Abstract
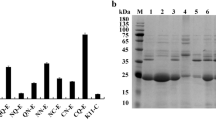
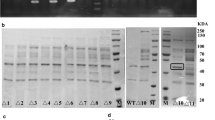
Bacillus acidopullulyticus pullulanase (BaPul13A) is a widely used debranching enzyme in the starch industry. A few details have been reported on the heterologous expression of BaPul13A in Escherichia coli (E. coli). This study compares different E. coli expression systems to improve the soluble expression level of BaPul13A. When pET22b(+)/pET28a(+) was used as the expression vector, the soluble expression of BaPul13A can be achieved by tightly controlling basal expression, whereas pET-20b(+)/pGEX4T2 leads to insoluble inclusion bodies. An efficient process control strategy aimed at minimizing the formation of inclusion bodies and enhancing the production of pullulanase was developed by a step decrease of the temperature in a 5-L fermentor. The highest total enzyme activity of BaPul13A reached 1,156.32 U/mL. This work reveals that the T7 promoter with lac operator and lacI gene collectively contribute to the soluble expression of BaPul13A, whereas either a T7 promoter alone or combined with the lac operator and lacI gene results in poor solubility. Basal expression in the initial growth phase of the host significantly affects the solubility of BaPul13A in E. coli.






Similar content being viewed by others
References
Baig F, Fernando LP, Salazar MA, Powell RR, Bruce TF, Harcum SW (2014) Dynamic transcriptional response of Escherichia coli to inclusion body formation. Biotechnol Bioeng 111(5):980–999. doi:10.1002/bit.25169
Basu A, Li X, Leong SS (2011) Refolding of proteins from inclusion bodies: rational design and recipes. Appl Microbiol Biotechnol 92(2):241–251. doi:10.1007/s00253-011-3513-y
Capitini C, Conti S, Perni M, Guidi F, Cascella R, De Poli A, Penco A, Relini A, Cecchi C, Chiti F (2014) TDP-43 inclusion bodies formed in bacteria are structurally amorphous, non-amyloid and inherently toxic to neuroblastoma cells. PLoS One 9(1):e86720. doi:10.1371/journal.pone.0086720
de Groot NS, Espargaro A, Morell M, Ventura S (2008) Studies on bacterial inclusion bodies. Future Microbiol 3(4):423–435. doi:10.2217/17460913.3.4.423
Duan X, Chen J, Wu J (2013) Optimization of pullulanase production in Escherichia coli by regulation of process conditions and supplement with natural osmolytes. Bioresour Technol 146:379–385. doi:10.1016/j.biortech.2013.07.074
Dubendorff JW, Studier FW (1991) Controlling basal expression in an inducible T7 expression system by blocking the target T7 promoter with lac repressor. J Mol Biol 219(1):45–59. doi:10.1016/0022-2836(91)90856-2
Garcia-Fruitos E, Vazquez E, Diez-Gil C, Corchero JL, Seras-Franzoso J, Ratera I, Veciana J, Villaverde A (2012) Bacterial inclusion bodies: making gold from waste. Trends Biotechnol 30(2):65–70. doi:10.1016/j.tibtech.2011.09.003
Gatti-Lafranconi P, Natalello A, Ami D, Doglia SM, Lotti M (2011) Concepts and tools to exploit the potential of bacterial inclusion bodies in protein science and biotechnology. FEBS J 278(14):2408–2418. doi:10.1111/j.1742-4658.2011.08163.x
Hoffmann F, Posten C, Rinas U (2001) Kinetic model of in vivo folding and inclusion body formation in recombinant Escherichia coli. Biotechnol Bioeng 72(3):315–322
Lappalainen A, Nikupaavola ML, Suortti T, Poutanen K (1991) Purification and characterization of Bacillus-Acidopullulyticus pullulanase for enzymatic starch modification. Starch-Starke 43(12):477–482. doi:10.1002/star.19910431207
Martinez-Alonso M, Garcia-Fruitos E, Ferrer-Miralles N, Rinas U, Villaverde A (2010) Side effects of chaperone gene co-expression in recombinant protein production. Microb Cell Fact 9:64. doi:10.1186/1475-2859-9-64
McNulty DE, Claffee BA, Huddleston MJ, Porter ML, Cavnar KM, Kane JF (2003) Mistranslational errors associated with the rare arginine codon CGG in Escherichia coli. Protein Expr Purif 27(2):365–374
Mertens N, Remaut E, Fiers W (1995) Tight transcriptional control mechanism ensures stable high-level expression from T7 promoter-based expression plasmids. Biotechnology (N Y) 13(2):175–179
