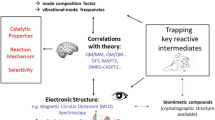
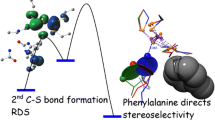
Abstract
The nonheme iron enzyme ScoE catalyzes the biosynthesis of an isonitrile substituent in a peptide chain. To understand details of the reaction mechanism we created a large active site cluster model of 212 atoms that contains substrate, the active oxidant and the first- and second-coordination sphere of the protein and solvent. Several possible reaction mechanisms were tested and it is shown that isonitrile can only be formed through two consecutive catalytic cycles that both use one molecule of dioxygen and α-ketoglutarate. In both cycles the active species is an iron(IV)-oxo species that in the first reaction cycle reacts through two consecutive hydrogen atom abstraction steps: first from the N–H group and thereafter from the C–H group to desaturate the NH-CH2 bond. The alternative ordering of hydrogen atom abstraction steps was also tested but found to be higher in energy. Moreover, the electronic configurations along that pathway implicate an initial hydride transfer followed by proton transfer. We highlight an active site Lys residue that is shown to donate charge in the transition states and influences the relative barrier heights and bifurcation pathways. A second catalytic cycle of the reaction of iron(IV)-oxo with desaturated substrate starts with hydrogen atom abstraction followed by decarboxylation to give isonitrile directly. The catalytic cycle is completed with a proton transfer to iron(II)-hydroxo to generate the iron(II)-water resting state. The work is compared with experimental observation and previous computational studies on this system and put in a larger perspective of nonheme iron chemistry.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Enzymes are versatile catalysts in Nature and often contain transition metal cofactors. In particular, iron is a common element due to its large natural abundance but also thanks to its position in the periodic table, which enables it to be involved in oxidation as well as reduction processes. The mononuclear nonheme iron dioxygenases are found in most forms of life, including plants, humans and bacteria and catalyse oxygen atom transfer reactions to substrates [1,2,3,4,5,6,7,8]. The most common reaction is typically aliphatic hydroxylation of a substrate and, for instance, is the precursor step in AlkB repair enzymes that dealkylate methylated DNA bases in the body [9]. Another reaction catalysed in the human body by nonheme iron dioxygenases relates to the biosynthesis of R-4-hydroxyproline, an essential component of collagen strands [10, 11].
Most nonheme iron dioxygenases have a central iron(II) atom linked to the protein through interactions with typically two His side chains and a carboxylate side chain of the protein in a facial orientation. A six-coordinated metal atom; therefore, has two binding positions available for substrate and co-substrate. A common co-substrate in the nonheme iron dioxygenases is α-ketoglutarate (αKG), which binds in the equatorial plane and reacts with dioxygen on the iron center to form an iron(IV)-oxo species and succinate through release of CO2 [12,13,14]. The iron(IV)-oxo species is believed to be the active oxidant of nonheme iron dioxygenases but short-lived. However, for several nonheme iron dioxygenases it was trapped and characterized spectroscopically [15,16,17,18,19]. Computational studies [20,21,22,23] confirmed the experimentally proposed catalytic cycle and showed that starting from an iron(II) ion ligated to three water molecules, αKG binding displaces two water ligands and bind to iron through the keto and carboxylate groups as a bidentate ligand. Subsequently, dioxygen replaces the last water ligand of the iron(II) center to form an end-on iron(III)-superoxo intermediate, which attacks the α-keto position of αKG to form a bicyclic ring structure. Thereafter, dioxygen bond cleavage takes place and releases CO2 to form an iron(IV)-oxo species and succinate. The steps leading up to the iron(IV)-oxo species encounter small reaction barriers of the order of 10 kcal mol−1 and lead to an iron(IV)-oxo species with large exothermicity. Consequently, the formation of the iron(IV)-oxo species is fast and no hard evidence of any other oxygen-bound intermediates exists. For cysteine dioxygenase a combination of UV–Vis absorption and electron paramagnetic resonance studies implicated a short-lived oxygen-bound intermediate, which was tentatively identified as either the iron(III)-superoxo species or the bicyclic ring structure [24]. Many computational studies investigated the structure and properties of the iron(IV)-oxo species of nonheme iron dioxygenases in detail using either density functional theory model complexes or quantum mechanics/molecular mechanics methods [25,26,27,28,29,30,31,32,33,34,35,36]. These studies identified the iron(IV)-oxo as a quintet spin ground state showed it to be an efficient oxidant of substrate hydroxylation reactions.
