Abstract
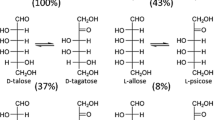
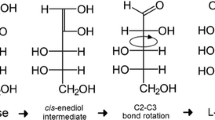
The activity of ribose-5-phosphate isomerases (RpiB) from Clostridium difficile for d-ribose isomerization was optimal at pH 7.5 and 40°C, while that from Thermotoga maritima for l-talose isomerization was optimal at pH 8.0 and 70°C. C. difficile RpiB exhibited activity only with aldose substrates possessing hydroxyl groups oriented in the right-handed configuration (Fischer projections) at the C2 and C3 positions, such as d-ribose, d-allose, l-talose, l-lyxose, d-gulose, and l-mannose. In contrast, T. maritima RpiB displayed activity only with aldose substrates possessing hydroxyl groups configured the same direction at the C2, C3, and C4 positions, such as the d- and l-forms of ribose, talose, and allose.



Similar content being viewed by others
References
Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M (1983) CHARMM: a program for macromolecular energy, minimization, and dynamics calculations. J Comput Chem 4:187–217
Goodsell DS, Morris GM, Olson AJ (1996) Automated docking of flexible ligands: applications of AutoDock. J Mol Recognit 9:1–5
Hossain MA, Wakabayashi H, Goda F, Kobayashi S, Maeba T, Maeta H (2000) Effect of the immunosuppressants FK506 and D-allose on allogenic orthotopic liver transplantation in rats. Transplant Proc 32:2021–2023
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291
Park CS, Yeom SJ, Kim HJ, Lee SH, Lee JK, Kim SW, Oh DK (2007a) Characterization of ribose-5-phosphate isomerase of Clostridium thermocellum producing d-allose from d-psicose. Biotechnol Lett 29:1387–1391
Park HY, Park CS, Kim HJ, Oh DK (2007b) Substrate specificity of a galactose 6-phosphate isomerase from Lactococcus lactis that produces d-allose from d-psicose. J Biotechnol 132:88–95
Poulsen TS, Chang YY, Hove-Jensen B (1999) d-Allose catabolism of Escherichia coli: involvement of alsI and regulation of als regulon expression by allose and ribose. J Bacteriol 181:7126–7130
Sali A, Blundell TL (1993) Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol 234:779–815
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Yeom SJ, Ji JH, Kim NH, Park CS, Oh DK (2009) Substrate specificity of a mannose-6-phosphate isomerase from Bacillus subtilis and its application in the production of l-ribose. Appl Environ Microbiol 75:4705–4710
Yoon RY, Yeom SJ, Kim HJ, Oh DK (2009a) Novel substrates of a ribose-5-phosphate isomerase from Clostridium thermocellum. J Biotechnol 139:26–32
Yoon RY, Yeom SJ, Park CS, Oh DK (2009b) Substrate specificity of a glucose-6-phosphate isomerase from Pyrococcus furiosus for monosaccharides. Appl Microbiol Biotechnol 83:295–303
Zhang R, Andersson CE, Savchenko A, Skarina T, Evdokimova E, Beasley S, Arrowsmith CH, Edwards AM, Joachimiak A, Mowbray SL (2003a) Structure of Escherichia coli ribose-5-phosphate isomerase: a ubiquitous enzyme of the pentose phosphate pathway and the Calvin cycle. Structure 11:31–42
Zhang RG, Andersson CE, Skarina T, Evdokimova E, Edwards AM, Joachimiak A, Savchenko A, Mowbray SL (2003b) The 2.2Å resolution structure of RpiB/AlsB from Escherichia coli illustrates a new approach to the ribose-5-phosphate isomerase reaction. J Mol Biol 332:1083–1094
Acknowledgments
This study was supported by a grant (R0A-2007-000-20015-0) from the National Research Lab. Program, Ministry of Education, Science and Technology.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yeom, SJ., Kim, BN., Park, CS. et al. Substrate specificity of ribose-5-phosphate isomerases from Clostridium difficile and Thermotoga maritima . Biotechnol Lett 32, 829–835 (2010). https://doi.org/10.1007/s10529-010-0224-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-010-0224-x