Mikami B, Iwamoto H, Malle D, Yoon HJ, Demirkan-Sarikaya E, Mezaki Y, Katsuya Y (2006) Crystal structure of pullulanase: evidence for parallel binding of oligosaccharides in the active site. J Mol Biol 359(3):690–707. doi:10.1016/j.jmb.2006.03.058
Miroux B, Walker JE (1996) Over-production of proteins in Escherichia coli: mutant hosts that allow synthesis of some membrane proteins and globular proteins at high levels. J Mol Biol 260(3):289–298. doi:10.1006/jmbi.1996.0399
Rajan RS, Illing ME, Bence NF, Kopito RR (2001) Specificity in intracellular protein aggregation and inclusion body formation. Proc Natl Acad Sci USA 98(23):13060–13065. doi:10.1073/pnas.181479798
Reetz MT, Carballeira JD (2007) Iterative saturation mutagenesis (ISM) for rapid directed evolution of functional enzymes. Nat Protoc 2(4):891–903. doi:10.1038/nprot.2007.72
Rokney A, Shagan M, Kessel M, Smith Y, Rosenshine I, Oppenheim AB (2009) E. coli transports aggregated proteins to the poles by a specific and energy-dependent process. J Mol Biol 392(3):589–601. doi:10.1016/j.jmb.2009.07.009
Sabate R, de Groot NS, Ventura S (2010) Protein folding and aggregation in bacteria. Cell Mol Life Sci 67(16):2695–2715. doi:10.1007/s00018-010-0344-4
Schlegel S, Rujas E, Ytterberg AJ, Zubarev RA, Luirink J, de Gier JW (2013) Optimizing heterologous protein production in the periplasm of E. coli by regulating gene expression levels. Microb Cell Fact 12:24. doi:10.1186/1475-2859-12-24
Schugerl K, Hubbuch J (2005) Integrated bioprocesses. Curr Opin Microbiol 8(3):294–300. doi:10.1016/j.mib.2005.01.002
Sorensen HP, Mortensen KK (2005) Advanced genetic strategies for recombinant protein expression in Escherichia coli. J Biotechnol 115(2):113–128. doi:10.1016/j.jbiotec.2004.08.004
Sorensen HP, Mortensen KK (2005) Soluble expression of recombinant proteins in the cytoplasm of Escherichia coli. Microb Cell Fact 4(1):1. doi:10.1186/1475-2859-4-1
Turkenburg JP, Brzozowski AM, Svendsen A, Borchert TV, Davies GJ, Wilson KS (2009) Structure of a pullulanase from Bacillus acidopullulyticus. Proteins 76(2):516–519. doi:10.1002/prot.22416
Ushasree MV, Vidya J, Pandey A (2014) Gene cloning and soluble expression of Aspergillus niger phytase in E. coli cytosol via chaperone co-expression. Biotechnol Lett 36(1):85–91. doi:10.1007/s10529-013-1322-3
Ventura S, Villaverde A (2006) Protein quality in bacterial inclusion bodies. Trends Biotechnol 24(4):179–185. doi:10.1016/j.tibtech.2006.02.007
Voulgaridou GP, Mantso T, Chlichlia K, Panayiotidis MI, Pappa A (2013) Efficient E. coli expression strategies for production of soluble human crystallin ALDH3A1. PLoS One 8(2):e56582. doi:10.1371/journal.pone.0056582
Wu W, Xing L, Zhou B, Lin Z (2011) Active protein aggregates induced by terminally attached self-assembling peptide ELK16 in Escherichia coli. Microb Cell Fact 10:9. doi:10.1186/1475-2859-10-9
Yang H, Liu L, Shin HD, Li J, Du G, Chen J (2013) Structure-guided systems-level engineering of oxidation-prone methionine residues in catalytic domain of an alkaline alpha-amylase from Alkalimonas amylolytica for significant improvement of both oxidative stability and catalytic efficiency. PLoS One 8(3):e57403. doi:10.1371/journal.pone.0057403
Acknowledgments
The authors would like to thank Wenwen Guo and Lu Li for the drawing figures. This work was supported by the National Basic Research Program of China (973 Program) (Grant No.: 2013CB733602) and the Fundamental Research Funds for the Central Universities (Grant No. JUSRP51401A).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Chen, A., Li, Y., Liu, X. et al. Soluble expression of pullulanase from Bacillus acidopullulyticus in Escherichia coli by tightly controlling basal expression. J Ind Microbiol Biotechnol 41, 1803–1810 (2014). https://doi.org/10.1007/s10295-014-1523-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-014-1523-3