Abstract
Background
Gastric cancer (GC) is one of the high-incidence and high-mortality cancers all over the world. Though genome-wide association studies (GWASs) have found some genetic loci related to GC, they could only explain a small fraction of the potential pathogenesis for GC.
Methods
We used multi-marker analysis of genomic annotation (MAGMA) to analyze pathways from four public pathway databases based on Chinese GWAS data including 2631 GC cases and 4373 controls. The differential expressions of selected genes in certain pathways were assessed on the basis of The Cancer Genome Atlas database. Immunohistochemistry was also conducted on 55 GC and paired normal tissues of Chinese patients to localize the expression of genes and further validate the differential expression.
Results
We identified three pathways including chemokine signaling pathway, potassium ion import pathway, and interleukin-7 (IL7) pathway, all of which were associated with GC risk. NMI in IL7 pathway and RAC1 in chemokine signaling pathway might be two new candidate genes involved in GC pathogenesis. Additionally, NMI and RAC1 were overexpressed in GC tissues than normal tissues.
Conclusion
Immune and inflammatory associated processes and potassium transporting might participate in the development of GC. Besides, NMI and RAC1 might represent two new key genes related to GC. Our findings might give new insight into the biological mechanism and immunotherapy for GC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Globally, gastric cancer (GC) is one of the most prevalent high-mortality cancers, and is reckoned to cause 951,000 new cases and 723,000 deaths each year [1]. In China, the disease burden of GC is extremely high, and it has been indicated that the number of new GC cases and deaths for 2015 are 679,100 and 498,000, respectively [2]. The most important known cause of GC is chronic inflammation induced by Helicobacter pylori (H. pylori) infection [3, 4], but malignant transformations were found to occur in only a small percentage of infected individuals. Therefore, apart from H. pylori infection, other factors such as genetic factors, dietary factors, Epstein-Barr virus infection, smoking and obesity may also contribute to incidence of GC [5, 6].
Germline variations of genomic sequence are implicated in the predisposition to most complex traits, including gastric cancer. Genetic variants in IL1B, encoding a cytokine with a pivotal role in GC development, were among the most studied variants based on candidate gene approach, and two variants (rs1143634 and rs16944) were associated with gastric cancer in an intermediate level of summary evidence [7]. Genome-wide association study (GWAS) is an effective method to simultaneously assess a large number of single nucleotide polymorphisms (SNPs) through high-throughput genetic analysis. Basing on this method, a series of genetic loci were identified to affect GC susceptibility, including rs4072037 (MUC1) [8], rs80142782 (ASH1L) [9], rs9841504 (ZBTB20) [10], rs13361707 (PRKAA1) [10], rs7712641 (lnc-POLR3G-4) [9], rs2294693 (UNC5CL) [11], rs1679709 (BTN3A2) [12], rs2294008 (PSCA) [13], rs2274223 (PLCE1) [8]. Nevertheless, the biological mechanisms behind the association between genetic variants and GC risk still remain unclear. Besides, a study in 2016 proposed that the estimate of heritability for GC was 22%, while those established risk loci could only explain a small proportion of GC heritability [14]. Therefore, there still exist large amounts of GC associated loci in the whole genome, and they have been missed most probably due to strict significance thresholds used in GWAS. Pathway-based GWAS analysis is a new and effective strategy that can detect the associations missed by traditional GWAS and explore the biological mechanisms of diseases. Recently, this new analysis strategy has given novel insights into pathogenesis of cancers such as oesophageal squamous cell carcinoma [15], pancreatic cancer [16] and breast cancer [17].
Multi-marker analysis of genomic annotation (MAGMA) is a fast and flexible tool for gene and gene-set analysis based on GWAS data [18]. MAGMA’s gene analysis uses a multiple regression method to validly incorporate linkage disequilibrium (LD) between variants and to discover multi-variant effects. Pathway-based analysis is based on gene-set analysis that is conducted using a gene-level regression model. MAGMA is a powerful tool to detect genes and pathways associated with diseases and to help us explore the potentially biological mechanisms [19, 20].
In this study, on the basis of Chinese GC GWAS, we applied MAGMA to conduct pathway analysis to identify crucial pathways and genes that contribute to GC susceptibility.