The biosynthesis of isonitrile groups by enzymes is unusual, but several have been reported that catalyse the reaction efficiently [37,38,39,40,41,42,43,44,44]. Most of those enzymes are flavin-dependent; however, recently, a nonheme iron dioxygenase was identified that biosynthesized an isonitrile group from R-3-((carboxymethyl)amino)butanoic acid through an oxidative decarboxylation reaction [43]. Thus, the isonitrile lipopeptide biosynthesis in Streptomyces coeruleorubidus is performed by a chain of enzymes, whereby the ScoE enzyme performs the oxidative decarboxylation reaction on an iron(II) center in the presence of dioxygen and αKG. A crystal structure (Fig. 1) was resolved at 1.8 Å resolution on a Zn(II)-containing and substrate-devoid system, which structurally resembles the well-characterized active site of taurine/αKG dependent dioxygenase [45, 46] and hence was identified as a nonheme iron dioxygenase.
The ScoE structure shows the metal bound to two His ligands (His132 and His295) and an aspartate group of Asp234, whereby the pdb structure contains Zn(II) instead of Fe(II). Both substrate and αKG are missing from the pdb file, but acetate is bound and likely is in the position of αKG. The substrate binding pocket is empty, but the Arg310 side chain has been hypothesized [45] to be involved in substrate binding and positioning in the active site. In addition, a number of aromatic residues lines the substrate binding pocket, namely Phe110, Phe137 and Phe239. These are probably involved with substrate positioning but their function remain unclear. Biochemical studies on ScoE confirmed the enzymatic reaction products using chromatography [45]. Further studies estimated that for each turnover of substrate at least four times the amount of succinate was produced, which implicates that two molecules of αKG are used in the process and significant amount of uncoupling happens [47]. The authors managed to trap the iron(IV)-oxo species and run Mössbauer experiments, which identify it as a high-spin (quintet) species with isomer shift δ = 0.31 mm s−1 and quadrupole splitting of ∆EQ = 0.59 mm s−1. The authors proposed a mechanism with two sequential substrate hydroxylation steps followed by a decarboxylation to form the isonitrile-containing product.
Recently, Li and Liu [48] reported a quantum mechanics/molecular mechanics study on the possible mechanism of ScoE. The authors showed that the mechanism starts from an iron(IV)-oxo species that reacts with substrate through two consecutive hydrogen atom abstraction steps that desaturate the substrate. The order of these first two hydrogen atom abstraction steps was shown to be unimportant and led to desaturated product with similar barriers. They proposed succinate to be released and the active site binds another molecule of αKG and dioxygen to form another iron(IV)-oxo species that abstracts a hydrogen atom from the substrate, which is followed by a decarboxylation step to form the isonitrile-containing product. Despite the fact that the results matched experiment well, the QM region in the QM/MM calculations only contained 107 atoms and had overall charge of − 2, which may lead to biased results. In particular, the authors found a competitive pathway for substrate hydroxylation and decarboxylation, while no substrate hydroxylation products are known from the literature [48]. As such we decided to do a computational study using density functional theory (DFT) cluster models with a substantially larger QM region of 212 atoms and overall neutral charge. The work shows competitive pathways of C–H versus N–H activation steps for an initial desaturation cycle that produces the desaturated substrate, water, succinate and CO2. Subsequently, a second catalytic cycle is followed by α-KG and dioxygen binding that leads to abstraction of the final hydrogen atom from the isonitrile group and decarboxylation to give final products. The DFT calculations highlight the important contribution of the active site Lys residue that brings a positive charge into the substrate binding pocket and affects the kinetics of the reaction through electrostatic interactions. The results are compared to the literature and rationalized with thermochemical cycles.
