Abstract
Background
Gastric cancer (GC) patients with PD-L1-negative tumor occasionally have a favorable response to anti-PD-1 mAb. The aim of the present study was to investigate the regulatory mechanism and immunosuppressive role of PD-L2 in GC.
Methods
We used immunohistochemistry to evaluate the expression of PD-L2 in primary tumors from 194 patients with GC. The mechanism of PD-L2 expression was assessed in TCGA stomach adenocarcinoma tissue dataset and in vitro assay using GC cell lines. The immunosuppressive role of PD-L2 was evaluated by cytotoxicity of CTL clone against PD-L2 expressing GC cells.
Results
PD-L2 was expressed on tumor cells (TCs) of 28.4% patients and PD-L2 expression on TCs was significantly associated with tumor progression. TCGA dataset revealed that IFN-γ and, to a lesser extent, IL-4 signature significantly correlated with PD-L2 expression. In vitro assay showed that IFN-γ and, also to a lesser extent, IL-4 can upregulate PD-L2 expression on GC cells. Anti-PD-L2 mAb significantly enhanced the cytotoxicity of CTL clone against GC cell lines expressing PD-L2.
Conclusions
PD-L2 is expressed on GC cells and PD-1/PD-L2 interaction are functionally involved in anti-tumor CTL activities. PD-L2 expression should be considered when determining the optimal immunotherapy for GC.
Similar content being viewed by others
Background
Gastric cancer (GC) is the fifth most frequently diagnosed cancer and the third leading cause of cancer death in the world, accounting for over 1,000,000 new cases and 783,000 deaths in 2018 worldwide [1]. Especially in East Asia including Japan and Korea, incidence rates are markedly elevated [1]. Combinations of surgical resection, diverse chemotherapy, and radiotherapy are used to treat advanced GC patients; however, the 5-year overall survival rates of patients with pathological stage IV disease is still 16.4% in Japan [2]. To improve the prognosis of advanced GC patients, the development of a novel therapeutic strategy is required.
Immune checkpoint blockade targeting the programmed cell death 1 (PD-1) axis has been approved to treat various human cancers including GC [3,4,5,6,7]. Although an immune checkpoint blockade with anti-PD-1 mAb has been also expected for advanced GC patients, its clinical efficacy is limited with an objective response rate of 11.9% [3, 8]. Therefore, more effective therapeutic strategies and predictive biomarkers to identify responders for immune checkpoint blockade targeting the PD-1 axis are urgently needed.
PD-1 is mainly expressed on activated T cells, B cells, and natural killer cells, and binds to programmed death ligand-1 (PD-L1) and PD-L2 [9, 10]. PD-L1 is expressed on both various types of tumor cells (TCs) and tumor-infiltrating immune cells (TIICs) [11,12,13]. On the other hand, although it was initially thought that PD-L2 is mainly expressed on macrophage in the presence of interleukin (IL)-4/IL-13 [14, 15], several studies recently reported that PD-L2 is also expressed on various types of TCs, depending on tumor microenvironment situation [16,17,18]. Activation of the PD-1 signaling pathway leads T cells to apoptosis and anergy, resulting in immunosuppression caused by T cell dysfunction [19, 20]. Immune checkpoint blockade targeting PD-1 inhibits the receptor interaction with both PD-L1 and PD-L2.
In clinical studies of anti-PD-1 mAb, it has been reported that patients with PD-L1 expressing tumor had a favorable clinical course [21, 22]. However, some patients with PD-L1 expressing tumor have been reported to have responded poorly to anti-PD-1 mAb, whereas patients with PD-L1-negative tumor occasionally have clinical efficacy in several types of cancer including GC [3, 7, 21,22,23,24,25,26,27]. Furthermore, Yearley et al. suggested that PD-L2 expression may provide information in predictive clinical response to anti-PD-1 mAb [16]. Although these findings indicate the immunosuppressive role of PD-L2, its role has not yet been fully elucidated. In the present study, we investigated the regulatory mechanism, expression status, and immunosuppressive role of PD-L2 in GC using surgically-resected specimens, The Cancer Genome Atlas (TCGA) stomach adenocarcinoma tissue dataset, the cytotoxicity of tumor antigen-specific cytotoxic T lymphocyte (CTL) clone against GC cells, and in vitro assay using GC cell lines.
Methods
Clinical samples
Surgically-resected specimens were obtained from 194 patients who had undergone surgical resection for GC at the Department of Gastrointestinal Tract Surgery, Fukushima Medical University Hospital, between January 2003 and December 2013. No patients had received pre-operative anti-tumor therapy such as radiotherapy or chemotherapy. Clinical and pathological information was retrospectively obtained by reviewing the patients’ medical records. Tumor grade and stage were defined in accordance with the Japanese GC Association (The 15th Edition). The study was conducted in accordance with the Declaration of Helsinki and was approved by the Institutional Ethical Committee of Fukushima Medical University (Reference Nos. 2329 and 2847). Written informed consent was obtained from all participants.
Immunohistochemistry staining
Four-μm-thick sections were deparaffinized and rehydrated. Endogenous peroxidases were blocked with 0.3% hydrogen peroxide in methanol and antigen retrieval was performed by autoclave for 10 min in Target Retrieval Solution (Agilent Technologies, Inc., Santa Clara, CA, USA) (120 °C, pH9.0). Thereafter, the slides were incubated at 4 °C overnight with the following primary antibodies: PD-L1 mAb (Cell Signaling Technology, Inc., Danvers, MA, USA) at 1:400, and PD-L2 mAb (Cell Signaling Technology, Inc.) at 1:200. Followed by incubation, the detection was performed with an HRP-coupled anti-rabbit polymer (Envision + System-HRP, Agilent Technologies, Inc.). The sections were then incubated with diaminobenzidine (Agilent Technologies, Inc.) at room temperature for 5 min, and counterstained with Mayer's Hematoxylin Solution (FUJIFILM Wako Pure Chemical Corp., Osaka, Japan), also at room temperature, for 1 min.
