Abstract
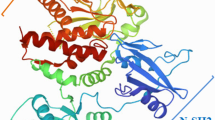
The Polo-like kinases (Plks) are a conserved subfamily of serine-threonine protein kinases that have significant roles in cell proliferation. The serine/threonine protein kinases or polo-like kinase 1 (PLK1) exist in centrosome during interphase and is an important regulatory enzyme in cell cycle progression during M phase. Mutations in mammalian PLK1 were found to be over expressed in various human cancers and it is disrupting the binding ability of polo box domain with target peptide. In this analysis we implemented a computational approach to filter the most deleterious and cancer associated mutation on PLK1 protein. We found W414F as the most deleterious and cancer associated by Polyphen 2.0, SIFT, I-mutant 3.0, PANTHER, PhD-SNP, SNP&GO, Mutpred and Dr Cancer tools. Molecular docking and molecular dynamics simulation (MDS) approach was used to investigate the structural and functional behavior of PLK1 protein upon mutation. MDS and docking results showed stability loss in mutant PLK1 protein. Due to mutation, PLK1 protein became more flexible and alters the dynamic property of protein which might affect the interaction with target peptide and leads to cell proliferation. Our study provided a well designed computational methodology to examine the cancer associated nsSNPs and their molecular mechanism. It further helps scientists to develop a drug therapy against PLK1 cancer-associated diseases.

Flow chart of in-silico screening of cancer associated mutation on PLK1 protein and its structural consequences studies.








Similar content being viewed by others
References
Barr FA, Sillje HH, Nigg EA (2004) Polo-like kinases and the orchestration of cell division. Nat Rev Mol Cell Biol 5:429–440
Van de Weerdt BC, Medema RH (2006) Polo-like kinases: a team in control of the division. Cell Cycle 5:853–864
Eckerdt F, Yuan J, Strebhardt K (2005) Polo-like kinases and oncogenesis. Oncogene 24:267–276
Strebhardt K, Ullrich A (2006) Targeting polo-like kinase 1 for cancer therapy. Nat Rev Cancer 6:321–330
Lee KS, Grenfell TZ, Yarm FR, Erikson RL (1998) Mutation of the polo-box disrupts localization and mitotic functions of the mammalian polo kinase plk. Proc Natl Acad Sci U S A 95:9301–9306
Nigg EA (1998) Polo-like kinases: positive regulators of cell division from start to finish. Curr Opin Cell Biol 10:776–783
Glover DM, Ohkura H, Tavares A (1996) Polo kinase: the choreographer of mitotic stage. J Cell Biol 135:1681–1684
Lane H, Nigg EA (1997) Cell-cycle control: POLO-like kinases join the outer circle. Trends Cell Biol 7:63–68
Llamazares S, Moreira A, Tavares A et al (1991) Polo encodes a protein kinase homolog required for mitosis in Drosophila. Genes Dev 5:2153–2165
Ohkura H, Hagan IM, Glover DM (1995) The conserved chizosaccharomyces pombe kinase plo1, required to form a bipolar spindle, the actin ring, and septum, can drive septum formation in G1 and G2 cells. Genes Dev 9:1059–1073
Lane HA, Nigg EA (1996) Antibody microinjection reveals an essential role for human polo-like kinase 1 (Plk1) in the functional maturation of mitotic centrosomes. J Cell Biol 135:1701–1713
Kumagai A, Dunphy WG (1996) Purification and molecular cloning of Plx1, a Cdc25-regulatory kinase from Xenopus egg extracts. Science 273:1377–1380
Toczyski DP, Galgoczy DJ, Hartwell LH (1997) CDC5 and CKII control adaptation to the yeast DNA damage checkpoint. Cell 90:1097–1106
Shirayama M, Zachariae W, Ciosk R (1998) The Polo-like kinase Cdc5p and the WD-repeat protein Cdc20p/fizzy are regulators and substrates of the anaphase promoting complex in Saccharomyces cerevisiae. EMBO J 17:1336–1349
Descombes P, Nigg EA (1998) The polo-like kinase Plx1 is required for M phase exit and destruction of mitotic regulators inXenopus egg extracts. EMBO J 17:1328–1335
Lee KS, Erikson RL (1997) Plk is a functional homolog of Saccharomyces cerevisiae Cdc5, and elevated Plk. Mol Cell Biol 17(6):3408-3417.