2 Methods
2.1 Model Set-Up
We use DFT cluster models based on the active site of the ScoE enzyme with procedures to generate the models as described before [49,51,51]. The model was created from the crystal structure coordinates deposited in the 6DCH protein databank (pdb) file [45, 46]. This is an enzyme monomer of ScoE with Zn(II), acetate, chloride and choline bound. Our model with the residues of the active site and second coordination sphere are specified in Scheme 1. We made some manual changes from the crystal structure coordinates and replaced the Zn(II) ion by Fe(IV), while acetate was replaced by succinate (modelled as acetate). The active site water molecule was replaced by an oxo group and positioned at a distance of 1.62 Å from the iron. The choline and chloride ions in the active site were removed and (R)-3-((carboxymethyl)amino)butanoic acid (CABA) was manually docked into the substrate binding pocket in such a way that its carboxylic acid groups form salt bridges with the side chains of Lys193 and Arg310. Subsequently, hydrogen atoms were added to the structure in Chimera [52], whereby all Asp and Glu side-chains were deprotonated, all Arg and Lys side-chains protonated and all His residues singly protonated.
Using previously described procedures on cluster models [53, 54], we created a large active site cluster model that incorporates the metal and its first-coordination sphere of ligands as well as a number of second-coordination sphere residues that determine the size and shape of the substrate binding pocket and incur hydrogen bonding interactions. Thus, the model includes the protein chain Phe130-Trp131-His132-Ala133-Asp134, whereby Trp131 and Ala133 were abbreviated to Gly. The axial histidine ligand (His295) of iron was truncated to methylimidazole and succinate to acetate. In addition, the side chains of the residues Tyr96, Tyr101, Phe110, Phe137, Lys193, Phe239 and Arg310 were included. Overall our model had 212 atoms in total and was overall charge neutral. As the model contains many internal hydrogen bonding interactions, no constraints were put on the system.
2.2 Procedures
The Gaussian-09 software package was used for all quantum chemical calculations discussed here [55]. Following previous experience with cluster models of nonheme iron dioxygenases [53, 56, 57], we utilized the unrestricted B3LYP density functional method [58, 59] in combination with a LANL2DZ (with electron core potential) on iron and 6-31G on the rest of the atoms: basis set BS1 [60, 61]. To correct the energetics, single point calculations with the LACV3P + (with electron core potential) on iron and 6–311 + G* on the rest of the atoms were performed (basis set BS2). The latter set of calculations included a continuum polarized conductor model (CPCM) with a dielectric constant mimicking chlorobenzene [62]. Single point calculations with the GD3 dispersion model of Grimme et al. were also performed, but gave very little changes to the relative energies and predicted the same selectivities [63].
Frequency calculations were performed on all local minima and transition states and it was confirmed that local minima had real frequencies only, while the transition states had a single imaginary mode for the correct vibration along the reaction coordinate. Intrinsic reaction coordinate scans confirmed the nature of the transition states and that they connect to the reactants and products.
Thermochemical cycles use isolated substrate and oxidant (with only first coordination sphere of ligands) and represent energies at UB3LYP/BS2 with solvent and zero-point energies included.
3 Results and Discussion
We started with a full geometry optimization at UB3LYP/BS1 of the iron(IV)-oxo reactant complex (5Re) of ScoE using the model described above in Scheme 1. The DFT optimized geometry of 5Re is given in Fig. 2. As can be seen the overlay of the 5Re structure with the original crystal structure coordinates is very good and the metal with its first-coordination sphere of ligands are virtually in the same position. There are also some obvious changes due to the absence of a substrate in the crystal structure coordinates and the presence of choline and chloride in the active site in the pdb structures. In particular, the charged side chains of Lys193 and Arg310 have bent and form salt bridge interactions with the carboxylate groups of the substrate and hold it in position. As such, in this substrate binding position it appears that both N–H and C–H bonds of substrate are accessible for activation by the iron(IV)-oxo species. The Fe–O bond is 1.650 Å in length, which is typical for high-spin iron(IV)-oxo species and matches the experimentally determined values for taurine/α-ketoglutarate dependent dioxygenase and halogenases excellently [16, 17]. Moreover, previous computational studies on the iron(IV)-oxo species of nonheme iron dioxygenases found similar values [12,13,14, 20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36, 64,66,66].
