Abstract
On the basis of spectroscopic and crystallographic data for dopamine β-monooxygenase and peptidylglycine α-hydroxylating monooxygenase (PHM), a variety of ligand sets have been used to model the oxygen-binding Cu site in these enzymes. Calculations which employed a combination of density functional and multireference second-order perturbation theory methods provided insights into the optimal ligand set for supporting η 1 superoxo coordination as seen in a crystal structure of a precatalytic Cu/O2 complex for PHM (Prigge et al. in Science 304:864–867, 2004). Anionic ligand sets stabilized η 2 dioxygen coordination and were found to lead to more peroxo-like Cu–O2 complexes with relatively exergonic binding free energies, suggesting that these adducts may be unreactive towards substrates. Neutral ligand sets (including a set of two imidazoles and a thioether), on the other hand, energetically favored η 1 dioxygen coordination and exhibited limited dioxygen reduction. Binding free energies for the 1:1 adducts with Cu supported by the neutral ligand sets were also higher than with their anionic counterparts. Deviations between the geometry and energetics of the most analogous models and the PHM crystal structures suggest that the protein environment influences the coordination geometry at the CuB site and increases the lability of water bound to the preoxygenated reduced form. Another implication is that a neutral ligand set will be critical in biomimetic models in order to stabilize η 1 dioxygen coordination.




Similar content being viewed by others
Notes
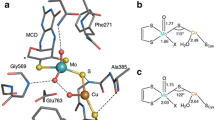
All calculations based directly on the PHM crystal structures used models in which bonds from Cu to ligating atoms were fixed at the distances and angles present in the crystal structure (see the “Choice of model systems” section). Consequently, the main geometric parameters in these models are those shown in Fig. 1.
References
Klinman JP (1996) Chem Rev 96:2541–2561
Stewart LC, Klinman JP (1988) Annu Rev Biochem 57:551–592
Eipper BA, Stoffers DA, Mains RE (1992) Annu Rev Neurosci 15:57–85
Eipper BA, Milgram SL, Husten EJ, Yun HY, Mains RE (1993) Protein Sci 2:489–497
McMahon A, Geertman R, Sabban EL (1990) J Neurosci Res 25:395–404
Stoffers DA, Green CB, Eipper BA (1989) Proc Natl Acad Sci USA 86:735–739
Tian GC, Berry JA, Klinman JP (1994) Biochemistry 33:226–234
Evans JP, Ahn K, Klinman JP (2003) J Biol Chem 278:49691–49698
Prigge ST, Kolhekar AS, Eipper BA, Mains RE, Amzel LM (1997) Science 278:1300–1305
Prigge ST, Kolhekar AS, Eipper BA, Mains RE, Amzel LM (1999) Nat Struct Biol 6:976–983
Prigge ST, Eipper BA, Mains RE, Amzel LM (2004) Science 304:864–867
Solomon EI, Chen P, Metz M, Lee SK, Palmer AE (2001) Angew Chem Int Ed Engl 40:4570–4590
Lewis EA, Tolman WB (2004) Chem Rev 104:1047–1076
Mirica LM, Ottenwaelder X, Stack TDP (2004) Chem Rev 104:1013–1045
Chen P, Bell J, Eipper BA, Solomon EI (2004) Biochemistry 43:5735–5747
Chen P, Solomon EI (2004) J Am Chem Soc 126:4991–5000
Champloy F, Benali-Cherif N, Bruno P, Blain I, Pierrot M, Reglier M, Michalowicz A (1998) Inorg Chem 37:3910–3918
Santra BK, Reddy PAN, Nethaji M, Chakravarty AR (2002) Inorg Chem 41:1328–1332
Kodera M, Kita T, Miura I, Nakayama N, Kawata T, Kano K, Hirota S (2001) J Am Chem Soc 123:7715–7716
Osako T, Nagatomo S, Tachi Y, Kitagawa T, Itoh S (2002) Angew Chem Int Ed Engl 41:4325–4328
Fujisawa K, Tanaka M, Morooka Y, Kitajima N (1994) J Am Chem Soc 116:12079–12080
Chen P, Root DE, Campochiaro C, Fujisawa K, Solomon EI (2003) J Am Chem Soc 125:466–474
Spencer DJE, Aboelella NW, Reynolds AM, Holland PL, Tolman WB (2002) J Am Chem Soc 124:2108–2109
Aboelella NW, Kryatov SV, Gherman BF, Brennessel WW, Young VG Jr, Sarangi R, Rybak-Akimova EV, Hodgson KO, Hedman B, Solomon EI, Cramer CJ, Tolman WB (2004) J Am Chem Soc 126:16896–16911
Reynolds AM, Gherman BF, Cramer CJ, Tolman WB (2005) Inorg Chem 44:6989–6997