Assessment of immunohistochemistry staining
Immunohistochemistry (IHC) evaluation was performed by two independent observers (Y.N. and K.S), who were blinded to the clinical data. The expression of PD-L1 and PD-L2 was evaluated by assessing membranous staining of TCs and TIICs; ≥ 1% was defined as positive and < 1% was defined as negative [7, 21, 26, 28].
RNA-seq analysis using TCGA dataset
The mRNA expression z-scores of genes (RNA-Seq V2 RSEM normalized, RNA-Seq data) were obtained from TCGA stomach adenocarcinoma tissue dataset (n = 269) through cBioPortal (https://www.cbioportal.org/) [29, 30]. In the present study, we evaluated the mRNA expression levels of PD-1 (PDCD1), PD-L1 (CD274), and PD-L2 (PDCD1LG2), Interferon (IFN)-γ gene signature, and interleukin (IL)-4 gene signature in GC. Samples with gene amplification for PD-L1 or PD-L2 were excluded for analysis. IFN-γ gene signature included indoleamine 2,3-dioxygenase 1 (IDO1), C-X-C motif chemokine ligand (CXCL)10, CXCL9, human leukocyte antigen (HLA)-DRA, signal transducer and activator of transcription 1 (STAT1) and IFN-γ [31], although TCGA RNA-seq data lacked HLA-DRA expression values. The BIOCARTA_IL4_PATHWAY was obtained from Gene Set Enrichment Analysis (GSEA) through Explore the Molecular Signatures Database (MSigDB) (https://software.broadinstitute.org/gsea/msigdb/index.jsp) and we used it as the IL-4 gene signature including AKT1, growth factor receptor-bound protein 2 (GRB2), IL-4, IL-4R, insulin receptor substrate 1 (IRS1), Janus kinase (JAK)1, JAK3, ribosomal protein S6 kinase B1 (RPS6KB1), SHC1, and STAT6 [32,33,34]. The signature score was calculated by averaging the expression levels of included genes [35, 36].
Tumor cell lines
MKN7, MKN45, OCUM-1, and NUGC-3 were purchased from the Japanese Collection of Research Bioresources Cell Bank (Osaka, Japan). NCI-N87 was purchased from the American Type Culture Collection (Manassas, VA, USA), and ECC10, GSU, HGC27, KE39, and NUGC-4 were purchased from the RIKEN BioResource Research Center (Ibaraki, Japan). These cell lines have no gene amplification or deletion for PD-L1 and PD-L2 according to the cell line data from each company. For PCR, KATO III was purchased from the American Type Culture Collection and OE19 was purchased from the Merck KGaA (Darmstadt, Germany). All cell lines, in which the absence of mycoplasma was confirmed, were cultured in RPMI-1640 containing l-glutamine (Merck KGaA) with 10% fetal bovine serum (Thermo Fisher Scientific, Inc., Waltham, MA, USA) and 1% Penicillin–Streptomycin (Thermo Fisher Scientific, Inc.), and were verified as authentic through short tandem repeat profiling.
Generation of CTL clone
HLA-A24 restricted, Kinesin family member 20A (KIF20A) peptide-specific CTL clone was established using HLA-A24 positive peripheral blood mononuclear cells (PBMC) from advanced GC patients as previously described [37]. Briefly, T cells were stimulated with KIF20A peptide-loaded, autologous mature dendritic cells every 7 days. After the third stimulation, the KIF20A peptide specificity of the CTL lines was tested by enzyme-linked immunospot (ELISpot) assay. CTL clones specific for KIF20A peptide were established from an HLA-A24 restricted, KIF20A peptide-specific CTL line using a limiting dilution method.
Cell treatment with cytokines and inhibitors
We decided the optimal doses of IFN-γ (R&D Systems, Minneapolis, MN, USA) and IL-4 (PeproTech, Inc., Rocky Hill, NJ, USA) for the PD-L1 and PD-L2 expression on tumor cell lines according to our previous study [37,38,39] (data not shown). Tumor cell lines were cultured in a 12-well plate and exposed to 10 ng/mL IFN-γ or 10 ng/mL IL-4. PBS was used as a negative control, and was added to all controls. The cells were used for western blotting for cell signaling pathway and gene expression microarray after 1 h incubation, and were used for flow cytometry and western blotting for PD-L1 and PD-L2 after incubation for 48 h.
For the blocking assay, the CTL clone and lymphocytes were incubated with or without 10 μg/mL anti-PD-1 mAb (Thermo Fisher Scientific, Inc.), and the tumor cell lines were incubated with or without 10 μg/mL anti-PD-L1 mAb (Thermo Fisher Scientific, Inc.) and/or 10 μg/mL anti-PD-L2 mAb (Thermo Fisher Scientific, Inc.) for 1 h before co-culture experiment, ELISpot assay, and cytotoxic assay. Each dose of these blocking antibodies was recommended according to the manufacture’s instructions.
Co-culture experiment
IL-2 activated lymphocytes were generated as previously described [40]. PD-1 expression on T cells in IL-2 activated lymphocytes were evaluated by flow cytometry using APC-H7 conjugated anti-human CD3 mAb (BD Biosciences, San Jose, CA, UAS) at 1:20, and PerCP-CyTM5.5 conjugated anti-human CD279 (PD-1) mAb (BD Biosciences) at 1:20. IL-2 activated lymphocytes treated with/without anti-PD-1 mAb were co-cultured with NUGC-3 treated with/without inhibitors at a 1:1 ratio in 24-well plates for 48 h. After a 48 h incubation, the proportion of apoptotic CD3-positive cells, T cells, were analyzed with PE-conjugated Annexin V and 7-Aminoactinomycin D (7-AAD) (PE Annexin V Apoptosis Detection Kit I, BD Biosciences) using flow cytometry.