The iron(IV)-oxo species is in a quintet spin ground state and its molecular valence orbitals are shown on the right-hand-side of Fig. 2, where we define the z-axis along the Fe–O bond. In the xy-plane the π*xy and σ*x2-y2 orbitals are located for the antibonding interactions of the metal with first-coordination sphere ligands namely the nitrogen atom of His132 and the carboxylate oxygen atoms of Asp134 and succinate. In addition, there are antibonding orbitals along the Fe–O bond between 2p orbitals on the oxo atom and the 3d orbitals on iron: π*xz and π*yz. The final orbital is along the z-axis and is antibonding between the 3dz2 orbital on iron with 2pz orbitals on the axial His group (His295) and the oxo group. In the quintet spin state these orbitals are occupied with four electrons as π*xy1 π*xz1 π*yz1 σ*x2-y21, while in the triplet the orbital occupation is π*xy2 π*xz1 π*yz1. Experimental studies on various nonheme iron dioxygenases characterized their iron(IV)-oxo species as a quintet spin state using electron paramagnetic resonance and Mössbauer spectroscopy studies [15,16,17,18,19].
Next we considered substrate (R)-3-((carboxymethyl)amino)butanoic acid (CABA) activation by the iron(IV)-oxo species of ScoE enzymes using the possible mechanisms shown in Scheme 2. Thus, it has been proposed that the first cycle of ScoE leads to desaturation of the substrate. This can happen through an initial N–H hydrogen atom abstraction via TS1A to form a radical intermediate IM1A followed by C–H hydrogen atom abstraction via TS2A to form the desaturated products IM2 (pathway 1 in Scheme 2). Alternatively, the reverse order of the hydrogen atom abstractions can take place with initial C–H hydrogen atom abstraction via TS1B to form radical intermediate IM1B that leads after N–H hydrogen atom abstraction via TS2B to the desaturated product IM2 (pathway 2 in Scheme 2). From the IM1B radical intermediate, we also considered OH rebound via TS3B to form intermediate IM3B (pathway 3 in Scheme 2) as often aliphatic hydroxylation is favourable over desaturation [67, 68]. The final pathway for the first ScoE cycle is a direct decarboxylation through 1,2-hydrogen transfer in IM2 via transition state TS3A to form isonitrile products and formate Prod1 (pathway 4 in Scheme 2).
In addition to this single dioxygen cycle, we considered the possibility where IM2 releases its distal water molecule and succinate and picks up new molecules of αKG and O2 to start a second oxygenation cycle that forms another iron(IV)-oxo intermediate (pathway 5 in Scheme 2). To this end, we took the structure of IM2 and replaced iron(II)(water) with iron(IV)-oxo to form 5Re2A. This keeps the charge and multiplicity of the model the same. Subsequently, we investigated a decarboxylation reaction via a mechanism reported before for cytochrome P450 OleT [69, 70] that starts with a hydrogen atom abstraction from substrate via transition state TS4A to form radical intermediate IM3A. The latter can proceed through decarboxylation via transition state TS5A to give decarboxylated products (Prod2) or react through OH rebound via transition state TS6A to form the alcohol product (IM5A) instead.