Reynolds AM, Lewis EA, Aboelella NW, Tolman WB (2005) Chem Commun 2014–2016
Blackburn NJ, Hasnain SS, Pettingill TM, Strange RW (1991) J Biol Chem 266:23120–23127
Reedy BJ, Blackburn NJ (1994) J Am Chem Soc 116:1924–1931
Boswell JS, Reedy BJ, Kulathila R, Merkler D, Blackburn NJ (1996) Biochemistry 35:12241–12250
Blackburn NJ, Rhames FC, Ralle M, Jaron S (2000) J Biol Inorg Chem 5:341–353
Kolhekar AS, Keutmann HT, Mains RE, Quon ASW, Eipper BA (1997) Biochemistry 36:10901–10909
Schrodinger LLC (2002) Jaguar 5.0. Schrodinger, Portland, OR
Johnson BG, Gill PMW, Pople JA (1993) J Chem Phys 98:5612–5626
Becke AD (1993) J Chem Phys 98:1372–1377
Lee CT, Yang WT, Parr RG (1988) Phys Rev B 37:785–789
Ducere J-M, Goursot A, Berthomieu D (2005) J Phys Chem A 109:400–408
Gherman BF, Cramer CJ (2004) Inorg Chem 43:7281–7283
Hay PJ, Wadt WR (1985) J Chem Phys 82:270–283
Wadt WR, Hay PJ (1985) J Chem Phys 82:284–298
Hay PJ, Wadt WR (1985) J Chem Phys 82:299–310
Bauschlicher CW, Partridge H (1995) J Chem Phys 103:1788–1791
Cramer CJ (2004) Essentials of computational chemistry. Theories and models, 2nd edn. Wiley, Chichester
Abraham MH, Andonian-Haftvan J, Whiting GS, Leo A, Taft RS (1994) J Chem Soc Perkin Trans 2 1777–1791
Thompson JD, Cramer CJ, Truhlar DG (2005) Theor Chem Acc 113:107–131
Marten B, Kim K, Cortis C, Friesner RA, Murphy RB, Ringnalda MN, Sitkoff D, Honig B (1996) J Phys Chem 100:11775–11788
Tannor DJ, Marten B, Murphy R, Friesner RA, Sitkoff D, Nicholls A, Ringnalda M, Goddard WA, Honig B (1994) J Am Chem Soc 116:11875–11882
Tissandier MD, Cowen KA, Feng WY, Gundlach E, Cohen MH, Earhart AD, Coe JV, Tuttle TR (1998) J Phys Chem A 102:7787–7794
Ross S (1985) The proton, applications to organic chemistry. Academic, Orlando
Lide DR (2004) Handbook of chemistry and physics, 85th edn. CRC, Boca Raton
Andersson K, Malmqvist PA, Roos BO (1992) J Chem Phys 96:1218–1226
Karlstrom G, Lindh R, Malmqvist PA, Roos BO, Ryde U, Veryazov V, Widmark PO, Cossi M, Schimmelpfennig B, Neogrady P, Seijo L (2003) Comput Math Sci 28:222–239
Barandiaran Z, Seijo L (1992) Can J Chem 70:409–415
Pierloot K, Dumez B, Widmark PO, Roos BO (1995) Theor Chim Acta 90:87–114
Carvajal MA, Novoa JJ, Alvarez S (2004) J Am Chem Soc 126:1465–1477
Fox BS, Beyer MK, Bondybey VE (2002) J Am Chem Soc 124:13613–13623
Schmiedekamp AM, Ryan MD, Deeth RJ (2002) Inorg Chem 41:5733–5743
Feller D, Glendening ED, de Jong WA (1999) J Chem Phys 110:1475–1491
Burda JV, Pavelka M, Simanek M (2004) THEOCHEM 683:183–193
Pavelka M, Burda JV (2005) Chem Phys 312:193–204
Cramer CJ, Tolman WB, Theopold KH, Rheingold AL (2003) Proc Natl Acad Sci USA 100:3635–3640
Vaska L (1976) Acc Chem Res 9:175–183
Record MT Jr, Anderson CF, Mills P, Mossing M, Roe JH (1985) Adv Biophys 20:109–135
von Hippel PH (1994) Science 263:769–770
Spolar RS, Record MT Jr (1994) Science 263:777–784
Dixit SB, Jayaram B (1998) J Biomol Struct Dyn 16:237–242
Kinsinger CR, Gherman BF, Gagliardi L, Cramer CJ (2005) J Biol Inorg Chem 10:778–789
Kamachi T, Kihara N, Shiota Y, Yoshizawa K (2005) Inorg Chem 44:4226–4236
McCracken J, Desai PR, Papadopoulos NJ, Villafranca JJ, Peisach J (1988) Biochemistry 27:4133–4137
Brenner MC, Klinman JP (1989) Biochemistry 28:4664–4670
Acknowledgements
We acknowledge support from the NIH through an NRSA postdoctoral fellowship to B.F.G. and from the University of Minnesota Department of Chemistry through a Gleysteen Scholarship to D.E.H. The National Science Foundation (CHE-0203346 to C.J.C.) and the NIH (GM47365 to W.B.T.) are also thanked for partial support for this work. We thank Edward Solomon and Peng Chen for stimulating discussions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Gherman, B.F., Heppner, D.E., Tolman, W.B. et al. Models for dioxygen activation by the CuB site of dopamine β-monooxygenase and peptidylglycine α-hydroxylating monooxygenase. J Biol Inorg Chem 11, 197–205 (2006). https://doi.org/10.1007/s00775-005-0066-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-005-0066-5