Japanese wild populations of medaka, Oryzias latipes. Zool Sci 20:1279–1291. doi:10.2108/zsj.20.1279
Taylor JS, Van de Peer Y, Meyer A (2001a) Genome duplication, divergent resolution and speciation. Trends Genet 17:299–301. doi:10.1016/S0168-9525(01)02318-6
Taylor JS, Van de Peer Y, Braasch I, Meyer A (2001b) Comparative genomics provides evidence for an ancient genome duplication event in fish. Philos Trans R Soc Lond B Biol Sci 356:1661–1679. doi:10.1098/rstb.2001.0975
Taylor JS, Braasch I, Frickey T, Meyer A, Van de Peer Y (2003) Genome duplication, a trait shared by 22000 species of ray-finned fish. Genome Res 13:382–390. doi:10.1101/gr.640303
Thorne JL, Kishino H, Painter IS (1998) Estimating the rate of evolution of the rate of molecular evolution. Mol Biol Evol 15:1647–1657
Van der Hoeven F, Sordino P, Fraudeau N, Izpisúa-Belmonte JC, Duboule D (1996) Teleost HoxD and HoxA genes: comparison with tetrapods and functional evolution of the HoxD complex. Mech Dev 4:9–21. doi:10.1016/0925-4773(95)00455-6
Van de Peer Y, Taylor JS, Braasch I, Meyer A (2001) The ghost of selection past: rates of evolution and functional divergence of anciently duplicated genes. J Mol Evol 53:436–446. doi:10.1007/s002390010233
Van de Peer Y, Taylor JS, Meyer A (2003) Are all fishes ancient polyploids? J Struct Funct Genomics 3:65–73. doi:10.1023/A:1022652814749
Van de Peer Y, Maere S, Meyer A (2009) The evolutionary significance of ancient genome duplications. Nat Rev Genet, in press. doi: 10.1038/nrg2600
Vandepoele K, De Vos W, Taylor JS, Meyer A, Van de Peer Y (2004) Major events in the genome evolution of vertebrates: paranome age and size differ considerably between ray-finned fishes and land vertebrates. Proc Natl Acad Sci U S A 101:1638–1643. doi:10.1073/pnas.0307968100
Venkatesh B (2003) Evolution and diversity of fish genomes. Curr Opin Genet Dev 13:588–592. doi:10.1016/j.gde.2003.09.001
Venkatesh B, Kirkness EF, Loh YH, Halpern AL, Lee AP, Johnson J, Dandona N, Viswanathan LD, Tay A, Venter JC, Strausberg RL, Brenner S (2007) Survey sequencing and comparative analysis of the elephant shark (Callorhinchus milii) genome. PLoS Biol 5:e101. doi:10.1371/journal.pbio.0050101
Vogel G (1998) Doubled genes may explain fish diversity. Science 281:1119–1121. doi:10.1126/science.281.5380.1119
Volff JN (2005) Genome evolution and biodiversity in teleost fish. Heredity 94:280–294. doi:10.1038/sj.hdy.6800635
Werthand CR, Windham MD (1991) A model for divergent, allopatric speciation of polyploid pteridophytes resulting from silencing of duplicate-gene expression. Am Nat 137:515–526. doi:10.1086/285180
Wheeler DA, Srinivasan M, Egholm M, Shen Y, Chen L, McGuire A, He W, Chen YJ, Makhijani V, Roth GT, Gomes X, Tartaro K, Niazi F, Turcotte CL, Irzyk GP, Lupski JR, Chinault C, Song XZ, Liu Y, Yuan Y, Nazareth L, Qin X, Muzny DM, Margulies M, Weinstock GM, Gibbs RA, Rothberg JM (2008) The complete genome of an individual by massively parallel DNA sequencing. Nature 452:872–876. doi:10.1038/nature06884
Wolfe KH (2001) Yesterday’s polyploids and the mystery of diploidization. Nat Rev Genet 2:333–341. doi:10.1038/35072009
Yamanoue Y, Miya M, Inoue JG, Matsuura K, Nishida M (2006) The mitochondrial genome of spotted green pufferfish Tetraodon nigroviridis (Teleostei: Tetraodontiformes) and divergence time estimation among model organisms in fishes. Genes Genet Syst 8:29–39. doi:10.1266/ggs.81.29
Yamanoue Y, Miya M, Matsuura K, Katoh M, Sakai H, Nishida M (2008) A new perspective on phylogeny and evolution of tetraodontiform fishes (Pisces: Acanthopterygii) based on whole mitochondrial genome sequences: Basal ecological diversification? BMC Evol Biol 8:212. doi:10.1186/1471-2148-8-212
Yamanoue Y, Miya M, Matsuura K, Miyazawa S, Tsukamoto N, Doi H, Takahashi H, Mabuchi K, Nishida M, Sakai H (2009) Explosive speciation of Takifugu: another use of fugu as a model system for evolutionary biology. Mol Biol Evol 26:623–629. doi:10.1093/molbev/msn283
Yang Z (1997) PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci 13:555–556. doi:10.1093/bioinformatics/13.5.555
Yu WP, Yew K, Rajasegaran V, Venkatesh B (2007) Sequencing and comparative analysis of fugu protocadherin clusters reveal diversity of protocadherin genes among teleosts. BMC Evol Biol 7:49. doi:10.1186/1471-2148-7-49
Zhang J (2003) Evolution by gene duplication: an update. Trends Ecol Evol 18:292–298. doi:10.1016/S0169-5347(03)00033-8
Zhou R, Cheng H, Tiersch TR (2001) Differential genome duplication and fish diversity. Rev Fish Biol Fish 11:331–337. doi:10.1023/A:1021395506705
Acknowledgments
The manuscript benefited from the comments of two anonymous reviewers. We thank our colleagues at the Ocean Research Institute and the Graduate School of Frontier Sciences of the University of Tokyo, and the National Institute of Genetics for helpful discussions and comments. This work was partially supported by Grants-in-Aid from the Japan Society for the Promotion of Science to MN.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sato, Y., Nishida, M. Teleost fish with specific genome duplication as unique models of vertebrate evolution. Environ Biol Fish 88, 169–188 (2010). https://doi.org/10.1007/s10641-010-9628-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10641-010-9628-7