Abstract
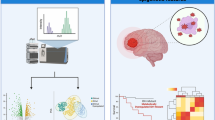
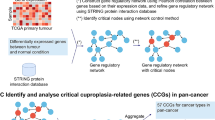
Malignant astrocytomas of World Health Organization (WHO) grade III or IV have a reduced median survival time, and possible pathways have been described for the progression of anaplastic astrocytomas and glioblastomas, but the molecular basis of malignant astrocytoma progression is still poorly understood. Microarray analysis provides the chance to accelerate studies by comparison of the expression of thousands of genes in these tumours and consequently identify targeting genes. We compared the transcriptional profile of 4,608 genes in tumours of 15 patients including 6 anaplastic astrocytomas (WHO grade III) and 9 glioblastomas (WHO grade IV) using microarray analysis. The microarray data were corroborated by real-time reverse transcription-polymerase chain reaction analysis of two selected genes. We identified 166 gene alterations with a fold change of 2 and higher whose mRNA levels differed (absolute value of the t statistic of 1.96) between the two malignant glioma groups. Further analyses confirmed same transcription directions for Olig2 and IL-13Rα2 in anaplastic astrocytomas as compared to glioblastomas. Microarray analyses with a close binary question reveal numerous interesting candidate genes, which need further histochemical testing after selection for confirmation. IL-13Rα2 and Olig2 have been identified and confirmed to be interesting candidate genes whose differential expression likely plays a role in malignant progression of astrocytomas.

Similar content being viewed by others
References
Bouvier C, Bartoli C, Guirre-Cruz L, Virard I, Colin C, Fernandez C et al (2003) Shared oligodendrocyte lineage gene expression in gliomas and oligodendrocyte progenitor cells. J Neurosurg 99(2):344–350
Chakravarti A, Zhai G, Suzuki Y, Sarkesh S, Black PM, Muzikansky A et al (2004) The prognostic significance of phosphatidylinositol 3-kinase pathway activation in human gliomas. J Clin Oncol 22(10):1926–1933
Debinski W, Gibo DM (2000) Molecular expression analysis of restrictive receptor for interleukin 13, a brain tumour-associated cancer/testis antigen. Mol Med 6(5):440–449
Freije WA, Castro-Vargas FE, Fang Z, Horvath S, Cloughesy T, Liau LM et al (2004) Gene expression profiling of gliomas strongly predicts survival. Cancer Res 64(18):6503–6510
Fukuda S, Kondo T, Takebayashi H, Taga T (2004) Negative regulatory effect of an oligodendrocytic bHLH factor OLIG2 on the astrocytic differentiation pathway. Cell Death Differ 11(2):196–202
Fuller GN, Hess KR, Rhee CH, Yung WK, Sawaya RA, Bruner JM et al (2002) Molecular classification of human diffuse gliomas by multidimensional scaling analysis of gene expression profiles parallels morphology-based classification, correlates with survival, and reveals clinically-relevant novel glioma subsets. Brain Pathol 12(1):108–116
Giulietti A, Overbergh L, Valckx D, Decallonne B, Bouillon R, Mathieu C (2001) An overview of real-time quantitative PCR: applications to quantify cytokine gene expression. Methods 25(4):386–401
Hoelzinger DB, Mariani L, Weis J, Woyke T, Berens TJ, McDonough WS et al (2005) Gene expression profile of glioblastoma multiforme invasive phenotype points to new therapeutic targets. Neoplasia 7(1):7–16
Ichimura K, Ichimura K, Ohgaki H, Kleihues P, Collins VP (2004) Molecular pathogenesis of astrocytic tumours. J Neurooncol 70(2):137–160
Joshi BH, Plautz GE, Puri RK (2000) Interleukin-13 receptor alpha chain: a novel tumour-associated transmembrane protein in primary explants of human malignant gliomas. Cancer Res 60(5):1168–1172
Kawakami K, Kawakami M, Snoy PJ, Husain SR, Puri RK (2001) In vivo overexpression of IL-13 receptor alpha2 chain inhibits tumourigenicity of human breast and pancreatic tumours in immunodeficient mice. J Exp Med 194(12):1743–1754
Kawakami K, Kawakami K, Taguchi J, Murata T, Puri RK (2005) Evidence that IL-13R alpha2 chain in human glioma cells is responsible for the antitumour activity mediated by receptor-directed cytotoxin therapy. J Immunother 28(3):193–202