3.1 Substrate Desaturation Via Pathways 1 and 2
Figure 3 shows the results of the calculations on the substrate desaturation pathways according to the mechanisms proceeding through pathways 1 and 2. The reactions are both stepwise via an intermediate, although they have a different electronic configuration. As can be seen the lowest energy hydrogen atom abstraction barrier is from the N–H group of the substrate. At UB3LYP/BS1 level of theory this barrier is only 3.1 kcal mol−1 in energy, but addition of zero-point energy and solvent corrections lowers it to close to zero and hence will proceed very fast. Upon hydrogen atom abstraction the system collapses to a radical intermediate 5IM1A that is lower in energy than the reactant complex by ∆E + ZPE = − 14.5 kcal mol−1. The second hydrogen atom abstraction barrier (from the C–H group in 5IM1A); however, encounters a significant energy of 23.7 kcal mol−1 above 5IM1A and consequently will be slow and rate-determining. Nevertheless, this pathway will lead to desaturated substrate (at 5IM2A) that is lower in energy than reactants by ∆E + ZPE = − 53.3 kcal mol−1.
Potential energy landscape for substrate desaturation via pathways 1 (in blue) and 2 (in red) as calculated with B3LYP. Energies (with respect to 5Re in kcal mol−1) represent ∆E + ZPE (∆G) values and are obtained at UB3LYP/BS2//UB3LYP/BS1 with solvent included. Optimized geometries of the transition states give bond lengths in angstroms and the imaginary frequency in cm−1
Next we calculated pathway 2 and find a C-H hydrogen atom abstraction barrier of 13.4 kcal mol−1 leading to an intermediate 5IM1B that is considerably more stable than the reactant complex by ∆E + ZPE = − 35.3 kcal mol−1. An analysis of the group spin densities and orbital occupations of 5IM1B; however, shows no radical character on the substrate and only unpaired spin density on the iron atom. As such, the substrate has relaxed to a closed-shell system with a positively charged nitrogen atom in a double bond configuration with the NH = CHCOO group. The 5IM1B intermediate; therefore is the result of a hydride transfer from substrate to iron(IV)-oxo species and gives an iron(II)-hydroxo complex with electronic configuration π*xy2 π*xz1 π*yz1 σ*x2− y21 σ*z21. The reaction from 5IM1B is followed by a small proton transfer step from substrate to iron(II)-hydroxo via 5TS2B then gives the iron(II)-water complex, i.e. 5IM2A, in another exothermic reaction. Although the 5TS2B transition state was fully optimized and characterized with an imaginary frequency of i735 cm−1 corresponding to the O–H–N stretch vibration, actually its energy is very low and close to that of 5IM1B. In particular, addition of zero-point, thermal and entropic corrections to the energy lowers 5TS2B slightly below the intermediate; hence its value in Fig. 3 is reported as ∆E + ZPE < − 34 kcal mol−1. The 5TS2B transition state is central with an O–H distance of 1.268 Å and an N–H distance of 1.273 Å for the transferring hydrogen atom. The orientation is relatively upright with an Fe–O–N angle of 153°.
Overall, the calculations on the large DFT cluster model of ScoE give relatively low barriers and particularly a low N–H abstraction barrier of only a few kcal mol−1. These results contradict the QM/MM results of Li and Liu [48] that predicted much higher barriers of 9.8 and 17.6 kcal mol−1 for hydrogen atom abstraction from the N–H and C–H groups, respectively, although the ordering is the same. The most probable reason for the difference in energetics is the missing Lys residue in the QM region of ScoE in the work of Li and Liu. In our cluster model, this Lys side chain is positively charged and forms an ionic interaction with the carboxylate group of the substrate, which in Ref [48] does not interact with any positively charged amino acid. Its stronger binding positions the substrate differently and assists the reaction for desaturation of the N–C bond of CABA substrate.