Cytotoxic assay
Cytotoxicity of the CTL clone was measured using a calcein-release assay as previously described [37,38,39]. Briefly, target cells were stained with 5 μM of calcein-AM (Dojindo Molecular Technologies, Inc., Kumamoto, Japan) for 30 min at 37 °C. The stained targets (5 × 103/well) were then co-cultured at various ratios of the CTL clone in 200 µL of culture medium for 4 h. Assays were performed in triplicate in a 96-well U-bottomed plate. After incubation, 100 μL of the supernatant was transferred into a 96-well flat-bottomed plate, and the fluorescence of each supernatant was measured using an Infinite 200 plate reader (Tecan Group Ltd., Männedorf, Switzerland). Spontaneous release was obtained from target cells incubated without effector cells, and maximum release was obtained from detergent-released target cells. The percentage of specific lysis was calculated according to the formula: %specific lysis = 100 × (experimental release − spontaneous release)/(maximum release − spontaneous release).
ELISpot assay
IFN-γ ELISpot assay was performed using a commercial kit (Mabtech, Stockholm, Sweden) to determine the CTL response [37, 38]. Briefly, 96-well plates with nitrocellulose membranes (Merck KGaA) were pre-coated with the primary anti-IFN-γ antibody at 4 °C overnight. After blocking with AIM-V medium containing 5% human serum, target cells (2 × 104/well) and KIF20A peptide-specific CTL clones (2 × 103/well) were co-cultured in 200 µL of culture medium at 37 °C for 24 h. These wells were treated with biotinylated secondary anti-IFN-γ mAb, followed by incubation with HRP-reagent and stained with TMB (Mabtech). The spots were then quantified with ImmunoSPOT S4 (Cellular Technology Ltd., Cleveland, OH, USA).
Western blotting
All samples were prepared and stained with antibodies, and the blots were visualized as previously described [40, 41]. The following were used as primary antibodies: STAT1 mAb (Cell Signaling Technology, Inc.) at 1:1000, phospho-STAT1 mAb (p-STAT1) (Cell Signaling Technology, Inc.) at 1:1000, PD-L1 mAb (Cell Signaling Technology, Inc.) at 1:1000, PD-L2 mAb (Cell Signaling Technology, Inc.) at 1:1000, STAT6 polyclonal Ab (Atras Antibodies, Bromma, Sweden) at 1:1000, p-STAT6 mAb (Thermo Fisher Scientific, Inc.) at 1:1000, β-actin mAb (Santa Cruz Biotechnology, Inc., Dallas, Texas, USA) at 1:2000. Horseradish peroxidase (HRP)-linked anti-rabbit antibody (Cell Signaling Technology, Inc.) at 1:2000 or the HRP-linked anti-mouse antibody (Santa Cruz Biotechnology, Inc.) at 1:2000 were used as secondary antibodies.
Flow cytometry
All samples were stained with antibodies, then measured and analyzed as previously described [40, 41]. The samples were stained with the following antibodies, which were purchased from Thermo Fisher Scientific, Inc.: PE-conjugated anti-human CD274 (B7-H1; PD-L1) at 1:20, and APC-conjugated anti-human CD273 (B7-DC; PD-L2) at 1:20. Isotype-matched immunoglobulin served as a negative control.
RT-PCR
Expression of KIF20A in GC cell lines was analyzed by reverse transcription PCR. Primers were used as described in Imai et al.: KIF20A, sense 5′-CTACAAGCACCCAAGGACTCT-3′ and antisense 5′-AGATGGAGAAGCGAATGTTT-3′ and ACTIN, sense 5′-CATCCACGAAACTACCTTCAACT-3′ and antisense 5′-TCTCCTTAGAGAGAAGTGGGGTG-3′ [42].
Pathway analysis
We used the GC cell lines including MKN7, NUGC-3, NUGC-4, and OCUM-1. Tumor cell lines were treated with or without IFN-γ, and the isolation of total RNA from treated tumor cell lines, as well as the analysis of microarray gene expression based on SuperPrint G3 Human Gene Expression 8 × 60 K v3 (Agilent, Inc., Santa Clara, CA), were performed at the Laboratory of Macrogen Japan Corp. (Tokyo, Japan). Upregulated genes by IFN-γ treatment were selected at a minimum fold change of 1.5 in each sample. The Database for Annotation, Visualization and Integrated Discovery v6.8 (https://david.ncifcrf.gov) was used for pathway (Kyoto Encyclopedia of Genes and Genomes PATHWAY) analysis on the list of selected genes [43].
Statistics
Comparisons of the IFN-γ or IL-4 gene signature with mRNA expression of PD-L1 or PD-L2, and of PD-L1 with PD-L2 mRNA expression, and of PD-1 with PD-L1 or PD-L2 mRNA expression were assessed using the scatter diagram and Pearson’s product-moment correlation coefficient. Fisher’s exact test, Chi-square test, and Mann–Whitney U test were used to determine differences between two variables, where appropriate. The significance of results in the cytotoxic assay was determined using one-way analysis of variance followed by a Tukey's post hoc test. Analyses were performed using SPSS Statistics Package version 25 (IBM, Chicago, IL, USA). All p values were two-sided, and those less than 0.05 were considered statistically significant.
Results
PD-L2 expression on TCs significantly correlated with tumor progression
We used IHC to evaluate the expression of PD-L1 and PD-L2 in primary tumors from 194 GC patients. The clinical features of the patients are summarized in Table 1, and representative immunostainings of PD-L1 and PD-L2 are presented in Fig. 1a. Regarding TCs, PD-L1 and PD-L2 were expressed in 64 (33.0%) and 55 (28.4%) patients, respectively, and both were co-expressed in 31 (16.0%) patients (Fig. 1b). On the other hand, regarding TIICs, PD-L1 and PD-L2 were, respectively, expressed in 133 (68.6%) and 155 (79.9%) of the patients, and both were co-expressed in 115 (59.3%) patients (Fig. 1b).
The relationship between PD-L1 or PD-L2 expression and clinicopathological factors was shown in Table 2. We confirmed that both PD-L1 and PD-L2 expression on TCs was strongly associated with venous invasion, lymphatic invasion, depth of invasion, and higher TNM stage.
mRNA expression of PD-L1 and PD-L2 is significantly correlated with the IFN-γ gene signature in GC
We previously reported that IFN-γ increased the gene expression level of PD-L2 in several cancer cell lines [35, 39]. Although there have been few articles providing the regulatory mechanism of PD-L2 expression on TCs, Rozali et al. and Loke et al. both reported that IL-4 increased PD-L2 expression on macrophages [14, 15]. Therefore, in the present study, we focused on the IFN-γ and IL-4 signaling pathways to elucidate the mechanism of PD-L2 expression on GC cells.