Seong YS, Kamijo K, Lee JS et al (2002) A spindle checkpoint arrest and a cytokinesis failure by the dominant-negative polo-box domain of Plk1 in U-2 OS cells. J Biol Chem 277:32282–32293
Song S, Grenfell TZ, Garfield S et al (2000) Essential function of the polo box of Cdc5 in subcellular localization and induction of cytokinetic structures. Mol Cell Biol 20:286–298
Liu J, Lewellyn AL, Chen LG, Maller JL (2004) The polo box is required for multiple functions of Plx1 in mitosis. J Biol Chem 279:21367–21373
Jeong K, Jeong JY, Lee HO (2010) Inhibition of Plk1 induces mitotic infidelity and embryonic growth defects in developing zebrafish embryos. Dev Biol 345:34–48
García-Alvarez B, de Cárcer G, Ibañez S et al (2007) Molecular and structural basis of polo-like kinase 1 substrate recognition: implications in centrosomal localization. Proc Natl Acad Sci U S A 104:3107–3112
Yun SM, Moulaei T, Lim D et al (2009) Structural and functional analyses of minimal phosphopeptides targeting the polo-box domain of polo-like kinase 1. Nat Struct Mol Biol 16:876–882
Strebhardt K (2001) In: Creighton TE (ed) PLK (Polo-like kinase): encyclopedia of molecular medicine. Wiley, New York, pp 2530–2532
Balu K, Purohit R (2013) Mutational analysis of TYR gene and its structural consequences in OCA1A. Gene 513:184–195
Kamaraj B, Purohit R (2013) In silico screening and molecular dynamics simulation of disease-associated nsSNP in TYRP1 gene and its structural consequences in OCA3. BioMed Res Int. doi:10.1155/2013/697051
Kamaraj B, Purohit R (2013) Computational screening of disease-associated mutations in oca2 gene. Cell Biochem Biophys. doi:10.1007/s12013-013-9697-2
Kumar A, Rajendran V, Sethumadhavan R et al (2013) Evidence of colorectal cancer-associated mutation in MCAK: a computational report. Cell Biochem Biophys. doi:10.1007/s12013-013-9572-1
Kumar A, Rajendran V, Sethumadhavan R et al (2013) Roadmap to determine the point mutations involved in cardiomyopathy disorder: a Bayesian approach. Gene 519:34–40
Adzhubei IA, Schmidt S, Peshkin L et al (2010) A method and server for predicting damaging missense mutations. Nat Methods 7:248–249
Kumar P, Henikoff S, Ng PC (2009) Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc 4:1073–1081
Capriotti E, Fariselli P, Rossi I, Casadio R (2008) A three-state prediction of single point mutations on protein stability changes. BMC Bioinforma 9S6
Thomas PD, Campbell MJ, Kejariwal A et al (2003) PANTHER: a library of protein families and subfamilies indexed by function. Genome Res 13:2129–2141
Capriotti E, Calabrese R, Casadio R (2006) Predicting the insurgence of human genetic diseases associated to single point protein mutations with support vector machines and evolutionary information. Bioinformatics 22:2729–2734
Calabrese R, Capriotti E, Fariselli P et al (2009) Functional annotations improve the predictive score of human disease-related mutations in proteins. Hum Mutat 30:1237–1244
Li B, Krishnan VG, Mort ME et al (2009) Automated inference of molecular mechanisms of disease from amino acid substitutions. Bioinformatics 25:2744–2750
Capriotti E, Altman RB (2011) A new disease-specific machine learning approach for the prediction of cancer-causing missense variants. Genomics 98:310–317
Purohit R, Rajendran V, Sethumadhavan R (2011) Relationship between mutation of serine residue at 315th position in M. tuberculosis catalase-peroxidase enzyme and isoniazid susceptibility: an in silico analysis. J Mol Model 17:869–877
Purohit R, Rajendran V, Sethumadhavan R (2011) Studies on adaptability of binding residues and flap region of TMC-114 resistance HIV-1 protease mutants. J Biomol Struct Dyn 29:137–152
Rajendran V, Sethumadhavan R (2013) Drug resistance mechanism of PncA in Mycobacterium tuberculosis. J Biomol Struct Dyn. doi:10.1080/07391102.2012.759885