The optimized geometry of the two hydrogen atom abstraction transition states along pathway 1 are given in Fig. 3. The N–H hydrogen atom abstraction barrier is central with almost equal O–H and N–H bond lengths of 1.243 and 1.246 Å, respectively. The Fe–O distance has elongated to 1.796 Å and the axial Fe–N bond to 2.204 Å. These changes mimic previous calculations on analogous reaction mechanisms well [26,27,28,29,30,31,32,33,34,35,36]. The transition state has a modest imaginary frequency of i619 cm−1, which is well lower than typical hydrogen atom abstraction transition states that usually have values well over i1000 cm−1 [72,73,73]. As such, the replacement of hydrogen by deuterium is not expected to give a major kinetic isotope effect as sometimes seen in nonheme iron reactivities [16]. Interestingly, the transition state 5TS2A has a much larger imaginary frequency of i1182 cm−1 and for that step; therefore, a considerable kinetic isotope effect for replacement of hydrogen by deuterium may be expected. Geometrically, 5TS2A is more reactant-like with a long O–H distance of 1.436 Å, while a much shorter C–H distance of 1.221 Å is found.
The optimized geometry of 5TS1B has a first coordination sphere similar to that seen for 5TS1A with an Fe–O and Fe–N distances of 1.729 and 2.055 Å, respectively. The transition state is early with a long O–H and short C–H distance of 1.557 and 1.177 Å. Early transition states often correspond with higher reaction barriers than late-transition states [71,73,74,74] and indeed 5TS1B is higher in energy than 5TS1A. Interestingly; however, the subsequent intermediates are reversed with a larger exothermicity for 5IM1B than for 5IM1A. To understand the change in ordering, we analyzed the electronic changes during the reaction mechanism and display the group spin densities of 5TS1A, 5TS1B and 5TS2A in Fig. 4. As can be seen 5TS1A is a hydrogen atom abstraction transition state where considerable amount of spin has accumulated already on the substrate (ρSub = − 0.98). This is unusual as normally in hydrogen atom abstraction transition states only partial spin on the substrate is found [71,73,74,74]. Therefore, this is an electronically early transition state where a full electron transfer has happened already. Previous studies on N–H activation by an iron(IV)-oxo heme cation radical model of nitric oxide synthase also showed early electron transfer processes [75]. To further check the changes in charge distributions we calculated the charge-transfer (QCT) of the substrate as the charge on the substrate in 5Re minus the charge in the transition state. As can be seen a large amount of charge-transfer has taken place in 5TS1A (QCT = 0.55), while it is much less in 5TS1B (QCT = 0.18).
The group spin densities of 5TS2A implicate a positive spin on the substrate (ρSub = 0.77), while its precursor intermediate (5IM1A) has negative amount (ρSub = − 1.00). We attempted to swap molecular orbitals for 5TS2A to create a system with negative spin on the substrate instead; however, the SCF converged back to a state with positive spin on substrate instead. The C–H abstraction transition state for pathway 2 also shows a positive spin density on the substrate (ρSub = 0.38) and hence is the result of a down-spin electron transfer from substrate to iron-oxo to give an electronic configuration π*xy2 π*xz1 π*yz1 σ*x2-y21 on iron coupled to an up-spin electron on substrate. This type of electron transfer mechanism is generally called the π-pathway, while the transfer of an up-spin electron to give an electronic configuration on iron of π*xy1 π*xz1 π*yz1 σ*x2−y21 σ*z21 is generally called a σ-pathway [76,78,79,79]. Usually the σ-pathway is the lowest in energy and rate-determining. We attempted the swap molecular orbitals and convert 5TS1B to a σ-type transition state, but during the SCF convergence it reverted back to the π-pathway configuration instead. It appears the geometry of 5TS1B is not appropriate for a σ-type transition state as usually these types of structures show attack from the top with an almost linear Fe–O–C angle of close to 180 degrees [76,78,79,80,81,82,82]. In 5TS1B the Fe–O–C angle; however, is only 138° Most likely the constraints in the substrate binding pocket and the substrate approach to the iron(IV)-oxo species affect the electron transfer pathways and trigger a π-pathway rather than the more common σ-pathway.