Materials and methods
Study populations
We conducted the analysis relying on three GC GWASs from ethnic Chinese. Two GWASs from Nanjing and Beijing populations (NJ-GWAS and BJ-GWAS) were implemented by our team [10], and the other GWAS from Shanxi and Linxian (NCI-GWAS) was based on Shanxi Upper Gastrointestinal Cancer Genetics Project supported by the National Cancer Institute (NCI) [8]. Details about the above three GWASs have been described elsewhere [8, 10]. In brief, all GC cases were histopathologically confirmed, and cancer-free controls from NJ-GWAS and BJ-GWAS were matched on age, sex and geographic region, while controls were matched on age and sex in NCI-GWAS. In total, 2631 GC cases and 4373 controls were included in our analysis. Basic demographic information of the participants was shown in Supplementary table 1.
Quality control, genotype imputation and meta-analysis
We excluded SNPs with call rate < 95%, minor allele frequency (MAF) < 0.01 or Hardy–Weinberg equilibrium (HWE) P value < 1 × 10−6. Then, we performed imputation with SHAPEIT [21] and IMPUTE2 [22] for those three GWASs separately. All populations from the 1000 Genomes Project Phase III were taken as the reference set. SNPTEST [23] was performed to evaluate the relationship between each variant and GC, then we used GWAMA [24] to conduct the meta-analysis. To get more reliable results, the SNPs were selected with more stringent inclusion criteria: HWE -value > 1 × 10−3 and imputed INFO score ≥ 0.5. Finally, 6,865,316 SNPs which were shared by all three GWASs and showed no obvious heterogeneity (I2 < 75%) were included in our following analysis.
Pathway-based analysis
Pathway data from four databases were downloaded online (http://software.broadinstitute.org/gsea/msigdb/collections.jsp). Totally, 186 gene sets form Kyoto Encyclopedia of Genes and Genomes (KEGG) database, 4436 gene sets from Gene Ontology (GO) database about biological process, 217 gene sets from BioCarta database and 674 gene sets from Reactome database were included in our analysis. We performed gene-based analysis on SNP P values, and raw genotype data from Nanjing and Beijing populations were set as a reference for linkage disequilibrium (LD). SNPs located in 10 kB upstream and downstream of a coding gene were mapped to the gene. The SNP-wise mean model which is equivalent to SKAT model using inverse variance weights [25] was adopted in this analysis. The maximum and minimum number of permutations per gene were set as 1 million and ten, respectively. We got two results about the P value for a certain gene, relying on different distributions (asymptotic sampling distribution and permutation-based sampling distribution). P value relying on asymptotic sampling distribution represented the result of gene-based analysis in this study. Taking advantage of the results of gene analysis, we implemented the gene-set analysis through a linear regression model. As Bonferroni correction may be too conservative when gene sets are strongly overlapping, we adopted a permutation-based empirical multiple testing correction which is provided by MAGMA. One hundred thousand permutations were performed during the gene-set analysis.
Differential expression analysis
Differential expression analysis was performed based on the data downloaded from The Cancer Genome Atlas (TCGA) database. Differential expression among paired (32 GC tissues and 32 adjacent normal tissues) and unpaired (413 GC tissues and 32 adjacent normal tissues) samples were both calculated in our analysis.
Immunohistochemistry (IHC)
Immunohistochemistry was performed on 55 pairs of GC and matched adjacent normal tissues of Chinese patients using tissue arrays (Shanghai Outdo Biotech Co., Ltd.). The tissue sections were sequential incubations with anti-N myc interactor antibody (for NMI, ab183724, Abcam) or anti-Rac1 antibody (for RAC1, ab33186, Abcam), and EnVision™ FLEX /HRP reagent (DM842, Dako Omnis). The intensity of staining and the frequency of the stained cells were estimated by two investigators who were blinded to the patients’ information. The frequency of positive cells was scored as follows: ≤5% = 0; >5 to ≤ 25% = 1; >25 to ≤ 50% = 2; >50 to ≤ 75% = 3; and > 75% = 4. Another score was given according to the intensity of staining as follows: negative = 0; weak = 1; moderate = 2; or strong = 3. For purposes of statistical analysis, NMI and RAC1 proteins’ intensity and frequency were transformed into a staining intensity score (SIS) calculated by multiplying the staining intensity score by the frequency score [26].