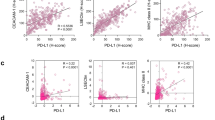
Analysis of TCGA stomach adenocarcinoma tissue dataset (n = 269) revealed that there were strong significant positive correlations between the IFN-γ gene signature and PD-L1 or PD-L2 (Fig. 2a), as well as weak but significant positive correlations between the IL-4 gene signature and PD-L1 or PD-L2 (Fig. 2b). These observations are in line with the results of our previous studies, which demonstrated the correlation between IFN-γ production and PD-L1 expression in the tumor microenvironment [35, 37, 39]. Furthermore, PD-L1 expression was found to be strongly associated with PD-L2 expression (Fig. 2c) and PD-1 expression was significantly correlated with both PD-L1 and PD-L2 expression (Fig. 2d).
Correlation of IFN-γ or IL-4 gene signature with the mRNA expression of PD-L1 or PD-L2 in TCGA stomach adenocarcinoma tissues dataset. a A heatmap showing IFN-γ gene signature and mRNA expressions of PD-L1 and PD-L2, and the correlation between IFN-γ gene signature and mRNA expressions of PD-L1 or PD-L2. b A heatmap showing IL-4 gene signature and mRNA expressions of PD-L1 and PD-L2, and the correlation between IL-4 gene signature and mRNA expressions of PD-L1 or PD-L2. c A heatmap showing mRNA expressions of PD-L1 and PD-L2, and the correlation between mRNA expressions of PD-L1 and PD-L2. d A heatmap showing mRNA expressions of PD-1, PD-L1, and PD-L2. The correlation between mRNA expressions of PD-1 and PD-L1 or PD-L2
PD-1/PD-L2 interaction inhibited the cytotoxicity of human CTL clone
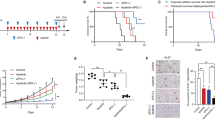
We next addressed how PD-1/PD-L2 interaction influenced the antitumor effects of T cells. At first, the proportion of apoptotic IL-2 activated T cells after co-culture with NUGC-3, which expresses both PD-L1 and PD-L2 (Fig. 3a), was analyzed using flow cytometry, as we recently reported that PD-1/PD-L1 interaction functionally led T cells expressing PD-1 to apoptosis [40]. To analyze the proportion of apoptotic T cells, CD3-positive cells were gated out of IL-2 activated lymphocytes, and the proportion of Annexin V positive cells was measured (Supplementary Fig. S1a). PD-1 expression was confirmed on IL-2 activated T cells (Supplementary Fig. S1b). Anti-PD-1 mAb, anti-PD-L1 mAb, and anti-PD-L2 mAb reduced the population of apoptotic T cells (Supplementary Fig. S1c), indicating that PD-1/PD-L2 as well as PD-1/PD-L1 interaction can affect the induction of T cell apoptosis, leading to an immunosuppressive role of PD-L2 in GC.
Effect of anti-PD-1 mAb, anti-PD-L1 mAb, and anti-PD-L2 mAb for anti-tumor specific CTL activity. a Surface expression of PD-L1 and PD-L2 was assessed by flow cytometry in MKN7 and NUGC-3. The black open curve is the specific ligand staining, and the gray filled curve represents the isotype control. b KIF20A expression was assessed by RT-PCR in GC cell lines. c The response of CTL clones treated with/without anti-PD-1 mAb against MKN7 or NUGC-3 treated with/without anti-PD-L1 mAb and/or anti-PD-L2 mAb was assessed by cytotoxic activity using calcein-release assay. Experiments were performed in triplicate, and error bars represent the mean ± SEM. of indicated samples. The p value by two-way ANOVA and Tukey’s test for multiple comparisons versus control; *p < 0.05; **p < 0.01; ***p < 0.001. E:T, effector: target
We subsequently performed ELISpot and cytotoxic assay using HLA-A24 restricted, KIF20A peptide-specific CTL clone in the presence of inhibitors for the PD-1 pathway, such as anti-PD-1 mAb, anti-PD-L1 mAb, and anti-PD-L2 mAb. The GC cell lines, MKN7 and NUGC-3, were used as targets because both cell lines are HLA-A24 positive [37] and express PD-L1, PD-L2, and KIF20A (Fig. 3a, b). Anti-PD-1 mAb, anti-PD-L1 mAb, and anti-PD-L2 mAb significantly enhanced the cytotoxicity of the CTL clone against MKN7 and NUGC3 (Fig. 3c). These inhibitors also enhanced the IFN-γ production of the CTL clone against MKN7 and NUGC3 (Supplementary Fig. S2). These results again revealed that PD-1/PD-L2 as well as PD-1/PD-L1 interactions can affect the anti-tumor activity of T cells, leading to an immunosuppressive role of PD-L2 in GC.
Upregulation of PD-L1 and PD-L2 by IFN-γ and IL-4 in GC cell lines
The optimal condition of IFN-γ treatment was based on our previous studies [37, 39] and that of IL-4 was determined by the same strategy (data not shown) [37, 38]. As a result, to analyze PD-L1 and PD-L2 expression, the GC cell lines were treated with either 10 ng/mL of IFN-γ or 10 ng/mL of IL-4 as described in the Methods section of the present study. Western blot analysis demonstrated that IFN-γ increased p-STAT1, which is a key molecule in the IFN-γ signaling pathway, and IL-4 increased p-STAT6, which is a key molecule in the IL-4 signaling pathway, in the GC cell lines (Fig. 4a). These observations validated that IFN-γ and IL-4 could efficiently act on these cell lines.