Kawakami K, Kawakami K, Kioi M, Liu Q, Kawakami M, Puri RK (2001) The interleukin-13 receptor alpha2 chain: an essential component for binding and internalization but not for interleukin-13-induced signal transduction through the STAT6 pathway. Blood 97(9):2673–2679
Kawakami M, Kawakami M, Kawakami K, Takahashi S, Abe M, Puri RK (2004) Analysis of interleukin-13 receptor alpha2 expression in human pediatric brain tumours. Cancer 101(5):1036–1042
Lang FF, Bruner JM, Fuller GN, Aldape K, Prados MD, Chang S et al (2003) Phase I trial of adenovirus-mediated p53 gene therapy for recurrent glioma: biological and clinical results. J Clin Oncol 21(13):2508–2518
Ligon KL, Alberta JA, Kho AT, Weiss J, Kwaan MR, Nutt CL et al (2004) The oligodendroglial lineage marker OLIG2 is universally expressed in diffuse gliomas. J Neuropathol Exp Neurol 63(5):499–509
Liu TF, Tatter SB, Willingham MC, Yang M, Hu JJ, Frankel AE (2003) Growth factor receptor expression varies among high-grade gliomas and normal brain: epidermal growth factor receptor has excellent properties for interstitial fusion protein therapy. Mol Cancer Ther 2(8):783–787
Lu QR, Park JK, Noll E, Chan JA, Alberta J, Yuk D et al (2001) Oligodendrocyte lineage genes (OLIG) as molecular markers for human glial brain tumours. Proc Natl Acad Sci U S A 98(19):10851–10856
Mariani L, Beaudry C, McDonough WS, Hoelzinger DB, Demuth T, Ross KR et al (2001) Glioma cell motility is associated with reduced transcription of proapoptotic and proliferation genes: a cDNA microarray analysis. J Neurooncol 53(2):161–176
Mischel PS, Shai R, Shi T, Horvath S, Lu KV, Choe G et al (2003) Identification of molecular subtypes of glioblastoma by gene expression profiling. Oncogene 22(15):2361–2373
Mokhtari K, Paris S, guirre-Cruz L, Privat N, Criniere E, Marie Y et al (2005) Olig2 expression, GFAP, p53 and 1p loss analysis contribute to glioma subclassification. Neuropathol Appl Neurobiol 31(1):62–69
Ohgaki H (2005) Genetic pathways to glioblastomas. Neuropathology 25(1):1–7
Pomeroy SL, Tamayo P, Gaasenbeek M, Sturla LM, Angelo M, McLaughlin ME et al (2002) Prediction of central nervous system embryonal tumour outcome based on gene expression. Nature 415(6870):436–442
Reifenberger G, Collins VP (2004) Pathology and molecular genetics of astrocytic gliomas. J Mol Med 82(10):656–670
Rich JN, Reardon DA, Peery T, Dowell JM, Quinn JA, Penne KL et al (2004) Phase II trial of gefitinib in recurrent glioblastoma. J Clin Oncol 22(1):133–142
Rickman DS, Bobek MP, Misek DE, Kuick R, Blaivas M, Kurnit DM et al (2001) Distinctive molecular profiles of high-grade and low-grade gliomas based on oligonucleotide microarray analysis. Cancer Res 61(18):6885–6891
Riemenschneider MJ, Koy TH, Reifenberger G (2004) Expression of oligodendrocyte lineage genes in oligodendroglial and astrocytic gliomas. Acta Neuropathol (Berl) 107(3):277–282
Sanson M, Thillet J, Hoang-Xuan K (2004) Molecular changes in gliomas. Curr Opin Oncol 16(6):607–613
Schramm A, Schulte JH, Klein-Hitpass L, Havers W, Sieverts H, Berwanger B et al (2005) Prediction of clinical outcome and biological characterization of neuroblastoma by expression profiling. Oncogene 24(53):7902–7912
Setoguchi T, Kondo T (2004) Nuclear export of OLIG2 in neural stem cells is essential for ciliary neurotrophic factor-induced astrocyte differentiation. J Cell Biol 166(7):963–968
Shai R, Shi T, Kremen TJ, Horvath S, Liau LM, Cloughesy TF et al (2003) Gene expression profiling identifies molecular subtypes of gliomas. Oncogene 22(31):4918–4923
Thome J, Gsell W, Rosler M, Kornhuber J, Frolich L, Hashimoto E et al (1997) Oxidative-stress associated parameters (lactoferrin, superoxide dismutases) in serum of patients with Alzheimer's disease. Life Sci 60(1):13–19
van den, Boom J, Wolter M, Kuick R, Misek DE, Youkilis AS, Wechsler DS et al (2003) Characterization of gene expression profiles associated with glioma progression using oligonucleotide-based microarray analysis and real-time reverse transcription-polymerase chain reaction. Am J Pathol 163(3):1033–1043
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A et al (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3(7):RESEARCH0034.