Statistical analysis
Multivariate logistic regression analysis in the additive model was used to estimate the association between genetic variants and GC risk. Nanjing and Beijing GWASs took age, sex, smoking status, alcohol consumption and the top genotype principal components (PCs) as covariates. NCI-GWAS was adjusted for age (10-year categories), sex and the top PCs. EIGENSTRAT 3.0 software was used to evaluate population structure. We adopted a fixed-effect model in the meta-analysis to explore the correlation between the single genetic variant and susceptibility of GC. As for differential expression analysis among mRNA level, t test was used to measure the difference between log-2 transformed expression values of GC and normal tissues. Moreover, differential expression analysis among protein level was conducted by Wilcoxon signed rank test on SIS.
Results
Based on 2631 GC cases and 4373 controls, we first obtained the gene-based results. Among 18,449 protein-coding genes on autosomes, we identified 1465 genes with P < 0.05 (Fig. 1). After Bonferroni correction, 20 genes reached statistical significance (P < 2.7 × 10−6, 0.05/18449) (Supplementary table 2), of which 19 genes were located in known GC susceptibility regions including 1q22, 5p13.1 and 10q23.33. IRGC in 19q13.32 was a new gene, but its expression level was similar between tumor and normal tissues (P = 0.15 and 0.52 for unpaired and paired samples from TCGA, respectively, Supplementary figure 1).
Manhattan plots that show the distribution and association of genes with GC risk. The x-axis is chromosomal position and the y-axis is − log10(P). the red horizontal line represents P = 2.7 × 10−6 while the green horizontal line represents P = 0.05. Red circles showed the association at P < 2.7 × 10−6, and the known GC-associated genes (MUC1 and PRKAA1) were marked by orange circles
Through pathway-based analysis, 244 candidate pathways were identified at the level of P < 0.05. Of the above pathways, 5 were from KEGG database, 200 from GO database, 11 from Biocarta database and 28 from Reactome database (Supplementary table 3–6). After permutation-based empirical multiple testing correction, no pathway remained. However, 3 pathways reached a less stringent threshold (corrected P < 0.5), and they were chemokine signaling pathway, potassium ion import pathway and interleukin-7 (IL7) pathway (Table 1). Stratification analysis of pathways showed no heterogeneities between the groups by age and sex (Supplementary table 7).
To discover new potentially causal genes, we examined the relationship between genes in each of the three pathways mentioned above and GC risk. Eighteen of 183 genes in chemokine signaling pathway were identified (P < 0.05), but only RAC1 and MAP2K1 were left after false discovery rate (FDR) correction (FDR corrected P < 0.05) (Table 2, Supplementary table 8). In potassium ion import pathway, although 4 of 28 genes were found to be associated with GC risk (P < 0.05), none of them reached the FDR corrected criterion (Supplementary table 9). Among 16 genes in IL7 pathway, 4 genes reached the P level at 0.05 and NMI passed FDR correction (Table 2, Supplementary table 10).
For the three genes passing FDR correction, we estimated the differential expression of them in paired and unpaired samples from TCGA. Results showed that NMI was overexpressed in GC for both of unpaired and paired samples, while increased expression of RAC1 in GC tissues was observed in unpaired tissues (Fig. 2). NMI and RAC1 were overexpressed in GC tissues in either early or advanced cases (Supplementary figure 2–3). However, the expression of MAP2K1 was similar between GC and normal tissues (Supplementary figure 1).
NMI localized to cytoplasm, whereas RAC1 localized primarily to cytoplasm and cellular matrix (Fig. 3). In GC tissues, NMI and RAC1 expressed in cancer cells but not in tumor-infiltrating lymphocytes. Among 55 Chinese GC patients, the protein levels of NMI and RAC1 were also increased in GC tissues as compared with paired normal tissues (P = 1.48 × 10−7 and 2.35 × 10−7, respectively), for either early stage (P = 2.04 × 10−3 and 2.30 × 10−3, respectively) or advanced stage of GC (P = 1.00 × 10−5 and 2.40 × 10−5, respectively) (Fig. 3).