Rajendran V, Purohit R, Sethumadhavan R (2012) In silico investigation of molecular mechanism of laminopathy cause by a pointmutation (R482W) in lamin A/C protein. Amino Acids 43:603–615
Purohit R (2013) Role of ELA region in auto-activation of mutant KIT receptor; a molecular dynamics simulation insight. J Biomol Struct Dyn. doi:10.1080/07391102.2013.803264
Balu K, Rajendran V, Sethumadhavan R et al (2013) Investigation of binding phenomenon of NSP3 and p130Cas mutants and their effect on cell signalling. Cell Biochem Biophys: 1–11
Yip YL, Scheib H, Diemand AV et al (2004) The Swiss-Prot variant page and the ModSNP database: a resource for sequence and structure information on human protein variants. Hum Mutat 23:464–470
Yip YL, Famiglietti M, Gos A et al (2008) Annotating single amino acid polymorphisms in the UniProt/Swiss-Prot knowledgebase. Hum Mutat 29:361–366
Boeckmann B, Bairoch A, Apweiler R et al (2003) The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Res 31:365–370
Berman HM, Westbrook J, Feng Z et al (2000) The protein data bank. Nucleic Acids Res 28:235–242
Kaplan W, Littlejohn TG (2001) Swiss-PDB viewer (deep view). Brief Bioinform 2:195–197
Hess B, Kutzner C, Spoel DVD, Lindahl E. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J Chem Theory Comput 4:435–447
Gallivan JP, Dougherty DA (1999) Cation-pi interactions in structural biology. Proc Natl Acad Sci U S A 96:9459–9464
Magyar C, Gromiha MM, Pujadas G et al (2005) SRide: a server for identifying stabilizing residues in proteins. Nucleic Acids Res 33:W303–W305
Berendsen HJC, Postma JPM, van Gunsteren WF et al (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 8:3684–3690
Cheatham TE III, Miller JL, Fox T et al (1995) Molecular dynamics simulations on solvated biomolecular systems: the particle mesh Ewald method leads to stable trajectories of DNA, RNA, and proteins. J Am Chem Soc 117:4193–4194
Turner PJ (2005) XMGRACE, version 5.1.19. Center for Coastal and Land-Margin Research, Oregon Graduate Institute of Science and Technology, Beaverton
Amadei A, Linssen AB, Berendsen HJ (1993) Essential dynamics of proteins. Proteins 17:412–425
Duhovny D, Nussinov R, Wolfson HJ (2002) Efficient unbound docking of rigid molecules. In Gusfield et al. (ed) Proceedings of the 2’nd Workshop on Algorithms in Bioinformatics(WABI) Rome, Italy, lecture notes in computer science 2452. Springer Verlag, pp 185–200
Schneidman-Duhovny D, Inbar Y, Nussinov R, Wolfson HJ (2005) PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res 33:W363–W367
Andrusier N, Nussinov R, Wolfson HJ (2007) FireDock: fast interaction refinement in molecular docking. Proteins 69(1):139–159
Mashiach E, Schneidman-Duhovny D, Andrusier N et al (2008) FireDock: a web server for fast interaction refinement in molecular docking. Nucleic Acids Res 36:W229–W232
Verma D, Jacobs DJ, Livesay DR (2012) Changes in lysozyme flexibility upon mutation are frequent, large and long-ranged. PLoS Comput Biol 8:e1002409
Ribeiro AA, de Alencastro RB (2013) Mixed Monte Carlo/molecular dynamics simulations of the prion protein. J Mol Graph Model 42:1–6
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22:2577–2637
Acknowledgments
Authors gratefully acknowledge the management of Vellore Institute of Technology for providing the facilities to carry out this work.
Author information
Authors and Affiliations
Corresponding author
Additional information
Balu Kamaraj and Vidya Rajendran equally contributed to this paper.
Rights and permissions
About this article
Cite this article
Kamaraj, B., Rajendran, V., Sethumadhavan, R. et al. In-silico screening of cancer associated mutation on PLK1 protein and its structural consequences. J Mol Model 19, 5587–5599 (2013). https://doi.org/10.1007/s00894-013-2044-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-013-2044-0