3.2 Substrate Hydroxylation Via Pathway 3
Usually a hydrogen atom abstraction by an iron(IV)-oxo species is followed by OH rebound to form an alcohol complex [83, 84]. In previous work on studies of competitive hydroxylation and desaturation processes by either heme or nonheme iron(IV)-oxo complexes it was shown that generally substrate hydroxylation is more exothermic than substrate desaturation [85,87,87]. This is mainly due to the strength of the C–O bond that is formed, although when a highly conjugated π-system is formed, such as an arene, the desaturation can become competitive with hydroxylation. In ScoE no conjugated π-system is formed, hence we considered also OH rebound from 5IM1B to form the alcohol complex 5IM3B.
Figure 5 shows the energy landscape for pathway 3 as compared to pathway 2 for the bifurcation channel from 5IM1B leading to desaturation and hydroxylation of the substrate. As can be seen the desaturation channel has the lowest barrier (via 5TS2B) and leads to the most exothermic products (5IM2A), which are lower in energy by ∆E + ZPE = 18.0 kcal mol−1 with respect to 5IM2A. By contrast, the OH rebound barrier is about 3 kcal mol−1 higher in energy than the desaturation barrier and much less exothermic, namely by only 6.9 kcal mol−1 with respect to 5IM2A. Therefore, the most likely pathway for ScoE after C–H hydrogen atom abstraction will lead to desaturation of the NH-CH2 bond rather than OH rebound to form the corresponding alcohol.
Potential energy landscape for substrate hydroxylation via pathway 3 (in blue) as compared to desaturation via pathway 2 (in red) as calculated with B3LYP. Energies (with respect to 5Re in kcal mol−1) represent ∆E + ZPE (∆G) values and are obtained at UB3LYP/BS2//UB3LYP/BS1 with solvent included. Optimized geometry of the transition state gives bond lengths in angstroms and the imaginary frequency in cm−1
The OH rebound transition state 5TS3B was fully characterized and its geometry is shown on the left-hand-side of Fig. 4. The transition state has a low imaginary frequency of only i96 cm−1, but displays a C–O stretch vibration. The rebound happens at a C–O distance of 1.984 Å, which is relatively short as previously values of magnitude around 2.4 Å were obtained [88, 89]. The reason for the short C–O distance may be the position of the positively charged Lys group along the same line at a distance of 1.695 Å that withdraws electron density into the newly formed bond and assists the rebound.
Overall, the calculations show that OH rebound is lesser likely than substrate desaturation although the difference in barriers is not huge and some hydroxylation of the substrate may occur but probably with lesser probability than substrate desaturation. This result is in excellent agreement with recent nuclear magnetic resonance studies of isotopically labelled compounds that identified a small signal for hydroxylated products [47].
3.3 Substrate Decarboxylation Via Pathway 4
Before starting a second dioxygen activation cycle, we considered direct 1,2-hydrogen atom transfer in the desaturated product 5IM2A to form formate and isocyanide product directly and the potential energy profile with corresponding transition state via 5TS3A is shown in Fig. 6. The direct 1,2-hydrogen atom shift is high in energy and a barrier of 65.9 kcal mol−1 above 5IM2A was located. This high energy implies that the direct 1,2-hydrogen atom transfer will be unlikely in an enzymatic context and a single reaction cycle with one molecule of dioxygen will be unable to give isocyanide products directly. Interestingly, the isocyanide product is also higher in energy than the desaturated product at 5IM2A by 10.7 kcal mol−1. Consequently, the desaturated product may have a finite lifetime and may live long enough for it to be detected experimentally.
Potential energy landscape for substrate decarboxylation via pathway 4 as calculated with B3LYP. Energies (with respect to 5Re in kcal mol−1) represent ∆E + ZPE (∆G) values and are obtained at UB3LYP/BS2//UB3LYP/BS1 with solvent included. Optimized geometry of the transition state gives bond lengths in angstroms and the imaginary frequency in cm−1
The optimized geometry of the 1,2-hydrogen shift transition state 5TS3A is shown in Fig. 5. It has the transferring hydrogen atom midway between the two carbon atoms in an almost equilateral triangle and C–H distances of 1.885 and 1.854 Å. The imaginary frequency is i585 cm−1 and shows the hydrogen transfer from C to the carboxylate carbon atom. The reason the barrier is high in energy may have to do with the active site Lys residue that is close to the transferring hydrogen atom.