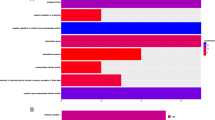
Results of Immunohistochemical staining of NMI and RAC1 proteins. a Immunohistochemical staining of NMI protein in two pairs of representative GC tissues and the adjacent normal tissues. b Immunohistochemical staining of RAC1 protein in two pairs of representative GC tissues and the adjacent normal tissues. c Distribution of staining intensity score of NMI protein in 55 paired GC tissues and the adjacent normal tissues. Red bars represent results of GC tissues, while blue bars represent results of the adjacent normal tissues. The first 25 paired samples came from early GC patients, and the last 30 paired samples came from advanced GC patients. d Distribution of staining intensity score of RAC1 protein in 55 paired GC tissues and the adjacent normal tissues. Red bars represent results of GC tissues, while blue bars represent results of the adjacent normal tissues. The first 25 paired samples came from early GC patients, and the last 30 paired samples came from advanced GC patients
Discussion
In this study, on the basis of 2631 GC cases and 4373 controls, we found 3 candidate pathways and 2 candidate casual genes associated with GC in Chinese populations. In a previous study, Lee et al. [27] performed an Identify Candidate Causal SNPs and Pathways (ICSNPathway) analysis based on NCI-GWAS, and reported several hypothetical biological mechanisms of GC, including ephrin receptor binding, drug and pyrimidine metabolism, cyanoamino acid metabolism, and lipid biosynthetic process, regulation of cell growth, and cation homeostasis. Using similar dataset and ICSNP pathway analysis, Zhu et al. [28] reported similar mechanisms including ephrin receptor binding. In contrast to the ICSNPathway aiming to discover candidate causal pathways that represent the way in which the candidate causal SNPs affect GC [29], MAGMA was used to identify novel pathways associated with GC in the current study. As a result, we found three new pathway for gastric cancer, including chemokine signaling pathway, potassium ion import pathway and Interleukin-7 (IL7) pathway. We also several identified pathways such as drug metabolism—other enzymes, pyrimidine metabolism, regulation of cell growth, and cellular cation homeostasis were repeated in our analysis, apart from three new discovered candidate pathways.
Three pathways were identified in our study. They were chemokine signaling, potassium ion import, and IL7 pathway. Chemokines are a large family of small cytokines and they were initially discovered as they could recruit immune cells to a site of inflammation during an immune response [30]. Subsequent researches reported chemokines could also provide directional guidance to normal cells like neurons and germ cells during embryonic development [31], besides, they could migrate cancer cells to distant sites during metastasis [32, 33]. Nowadays, researchers have paid great attention to the relationship between chemokines and cancer, and they found that chemokines were involved in many other cancer-related processes, including facilitating growth and survival of cancer cells [34, 35] and formation of cancer blood vessels [36]. It has been reported that chemokines persisting at an inflammatory site were crucial in neoplastic progression [37], and once GC occurred, certain chemokines could be produced by tumor and might play a great role in promoting the development and progression of GC. IL-7 is an important cytokine for adaptive immune system, because IL-7 plays an essential role in the development of B and T cells such as generation of T cells in the thymus and peripheral homeostasis of T Cells [38]. A healthy immune system could help clear tumor cells effectively, and suppressed immunity might increase the risk for cancers. Additionally, cancers could suppress normal immunological surveillance function to escape being identified by immune system during the progression of cancers [39]. According to the evidence above, we inferred that both chemokine signaling pathway and IL7 pathway might participate in occurrence of cancers through immune and inflammatory processes, and facilitate the development of cancers through multiple mechanisms.
Nowadays, immunotherapy has been a new treatment for cancers and has achieved great success [40,41,42,43]. For GC, several common immunotherapies such as cancer vaccines therapy, T cell based adoptive transfer therapy and checkpoint inhibition therapy have been used to induce antitumor immunity to kill cancer cells and improve survival [44]. Although chemokines and cytokines have been commonly applied to the treatment of cancers, they were rarely used for treating GC [45,46,47]. Recent studies have found IL7 has great potential in cancer immunotherapy because of its function in immune reconstitution, enhancing the function of effector immune cells and fighting against the immunosuppressive network [48]. Therefore, in consideration of those findings of the recent studies mentioned above and our results, there are reasons to believe that new ways of immunotherapy for GC might be discovered centering on chemokines, IL7 and other cytokines in the future.
In addition to the above two pathways, potassium ion import pathway was the third one we discovered. Potassium has been reported to be associated with apoptosis, for example, loss of potassium ions is linked to shrinkage which is a basic morphological characteristic of apoptosis [49]. Besides, low level of cytoplasmic potassium is capable of activating caspases and nucleases, which is pivotal in apoptosis [50, 51]. Abnormal apoptosis process could break the equilibrium between cell proliferation and cell death, which might promote oncogenesis because “undead” cells are accumulated [52]. After oncogenesis, accumulation of potassium in T cells would suppress the activity of T cells [53]. Enhancing activity of potassium channel may enable T cells to attack cancer cells more effectively and powerfully [54]. Besides, potassium channels were reported to be correlated with multidrug resistance in gastric cancer cells [55]. Thus, process of potassium import may influence susceptibility of GC and provide new ideas for treating GC.