Wang J, Jani-Sait SN, Escalon EA, Carroll AJ, de Jong PJ, Kirsch IR et al (2000) The t(14;21)(q11.2;q22) chromosomal translocation associated with T-cell acute lymphoblastic leukemia activates the BHLHB1 gene. Proc Natl Acad Sci U S A 97(7):3497–3502
Wennerberg K, Rossman KL, Der CJ (2005) The Ras superfamily at a glance. J Cell Sci 118(Pt 5):843–846
Yang YH, Dudoit S, Luu P, Lin DM, Peng V, Ngai J et al (2002) Normalization for cDNA nicroarray data: a robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res 30(4):e15
Zhou Q, Anderson DJ (2002) The bHLH transcription factors OLIG2 and OLIG1 couple neuronal and glial subtype specification. Cell 109(1):61–73
Acknowledgement
For these studies, the first author (OB) has been financed by the Kempkes Foundation Marburg, Germany. We like to thank Prof. Eilers and Prof. Neubauer and their co-workers very much for their intensive assistance, help and collaboration to finish these studies. Special thanks to Dr. Krause and his microarray facility.
Author information
Authors and Affiliations
Corresponding author
Additional information
Comments
Dietmar Krex, Dresden, Germany
In the present study by Bozinov et al., microarray analysis was performed in six anaplastic astrocytomas and nine GBMs (WHO grade III and IV, respectively) to identify molecular markers associated with progression and further dedifferentiation in malignant gliomas. Although there are numerous published data about the molecular differences between low- and high-grade gliomas, only little is known about molecular markers that are involved in late-stage malignization of glial tumours. Microarray-based approaches are reasonable to screen for candidate genes, although specificity of these analyses is low while costs are high. The latter is also a reason why in most studies only small numbers of patients are studied. To validate the results, and to identify promising candidate genes, data can be pooled from various analyses. Therefore, there is a clear need for more array-based approaches. Of course, like in the present study, results additionally need to be confirmed by another experimental approach. Bozinov et al. have picked two interesting genes Olig2 and IL-13Rα2 that were confirmed to be up- and downregulated, respectively, in glioblastomas as compared to anaplastic astrocytomas. Particularly the downregulation of the IL-13Rα2 gene is challenging and needs confirmation in larger series, as it is in contrast to several preclinical and clinical studies and indicates that the IL-13 receptor might not be a perfect target for a molecular therapy in a pure glioblastoma population.
Comments
Kaoru Kurisu, Hiroshima, Japan
The authors emphasized the usefulness of microarray analysis in the determination of affected genes. In addition, they pointed out new transcriptional networks which concern the up and downregulation of signaling pathways. This approach is very beneficial to pick up the affected genes, but after obtaining the results, we have to check the function of them as mentioned by the authors in the conclusion. On the point of treatment application of these results, Olig2 and IL-13Rα2 are very fascinating to develop new molecular targeting to control tumor growth, invasion and dissemination on so on. Of course, after initial treatment, neoplasm can get another function as tolerance against the first treatment drugs and molecules. Therefore, we have to identify the advanced stage function of these candidates. In our department, survival was identified as a good prognostic factor in astrocytic tumors (Cancer. 2003; 97(4):1077–83) and its intracellular expression pattern can indicate the prognosis of high-grade astrocytomas (J Neurooncol. 2007 Apr; 82(2):193–8). Therefore, expression pattern or intracellular localization of the candidate factors is very important to understand the biological behavior of them. I hope the authors will develop their integrated works on these affected genes and related factors to overcome the worst neoplasm of central nervous system, glioblastoma.
An erratum to this article can be found at http://dx.doi.org/10.1007/s10143-008-0125-9
Rights and permissions
About this article
Cite this article
Bozinov, O., Köhler, S., Samans, B. et al. Candidate genes for the progression of malignant gliomas identified by microarray analysis. Neurosurg Rev 31, 83–90 (2008). https://doi.org/10.1007/s10143-007-0107-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10143-007-0107-3