Effect of IFN-γ and IL-4 for the expression of PD-L1 and PD-L2 in GC cells. GC cell lines were treated without (control) or with 10 ng/mL IFN-γ or 10 ng/mL IL-4 for 1 h (a) or 48 h (b). Western blot analysis of relevant molecules in signaling pathways (a) and flow cytometric analysis of PD-L1 and PD-L2 expression (b) were performed, and representative results out of three independent experiments are shown. c Pathway analysis was performed in GC cell lines treated with IFN-γ as described in the Methods section
Treatment with IFN-γ increased PD-L1 expression in all the tested cell lines and PD-L2 expression in three out of 10 cell lines (Fig. 4b and Supplementary Fig. S3). On the other hand, treatment with IL-4 did not affect PD-L1 expression in any tested cell lines and increased PD-L2 expression in two out of 10 cell lines (Fig. 4b and Supplementary Fig. S3). To analyze the regulatory mechanism of PD-L2 expression, we focused on four cell lines, MKN7, NUGC-3, NUGC-4, and OCUM-1, because treatment with IFN-γ increased both PD-L1 and PD-L2 expression in MKN7 and NUGC-3, but only PD-L1 expression in NUGC-4 and OCUM-1 (Fig. 4b and Supplementary Fig. S3). The microarray gene expression analysis was performed for these cell lines after IFN-γ treatment. As a result, IFN-γ related genes in the JAK-STAT pathway were significantly increased in all tested cells and, of note, signaling pathways of TNF and NF-κB were enhanced in MKN7 and NUGC-3, but not in NUGC-4 and OCUM-1 (Fig. 4c). These results may indicate that PD-L2 expression is regulated by the TNF and the NF-κB signaling pathways, as well as the JAK-STAT pathway.
Discussion
It was revealed in an analysis of the ATTRACTION-2 study for GC that some patients with PD-L1-negative tumors have a favorable response to anti-PD-1 mAb in GC [3]. Since PD-1 interacts with PD-L2 as well as PD-L1, PD-L2 expression on TCs may be involved in the clinical response. However, the immunosuppressive role of PD-L2 in GC has not yet been fully elucidated. In the present study, we revealed that a subset of GC patients expressed PD-L2 relating to tumor progression, and both IFN-γ and IL-4 can up-regulate PD-L2 expression in GC cells. Furthermore, anti-PD-L2 mAb enhanced the anti-tumor activity of CTLs against GC cells expressing PD-L2. These results indicate that PD-1/PD-L2 interaction is significantly related to CTL function in GC and, to our knowledge, this is the first report to elucidate the immunosuppressive role of PD-L2 from the clinical and translational point of view in GC.
In our IHC cohort, 28.4% GC patients expressed PD-L2 on TCs (Fig. 1b) and PD-L2 expression on TCs was significantly associated with tumor progression (Table 2). TCGA stomach adenocarcinoma dataset also revealed that PD-L2 was expressed and was significantly co-expressed with PD-L1 in GC patients (Fig. 2c). Tanegashima et al. also reported that the PD-1/PD-L2 interaction appears to play a more important role in evading antitumor immunity than the PD-1/PD-L1 interaction in renal cell carcinoma and lung squamous cell carcinoma [44]. This report and our results in the present study indicated that a subset of GC patients expressed PD-L2 and PD-L2 as well as PD-L1 may suppress the immune response in the tumor microenvironment of GC.
PD-1 is expressed on CTL and binds to PD-L1 and PD-L2 on target cells [9]. Activation of the PD-1 pathway leads CTL-dysfunction, resulting in tumor progression [19, 20]. In the present study, we indicated that PD-L1 and PD-L2 were co-expressed on TCs of 16.0% GC patients in our IHC cohort (Fig. 1b), and, furthermore, PD-1 expression was significantly correlated with PD-L1 and PD-L2 expressions in TCGA stomach adenocarcinoma tissues dataset (Fig. 2d). These reports and our results indicated that GC cells may express both PD-L1 and PD-L2 in the situation that CTL exist in the tumor microenvironment. To increase the efficacy of immunotherapy for patients with GC, therefore, it is important to elucidate the expression mechanism and the immune suppressive role of PD-L2 in GC.
The mechanism to regulate the PD-L1 expression has been well demonstrated such as innate and acquired expression [45], and we recently reported that PD-L1 expression on GC cells significantly correlated with the presence of CD8 T cells in the tumor microenvironment and with the IFN-γ expression in TCs [39]. However, PD-L2 expression on human TCs has been demonstrated in limited studies, and the mechanism to regulate the PD-L2 expression in human TCs has not been fully elucidated [15, 16, 44]. Rozali et al. and Loke et al. both reported that IL-4 increased PD-L2 expression in macrophages [14, 15], and we and Garcia-Diaz et al. reported that IFN-γ upregulates the expression of PD-L2 in TCs [35, 39, 46]. Therefore, in the present study, we focused on IFN-γ and IL-4 as key regulators for the PD-L2 expression in GC.
Our results from TCGA stomach adenocarcinoma dataset (Fig. 2a, b) revealed that IFN-γ and, to a lesser extent, IL-4 signatures significantly correlated with PD-L2 expression. Moreover, in vitro assay using GC cells (Fig. 4b and Supplementary Fig. S3) showed that IFN-γ and, to a lesser extent, IL-4 can upregulate PD-L2 expression on GC cells. Therefore, it is likely that IFN-γ and, to a lesser extent, IL-4 are involved in the regulatory mechanisms for PD-L2 expression in GC cells, although other unknown mechanisms also exist. Garcia-Diaz et al. recently reported that IFN-β as well as IFN-γ upregulate PD-L2 in melanoma cells through STAT3, which binds to the PD-L2 promoter [46]. We investigated STAT3 expression in GC cells treated with IFN-γ or IL-4 using western blot, however, no correlation was observed between STAT3 and PD-L2 expression (data not shown). We speculate from the present study that the TNF and the NF-κB signaling pathways are involved in regulatory mechanisms for PD-L2 expression in GC cells, since these pathways were enhanced in MKN7 and NUGC-3 but not in NUGC-4 and OCUM-1 after IFN-γ stimulation (Fig. 4c), in which PD-L2 expression was increased by IFN-γ in the former two cell lines but not in the latter two. We need further investigation to fully elucidate the mechanism of PD-L2 expression in GC cells.