NMI and RAC1 were two new candidate genes discovered in our study. NMI in 2q23.3 could interact with N-myc and C-myc which are two members of the oncogene Myc family and induce transcription activity as a transcription cofactor [56]. Researchers have found that overexpression of NMI retards invasion and growth of cancer cells through inhibiting the Wnt/beta-catenin signaling [57]. Besides, NMI could suppress tumor invasion and metastasis by inhibiting NF-κB pathways in human gastric cancer cells [58]. NMI in IL7 pathway can augment the recruitment of CBP/p300 to transcription factor STAT5, which can enhance transcription of downstream target genes [59]. STAT5 proteins are required for normal T cell proliferation and NK function. Additionally, they critically regulate vital cellular functions such as proliferation, differentiation, and survival [60]. RAC1 located at 7p22.1 is a GTPase that belongs to the RAS superfamily whose members can regulate cellular events, such as cell growth, cytoskeletal reorganization, and the activation of protein kinases [61]. RAC1 in chemokine signaling pathway can interact with downstream effector PAK1. Activation of PAK1 affects mitogen activated protein kinase (MAPK), phospoinositide 3-kinase (PI3K), and Wnt/beta-catenin signaling pathways associated with inflammation and malignant transformation [62]. RAC1 is also a key activator of NF-κB and promote tumor growth by inducing the expression of inflammatory cytokines [63]. It has been proved that RAC1 was overexpressed in cancer tissues and associated with poor survival [64, 65]. In addition, researchers have indicated RAC1 can serve as therapeutic targets for cancers including GC [66, 67]. Therefore, both of the two candidate genes discovered in our study are vital in the progression of cancers and might be treated as drug targets for GC.
In conclusion, immune and inflammatory associated processes and potassium transporting might participate in the occurrence and development of GC. Besides, NMI and RAC1 were two new vital genes related to GC. Our findings might give new insight into the mechanism and treatment of GC. However, due to the genetic heterogeneity among different populations [7, 68], there might be inconsistent results according to the ethnicities as a result of differences in genetic basis as well as environmental exposures. In the future, additional studies are needed to confirm the identified pathways and genes associated with risk for GC in populations of East Asian and elsewhere, and functional experiments should be conducted to observe the effect of the two candidate genes on the development of GC. Additionally, more efforts are warranted to explore new therapies for GC based on our findings, and potential therapies may refer to immunotherapy centering on chemokines, IL7 and other cytokines, to enhance activity of potassium channel, and therapeutic targets on NMI and RAC1.
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65(2):87–108.
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66(2):115–32.
Bornschein J, Selgrad M, Warnecke M, Kuester D, Wex T, Malfertheiner P. H. pylori infection is a key risk factor for proximal gastric cancer. Digestive Dis Sci. 2010;55(11):3124–31.
Blankfield RP. Helicobacter pylori infection and the development of gastric cancer. N EnglJ Med. 2002;346(1):65–7.
Cheng XJ, Lin JC, Tu SP. Etiology and prevention of gastric cancer. Gastrointest Tumors. 2016;3(1):25–36.
Van Cutsem E, Sagaert X, Topal B, Haustermans K, Prenen H. Gastric cancer. Lancet. 2016;388(10060):2654–64.
Mocellin S, Verdi D, Pooley KA, Nitti D. Genetic variation and gastric cancer risk: a field synopsis and meta-analysis. Gut. 2015;64(8):1209–19.
Abnet CC, Freedman ND, Hu N, Wang Z, Yu K, Shu XO, et al. A shared susceptibility locus in PLCE1 at 10q23 for gastric adenocarcinoma and esophageal squamous cell carcinoma. Nat Genet. 2010;42(9):764–7.
Wang Z, Dai J, Hu N, Miao X, Abnet CC, Yang M, et al. Identification of new susceptibility loci for gastric non-cardia adenocarcinoma: pooled results from two Chinese genome-wide association studies. Gut. 2017;66(4):581–7.
Shi Y, Hu Z, Wu C, Dai J, Li H, Dong J, et al. A genome-wide association study identifies new susceptibility loci for non-cardia gastric cancer at 3q13.31 and 5p13.1. Nat Genet. 2011;43(12):1215–8.
Hu N, Wang Z, Song X, Wei L, Kim BS, Freedman ND, et al. Genome-wide association study of gastric adenocarcinoma in Asia: a comparison of associations between cardia and non-cardia tumours. Gut. 2016;65(10):1611–8.
Zhu M, Yan C, Ren C, Huang X, Zhu X, Gu H, et al. Exome array analysis identifies variants in SPOCD1 and BTN3A2 that affect risk for gastric cancer. Gastroenterology. 2017;152(8):2011–21.
Sakamoto H, Yoshimura K, Saeki N, Katai H, Shimoda T, Matsuno Y, et al. Genetic variation in PSCA is associated with susceptibility to diffuse-type gastric cancer. Nat Genet. 2008;40(6):730–40.
Mucci LA, Hjelmborg JB, Harris JR, Czene K, Havelick DJ, Scheike T, et al. Familial risk and heritability of cancer among twins in Nordic countries. Jama. 2016;315(1):68–76.
Hyland PL, Zhang H, Yang Q, Yang HH, Hu N, Lin SW, et al. Pathway, in silico and tissue-specific expression quantitative analyses of oesophageal squamous cell carcinoma genome-wide association studies data. Int J Epidemiol. 2016;45(1):206–20.
Li D, Duell EJ, Yu K, Risch HA, Olson SH, Kooperberg C, et al. Pathway analysis of genome-wide association study data highlights pancreatic development genes as susceptibility factors for pancreatic cancer. Carcinogenesis. 2012;33(7):1384–90.
Menashe I, Maeder D, Garcia-Closas M, Figueroa JD, Bhattacharjee S, Rotunno M, et al. Pathway analysis of breast cancer genome-wide association study highlights three pathways and one canonical signaling cascade. Cancer Res. 2010;70(11):4453–9.
de Leeuw CA, Mooij JM, Heskes T, Posthuma D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput Biol. 2015;11(4):e1004219.
Sniekers S, Stringer S, Watanabe K, Jansen PR, Coleman JRI, Krapohl E, et al. Genome-wide association meta-analysis of 78,308 individuals identifies new loci and genes influencing human intelligence. Nat Genet. 2017;49(7):1107–12.
Jansen A, Dieleman GC, Smit AB, Verhage M, Verhulst FC, Polderman TJC, et al. Gene-set analysis shows association between FMRP targets and autism spectrum disorder. Eur J Hum Genet EJHG. 2017;25(7):863–8.
Delaneau O, Marchini J, Zagury JF. A linear complexity phasing method for thousands of genomes. Nat Methods. 2011;9(2):179–81.
Howie BN, Donnelly P, Marchini J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 2009;5(6):e1000529.
Marchini J, Howie B, Myers S, McVean G, Donnelly P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat Genet. 2007;39(7):906–13.
Magi R, Morris AP. GWAMA: software for genome-wide association meta-analysis. BMC Bioinform. 2010;11:288.
Timbers TA, Garland SJ, Mohan S, Flibotte S, Edgley M, Muncaster Q, et al. Accelerating gene discovery by phenotyping whole-genome sequenced multi-mutation strains and using the sequence kernel association test (SKAT). PLoS Genet. 2016;12(8):e1006235.
Chen Y, Tian T, Mao MJ, Deng WY, Li H. CRBP-1 over-expression is associated with poor prognosis in tongue squamous cell carcinoma. BMC Cancer. 2018;18(1):514.
Lee JH, Kim Y, Choi JW, Kim YS. Genetic variants and risk of gastric cancer: a pathway analysis of a genome-wide association study. SpringerPlus. 2015;4:215.
Zhu H, Yang M, Zhang H, Chen X, Yang X, Zhang C, et al. Genome-wide association pathway analysis to identify candidate single nucleotide polymorphisms and molecular pathways for gastric adenocarcinoma. Tumour Biol J Int Soc Oncodev Biol Med. 2015;36(7):5635–9.
Zhang K, Chang S, Cui S, Guo L, Zhang L, Wang J. ICSNPathway: identify candidate causal SNPs and pathways from genome-wide association study by one analytical framework. Nucleic acids research. 2011;39(Web Server issue):W437–43.