To our knowledge, this is the first report to directly show the functional consequence of the immunosuppressive role of PD-L2 in a human model, although two reports recently proved the antitumor activity of PD-L2 using a preclinical animal model [44, 47]. In the present study, using tumor antigen-specific CTL clone, we clearly showed that anti-PD-L2 mAb significantly enhanced the cytotoxicity of CTLs against GC cell lines expressing PD-L2 (Fig. 3c). Taken together, the blockade of PD-1/PD-L2 interaction, in addition to PD-1/PD-L1 interaction, may be an optimum for the immune checkpoint inhibitors in GC patients.
In conclusion, PD-L2 is expressed on GC cells and PD-1/PD-L2 as well as PD-1/PD-L1 interactions are functionally involved in anti-tumor CTL activities. Therefore, PD-L2 expression also should be considered when determining the optimal immunotherapy for patients with GC.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. https://doi.org/10.3322/caac.21492 (Epub 2018/09/13, PubMed PMID: 30207593).
Katai H, Ishikawa T, Akazawa K, Isobe Y, Miyashiro I, Oda I, et al. Five-year survival analysis of surgically resected gastric cancer cases in Japan: a retrospective analysis of more than 100,000 patients from the nationwide registry of the Japanese Gastric Cancer Association (2001–2007). Gastric Cancer. 2018;21(1):144–54. https://doi.org/10.1007/s10120-017-0716-7 (Epub 2017/04/19, PubMed PMID: 28417260).
Kang YK, Boku N, Satoh T, Ryu MH, Chao Y, Kato K, et al. Nivolumab in patients with advanced gastric or gastro-oesophageal junction cancer refractory to, or intolerant of, at least two previous chemotherapy regimens (ONO-4538–12, ATTRACTION-2): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet. 2017;390(10111):2461–71. https://doi.org/10.1016/s0140-6736(17)31827-5 (Epub 2017/10/11, PubMed PMID: 28993052).
Hodi FS, O'Day SJ, McDermott DF, Weber RW, Sosman JA, Haanen JB, et al. Improved survival with ipilimumab in patients with metastatic melanoma. N Engl J Med. 2010;363(8):711–23. https://doi.org/10.1056/NEJMoa1003466 (PubMed PMID: 20525992; PubMed Central PMCID: PMCPMC3549297).
Topalian SL, Hodi FS, Brahmer JR, Gettinger SN, Smith DC, McDermott DF, et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N Engl J Med. 2012;366(26):2443–544. https://doi.org/10.1056/NEJMoa1200690 (Epub 2012/06/05, PubMed PMID: 22658127; PubMed Central PMCID: PMCPMC3544539).
Brahmer JR, Tykodi SS, Chow LQ, Hwu WJ, Topalian SL, Hwu P, et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N Engl J Med. 2012;366(26):2455–65. https://doi.org/10.1056/NEJMoa1200694 (Epub 2012/06/05, PubMed PMID: 22658128; PubMed Central PMCID: PMCPMC3563263).
Motzer RJ, Escudier B, McDermott DF, George S, Hammers HJ, Srinivas S, et al. Nivolumab versus everolimus in advanced renal-cell carcinoma. N Engl J Med. 2015;373(19):1803–13. https://doi.org/10.1056/NEJMoa1510665 (Epub 2015/09/26, PubMed PMID: 26406148; PubMed Central PMCID: PMCPMC5719487).
Muro K, Chung HC, Shankaran V, Geva R, Catenacci D, Gupta S, et al. Pembrolizumab for patients with PD-L1-positive advanced gastric cancer (KEYNOTE-012): a multicentre, open-label, phase 1b trial. Lancet Oncol. 2016;17(6):717–26. https://doi.org/10.1016/s1470-2045(16)00175-3 (Epub 2016/05/10, PubMed PMID: 27157491).
Terme M, Ullrich E, Aymeric L, Meinhardt K, Desbois M, Delahaye N, et al. IL-18 induces PD-1-dependent immunosuppression in cancer. Can Res. 2011;71(16):5393–9. https://doi.org/10.1158/0008-5472.CAN-11-0993 (PubMed PMID: 21724589).
Fanoni D, Tavecchio S, Recalcati S, Balice Y, Venegoni L, Fiorani R, et al. New monoclonal antibodies against B-cell antigens: possible new strategies for diagnosis of primary cutaneous B-cell lymphomas. Immunol Lett. 2011;134(2):157–60. https://doi.org/10.1016/j.imlet.2010.09.022 (PubMed PMID: 20951741).
Zou W, Chen L. Inhibitory B7-family molecules in the tumour microenvironment. Nat Rev Immunol. 2008;8(6):467–77. https://doi.org/10.1038/nri2326 (PubMed PMID: 18500231).
Song M, Chen D, Lu B, Wang C, Zhang J, Huang L, et al. PTEN loss increases PD-L1 protein expression and affects the correlation between PD-L1 expression and clinical parameters in colorectal cancer. PLoS ONE. 2013;8(6):e65821. https://doi.org/10.1371/journal.pone.0065821 (PubMed PMID: 23785454; PubMed Central PMCID: PMCPMC3681867).
Herbst RS, Soria JC, Kowanetz M, Fine GD, Hamid O, Gordon MS, et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature. 2014;515(7528):563–7. https://doi.org/10.1038/nature14011 (Epub 2014/11/28, PubMed PMID: 25428504; PubMed Central PMCID: PMCPMC4836193).
Loke P, Allison JP. PD-L1 and PD-L2 are differentially regulated by Th1 and Th2 cells. Proc Natl Acad Sci USA. 2003;100(9):5336–411. https://doi.org/10.1073/pnas.0931259100 (Epub 2003/04/17, PubMed PMID: 12697896; PubMed Central PMCID: PMCPMC154346).
Rozali EN, Hato SV, Robinson BW, Lake RA, Lesterhuis WJ. Programmed death ligand 2 in cancer-induced immune suppression. Clin Dev Immunol. 2012;2012:656340. https://doi.org/10.1155/2012/656340 (Epub 2012/05/23, PubMed PMID: 22611421; PubMed Central PMCID: PMCPMC3350956).