Palomino DC, Marti LC. Chemokines and immunity. Einstein (Sao Paulo). 2015;13(3):469–73.
Lewellis SW, Knaut H. Attractive guidance: how the chemokine SDF1/CXCL12 guides different cells to different locations. Semin Cell Dev Biol. 2012;23(3):333–40.
Roy I, McAllister DM, Gorse E, Dixon K, Piper CT, Zimmerman NP, et al. Pancreatic cancer cell migration and metastasis is regulated by chemokine-biased agonism and bioenergetic signaling. Cancer Res. 2015;75(17):3529–42.
Singh R, Lillard JW Jr, Singh S. Chemokines: key players in cancer progression and metastasis. Front Biosci (Schol Ed). 2011;3:1569–82.
Wani N, Nasser MW, Ahirwar DK, Zhao H, Miao Z, Shilo K, et al. C-X-C motif chemokine 12/C-X-C chemokine receptor type 7 signaling regulates breast cancer growth and metastasis by modulating the tumor microenvironment. Breast Cancer Res BCR. 2014;16(3):R54.
Bolitho C, Hahn MA, Baxter RC, Marsh DJ. The chemokine CXCL1 induces proliferation in epithelial ovarian cancer cells by transactivation of the epidermal growth factor receptor. Endocrine-Related Cancer. 2010;17(4):929–40.
Kiefer F, Siekmann AF. The role of chemokines and their receptors in angiogenesis. Cell Mol Life Sci CMLS. 2011;68(17):2811–30.
Coussens LM, Werb Z. Inflammation and cancer. Nature. 2002;420(6917):860–7.
Surh CD, Sprent J. Homeostasis of naive and memory T cells. Immunity. 2008;29(6):848–62.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–74.
Rosenberg SA, Restifo NP. Adoptive cell transfer as personalized immunotherapy for human cancer. Science. 2015;348(6230):62–8.
Amedei A, Prisco D, MM DE. The use of cytokines and chemokines in the cancer immunotherapy. Recent Patents Anti-Cancer Drug Discov. 2013;8(2):126–42.
Ruella M, Kalos M. Adoptive immunotherapy for cancer. Immunol Rev. 2014;257(1):14–38.
Ye Z, Li Z, Jin H, Qian Q. Therapeutic cancer vaccines. Adv Exp Med Biol. 2016;909:139–67.
Subhash VV, Yeo MS, Tan WL, Yong WP. Strategies and advancements in harnessing the immune system for gastric cancer immunotherapy. J Immunol Res. 2015;2015:308574.
Valentine FT, Golomb FM, Harris M, Roses DF. A novel immunization strategy using cytokine/chemokines induces new effective systemic immune responses, and frequent complete regressions of human metastatic melanoma. Oncoimmunology. 2018;7(2):e1386827.
Alfaro C, Sanmamed MF, Rodriguez-Ruiz ME, Teijeira A, Onate C, Gonzalez A, et al. Interleukin-8 in cancer pathogenesis, treatment and follow-up. Cancer Treat Rev. 2017;60:24–31.
Fridlender ZG, Buchlis G, Kapoor V, Cheng G, Sun J, Singhal S, et al. CCL2 blockade augments cancer immunotherapy. Cancer Res. 2010;70(1):109–18.
Gao J, Zhao L, Wan YY, Zhu B. Mechanism of action of IL-7 and its potential applications and limitations in cancer immunotherapy. Int J Mol Sci. 2015;16(5):10267–80.
Bortner CD, Cidlowski JA. Cell shrinkage and monovalent cation fluxes: role in apoptosis. Arch Biochem Biophys. 2007;462(2):176–88.
Cain K, Langlais C, Sun XM, Brown DG, Cohen GM. Physiological concentrations of K + inhibit cytochrome c-dependent formation of the apoptosome. J Biol Chem. 2001;276(45):41985–90.
Hughes FM Jr, Bortner CD, Purdy GD, Cidlowski JA. Intracellular K + suppresses the activation of apoptosis in lymphocytes. J Biol Chem. 1997;272(48):30567–76.
Cotter TG. Apoptosis and cancer: the genesis of a research field. Nat Rev Cancer. 2009;9(7):501–7.
Eil R, Vodnala SK, Clever D, Klebanoff CA, Sukumar M, Pan JH, et al. Ionic immune suppression within the tumour microenvironment limits T cell effector function. Nature. 2016;537(7621):539–43.
Chandy KG, Norton RS. Immunology channelling potassium to fight cancer. Nature. 2016;537(7621):497–9.
Han Y, Shi Y, Han Z, Sun L, Fan D. Detection of potassium currents and regulation of multidrug resistance by potassium channels in human gastric cancer cells. Cell Biol Int. 2007;31(7):741–7.
Bao J, Zervos AS. Isolation and characterization of Nmi, a novel partner of Myc proteins. Oncogene. 1996;12(10):2171–6.
Fillmore RA, Mitra A, Xi Y, Ju J, Scammell J, Shevde LA, et al. Nmi (N-Myc interactor) inhibits Wnt/beta-catenin signaling and retards tumor growth. Int J Cancer. 2009;125(3):556–64.
Hou J, Wang T, Xie Q, Deng W, Yang JY, Zhang SQ, et al. N-Myc-interacting protein (NMI) negatively regulates epithelial-mesenchymal transition by inhibiting the acetylation of NF-kappaB/p65. Cancer Lett. 2016;376(1):22–33.
Zhu M, John S, Berg M, Leonard WJ. Functional association of Nmi with Stat5 and Stat1 in IL-2- and IFNgamma-mediated signaling. Cell. 1999;96(1):121–30.
Rani A, Murphy JJ. STAT5 in cancer and immunity. J Interferon Cytokine Res. 2016;36(4):226–37.
Eller-Borges R, Batista WL, da Costa PE, Tokikawa R, Curcio MF, Strumillo ST, et al. Ras, Rac1, and phosphatidylinositol-3-kinase (PI3K) signaling in nitric oxide induced endothelial cell migration. Nitric Oxide Biol Chem. 2015;47:40–51.
Dammann K, Khare V, Gasche C. Tracing PAKs from GI inflammation to cancer. Gut. 2014;63(7):1173–84.
Shi Y, Bollam SR, White SM, Laughlin SZ, Graham GT, Wadhwa M, et al. Rac1-mediated DNA damage and inflammation promote Nf2 tumorigenesis but also limit cell-cycle progression. Dev Cell. 2016;39(4):452–65.
Leng R, Liao G, Wang H, Kuang J, Tang L. Rac1 expression in epithelial ovarian cancer: effect on cell EMT and clinical outcome. Med Oncol. 2015;32(2):329.
Zhou Y, Liao Q, Han Y, Chen J, Liu Z, Ling H, et al. Rac1 overexpression is correlated with epithelial mesenchymal transition and predicts poor prognosis in non-small cell lung cancer. J Cancer. 2016;7(14):2100–9.
Ji J, Feng X, Shi M, Cai Q, Yu Y, Zhu Z, et al. Rac1 is correlated with aggressiveness and a potential therapeutic target for gastric cancer. Int J Oncol. 2015;46(3):1343–53.
Bid HK, Roberts RD, Manchanda PK, Houghton PJ. RAC1: an emerging therapeutic option for targeting cancer angiogenesis and metastasis. Mol Cancer Ther. 2013;12(10):1925–34.
Kim J, Kim Y, Lee KA. Ethnic differences in gastric cancer genetic susceptibility: allele flips of interleukin gene. World J Gastroenterol. 2014;20(16):4558–65.
Acknowledgements
The authors thank all the participants of the Nanjing/Beijing and the National Cancer Institute gastric cancer studies. This study was supported by grants from the National key research and development program of China (Grant no. 2016YFC1302703); National Natural Science Foundation of China (81521004, 81422042, 81373090, 81703297); Jiangsu Province’s Key Medical Talents Program (QNRC2016352); the key grant of natural science foundation of Jiangsu higher education institutions (15KJA330002); the Priority Academic Program Development of Jiangsu Higher Education Institutions (Public Health and Preventive Medicine) and Top-notch Academic Programs Project of Jiangsu Higher Education Institutions (PPZY2015A067).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Human rights statement
All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1964 and later versions.
Informed consent
Informed consent to be included in the study, or the equivalent, was obtained from all patients.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yu, F., Tian, T., Deng, B. et al. Multi-marker analysis of genomic annotation on gastric cancer GWAS data from Chinese populations. Gastric Cancer 22, 60–68 (2019). https://doi.org/10.1007/s10120-018-0841-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10120-018-0841-y