Yearley JH, Gibson C, Yu N, Moon C, Murphy E, Juco J, et al. PD-L2 expression in human tumors: relevance to anti-PD-1 therapy in cancer. Clin Cancer Res. 2017;23(12):3158–67. https://doi.org/10.1158/1078-0432.Ccr-16-1761 (Epub 2017/06/18, PubMed PMID: 28619999).
Ohigashi Y, Sho M, Yamada Y, Tsurui Y, Hamada K, Ikeda N, et al. Clinical significance of programmed death-1 ligand-1 and programmed death-1 ligand-2 expression in human esophageal cancer. Clin Cancer Res. 2005;11(8):2947–53. https://doi.org/10.1158/1078-0432.Ccr-04-1469 (Epub 2005/04/20, PubMed PMID: 15837746).
Wu Y, Cao D, Qu L, Cao X, Jia Z, Zhao T, et al. PD-1 and PD-L1 co-expression predicts favorable prognosis in gastric cancer. Oncotarget. 2017;8(38):64066–82. https://doi.org/10.18632/oncotarget.19318 (Epub 2017/10/04, PubMed PMID: 28969052; PubMed Central PMCID: PMCPMC5609984).
Dong H, Strome SE, Salomao DR, Tamura H, Hirano F, Flies DB, et al. Tumor-associated B7–H1 promotes T-cell apoptosis: a potential mechanism of immune evasion. Nat Med. 2002;8(8):793–800. https://doi.org/10.1038/nm730 (PubMed PMID: 12091876).
Freeman GJ, Long AJ, Iwai Y, Bourque K, Chernova T, Nishimura H, et al. Engagement of the PD-1 immunoinhibitory receptor by a novel B7 family member leads to negative regulation of lymphocyte activation. J Exp Med. 2000;192(7):1027–34 (PubMed PMID: 11015443; PubMed Central PMCID: PMCPMC2193311).
Borghaei H, Paz-Ares L, Horn L, Spigel DR, Steins M, Ready NE, et al. Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N Engl J Med. 2015;373(17):1627–39. https://doi.org/10.1056/NEJMoa1507643 (Epub 2015/09/29, PubMed PMID: 26412456; PubMed Central PMCID: PMCPMC5705936).
Carbone DP, Reck M, Paz-Ares L, Creelan B, Horn L, Steins M, et al. First-line nivolumab in stage IV or recurrent non-small-cell lung cancer. N Engl J Med. 2017;376(25):2415–26. https://doi.org/10.1056/NEJMoa1613493 (PubMed PMID: 28636851).
Garon EB, Rizvi NA, Hui R, Leighl N, Balmanoukian AS, Eder JP, et al. Pembrolizumab for the treatment of non-small-cell lung cancer. N Engl J Med. 2015;372(21):2018–28. https://doi.org/10.1056/NEJMoa1501824 (Epub 2015/04/22, PubMed PMID: 25891174).
Robert C, Schachter J, Long GV, Arance A, Grob JJ, Mortier L, et al. Pembrolizumab versus ipilimumab in advanced melanoma. N Engl J Med. 2015;372(26):2521–32. https://doi.org/10.1056/NEJMoa1503093 (Epub 2015/04/22, PubMed PMID: 25891173).
Tumeh PC, Harview CL, Yearley JH, Shintaku IP, Taylor EJ, Robert L, et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature. 2014;515(7528):568–71. https://doi.org/10.1038/nature13954 (PubMed PMID: 25428505; PubMed Central PMCID: PMCPMC4246418).
Brahmer J, Reckamp KL, Baas P, Crino L, Eberhardt WE, Poddubskaya E, et al. Nivolumab versus docetaxel in advanced squamous-cell non-small-cell lung cancer. N Engl J Med. 2015;373(2):123–35. https://doi.org/10.1056/NEJMoa1504627 (Epub 2015/06/02, PubMed PMID: 26028407; PubMed Central PMCID: PMCPMC4681400).
Bellmunt J, de Wit R, Vaughn DJ, Fradet Y, Lee JL, Fong L, et al. Pembrolizumab as second-line therapy for advanced urothelial carcinoma. N Engl J Med. 2017;376(11):1015–26. https://doi.org/10.1056/NEJMoa1613683 (Epub 2017/02/18, PubMed PMID: 28212060; PubMed Central PMCID: PMCPMC5635424).
Herbst RS, Baas P, Kim DW, Felip E, Perez-Gracia JL, Han JY, et al. Pembrolizumab versus docetaxel for previously treated, PD-L1-positive, advanced non-small-cell lung cancer (KEYNOTE-010): a randomised controlled trial. Lancet. 2016;387(10027):1540–50. https://doi.org/10.1016/s0140-6736(15),01281-7 (Epub 2015/12/30, PubMed PMID: 26712084).
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6(269):l1. https://doi.org/10.1126/scisignal.2004088 (Epub 2013/04/04, PubMed PMID: 23550210; PubMed Central PMCID: PMCPMC4160307).
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2(5):401–4. https://doi.org/10.1158/2159-8290.Cd-12-0095 (Epub 2012/05/17, PubMed PMID: 22588877; PubMed Central PMCID: PMCPMC3956037).
Ayers M, Lunceford J, Nebozhyn M, Murphy E, Loboda A, Kaufman DR, et al. IFN-gamma-related mRNA profile predicts clinical response to PD-1 blockade. J Clin Investig. 2017;127(8):2930–40. https://doi.org/10.1172/jci91190 (Epub 2017/06/27, PubMed PMID: 28650338; PubMed Central PMCID: PMCPMC5531419).
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102(43):15545–50. https://doi.org/10.1073/pnas.0506580102 (Epub 2005/10/04, PubMed PMID: 16199517; PubMed Central PMCID: PMCPMC1239896).
Liberzon A, Subramanian A, Pinchback R, Thorvaldsdottir H, Tamayo P, Mesirov JP. Molecular signatures database (MSigDB) 3.0. Bioinformatics (Oxford, England). 2011;27(12):1739–40. https://doi.org/10.1093/bioinformatics/btr260 (Epub 2011/05/07, PubMed PMID: 21546393; PubMed Central PMCID: PMCPMC3106198).
Liberzon A, Birger C, Thorvaldsdottir H, Ghandi M, Mesirov JP, Tamayo P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell systems. 2015;1(6):417–25. https://doi.org/10.1016/j.cels.2015.12.004 (Epub 2016/01/16, PubMed PMID: 26771021; PubMed Central PMCID: PMCPMC4707969).
Nakayama Y, Mimura K, Tamaki T, Shiraishi K, Kua LF, Koh V, et al. PhosphoSTAT1 expression as a potential biomarker for antiPD1/antiPDL1 immunotherapy for breast cancer. Int J Oncol. 2019;54(6):2030–8. https://doi.org/10.3892/ijo.2019.4779 (Epub 2019/05/14, PubMed PMID: 31081058; PubMed Central PMCID: PMCPMC6521934).
Kikuchi T, Mimura K, Okayama H, Nakayama Y, Saito K, Yamada L, et al. A subset of patients with MSS/MSI-low-colorectal cancer showed increased CD8(+) TILs together with up-regulated IFN-gamma. Oncol Lett. 2019;18(6):5977–85. https://doi.org/10.3892/ol.2019.10953 (Epub 2019/12/04, PubMed PMID: 31788072; PubMed Central PMCID: PMCPMC6865144).
Mimura K, Kua LF, Shiraishi K, Kee Siang L, Shabbir A, Komachi M, et al. Inhibition of mitogen-activated protein kinase pathway can induce upregulation of human leukocyte antigen class I without PD-L1-upregulation in contrast to interferon-gamma treatment. Cancer Sci. 2014;105(10):1236–44. https://doi.org/10.1111/cas.12503 (PubMed PMID: 25154680).
Mimura K, Shiraishi K, Mueller A, Izawa S, Kua LF, So J, et al. The MAPK pathway is a predominant regulator of HLA-A expression in esophageal and gastric cancer. J Immunol (Baltimore, Md. 1950). 2013;191(12):6261–72. https://doi.org/10.4049/jimmunol.1301597 (PubMed PMID: 24244023; PubMed Central PMCID: PMCPMC3856928).
Mimura K, Teh JL, Okayama H, Shiraishi K, Kua LF, Koh V, et al. PD-L1 expression is mainly regulated by interferon gamma associated with JAK-STAT pathway in gastric cancer. Cancer Sci. 2018;109(1):43–53. https://doi.org/10.1111/cas.13424 (Epub 2017/10/17, PubMed PMID: 29034543; PubMed Central PMCID: PMCPMC5765310).
Thar Min AK, Okayama H, Saito M, Ashizawa M, Aoto K, Nakajima T, et al. Epithelial-mesenchymal transition-converted tumor cells can induce T-cell apoptosis through upregulation of programmed death ligand 1 expression in esophageal squamous cell carcinoma. Cancer Med. 2018. https://doi.org/10.1002/cam4.1564 (Epub 2018/06/02, PubMed PMID: 29855157; PubMed Central PMCID: PMCPMC6051241).
Ashizawa M, Okayama H, Ishigame T, Thar Min AK, Saito K, Ujiie D, et al. miRNA-148a-3p regulates immunosuppression in DNA mismatch repair-deficient colorectal cancer by targeting PD-L1. Mol Cancer Res. 2019;17(6):1403–13. https://doi.org/10.1158/1541-7786.Mcr-18-0831 (Epub 2019/03/16, PubMed PMID: 30872332).
Imai K, Hirata S, Irie A, Senju S, Ikuta Y, Yokomine K, et al. Identification of HLA-A2-restricted CTL epitopes of a novel tumour-associated antigen, KIF20A, overexpressed in pancreatic cancer. Br J Cancer. 2011;104(2):300–7. https://doi.org/10.1038/sj.bjc.6606052 (Epub 2010/12/24, PubMed PMID: 21179034; PubMed Central PMCID: PMCPMC3031900).
da Huang W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57. https://doi.org/10.1038/nprot.2008.211 (Epub 2009/01/10, PubMed PMID: 19131956).
Tanegashima T, Togashi Y, Azuma K, Kawahara A, Ideguchi K, Sugiyama D, et al. Immune suppression by PD-L2 against spontaneous and treatment-related antitumor immunity. Clin Cancer Res. 2019;25(15):4808–19. https://doi.org/10.1158/1078-0432.Ccr-18-3991 (Epub 2019/05/12, PubMed PMID: 31076547).
Pardoll DM. The blockade of immune checkpoints in cancer immunotherapy. Nat Rev Cancer. 2012;12(4):252–64. https://doi.org/10.1038/nrc3239 (PubMed PMID: 22437870).
Garcia-Diaz A, Shin DS, Moreno BH, Saco J, Escuin-Ordinas H, Rodriguez GA, et al. Interferon receptor signaling pathways regulating PD-L1 and PD-L2 expression. Cell Rep. 2017;19(6):1189–201. https://doi.org/10.1016/j.celrep.2017.04.031 (Epub 2017/05/13, PubMed PMID: 28494868).
Umezu D, Okada N, Sakoda Y, Adachi K, Ojima T, Yamaue H, et al. Inhibitory functions of PD-L1 and PD-L2 in the regulation of anti-tumor immunity in murine tumor microenvironment. Cancer Immunol Immunother. 2019;68(2):201–11. https://doi.org/10.1007/s00262-018-2263-4(Epub 2018/10/26, PubMed PMID: 30357491).
Funding
This work was supported by the Japan Society for the Promotion of Science KAKENHI Grant Number 16K10511 (KK).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human rights statement and informed consent
All procedures followed were in accordance with the Institutional Ethical Committee of Fukushima Medical University School of Medicine (Reference Nos. 2329 and 2847) and with the Helsinki Declaration of 1964 and later versions. Written informed consent was obtained from all participants.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nakayama, Y., Mimura, K., Kua, LF. et al. Immune suppression caused by PD-L2 expression on tumor cells in gastric cancer. Gastric Cancer 23, 961–973 (2020). https://doi.org/10.1007/s10120-020-01079-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10120-020-01079-z