3.4 Substrate Decarboxylation Via Pathway 5
The DFT calculations reported above imply that a single molecule of dioxygen on an iron center with α-KG as co-substrate cannot form isonitrile products directly. Consequently, a second cycle with dioxygen and α-KG is needed. We; therefore, assumed that substrate would remain in the binding pocket and the enzyme will release succinate and bind new molecules of α-KG and dioxygen. To this end, we took structure 5IM2A and replaced the iron(II)-water with iron(IV)-oxo and assumed that in between these two stages water and succinate are released and new molecules of α-KG and dioxygen bind and react to form iron(IV)-oxo and succinate to give the second reactant complex 5Re2A. The optimized geometry of 5Re2A and the hydrogen atom abstraction pathway to form radical intermediate 5IM3A is shown in Fig. 7. The second reactant structure 5Re2A is very similar to 5ReA discussed above with Fe–O and Fe–N distances of 1.654 and 2.068 Å. These distances are within 0.01 Å from those of 5ReA described above and hence the structures are very similar.
Potential energy landscape for substrate decarboxylation via pathway 5 as calculated with B3LYP. Energies (with respect to 5Re2A in kcal mol−1) represent ∆E + ZPE (∆G) values and are obtained at UB3LYP/BS2//UB3LYP/BS1 with solvent included. Optimized geometry of 5Re2A, 5TS4A and 5TS5A give bond lengths in angstroms and the imaginary frequency in cm−1
Next, we explored hydrogen atom abstraction from the C–H bond of the desaturated substrate from 5Re2A. In principle, an sp2 hybridized C–H bond is strong and breaking this bond will require a large bond dissociation energy. This is unusual as iron(IV)-oxo species of heme and nonheme iron monooxygenases typically react with sp3 hybridized C–H bonds only and do not react with olefin C–H bonds through hydroxylation pathways [90,92,93,93]. Nevertheless, a hydrogen atom abstraction transition state was located for the ScoE model discussed here and has a barrier of ∆E + ZPE = 10.0 kcal mol−1. As such it is lower in energy than the first C–H hydrogen atom abstraction discussed above in Fig. 3. Our DFT calculated hydrogen atom abstraction barrier is in perfect agreement with the earlier QM/MM result of Li and Liu [48], who reported a barrier of 9.8 kcal mol−1 for the second step. Clearly, the choice of the QM region and the second-coordination sphere has little effect on the hydrogen atom abstraction for cycle 2 of ScoE and gives the same energetics and conclusions. We then explored the decarboxylation and OH rebound pathways from 5IM3A. The OH rebound has a relatively large barrier and a constraint geometry scan locates it at approximately 17 kcal mol−1 above 5IM3A. Most likely, the positive charge of Lys and the hydrogen bonding network surrounding the OH group hampers radical rebound and makes it high in energy Lower in energy; however, is the decarboxylation transition state 5TS5A at ∆E + ZPE = 11.9 kcal mol−1 above 5IM3A. As such decarboxylation to form the isonitrile product will be favourable over OH rebound to form an alcohol product. The 5TS5A transition state has a long C–C bond of 2.230 Å and an imaginary frequency of i232 cm−1 representing the C–C stretch vibration. The interaction between the side chain of Lys193 and the iron(III)-hydroxo group maintains short at 1.643 Å.
Figure 7 shows the optimized geometry of the hydrogen atom abstraction transition state for the second cycle. The transition state is central with an O–H distance of 1.307 Å and a C–H distance of 1.239 Å. The hydrogen atom transfer is accompanied by an imaginary frequency of i1054 cm−1, which will result in a significant kinetic isotope effect upon replacement of the transferring hydrogen atom by deuterium. The transition state is of σ-type with 3.99 spin density on the FeO group and negative spin accumulating on the substrate (ρSub = − 0.42). This transition state connect to a radical intermediate 5IM3A that indeed has radical character on the substrate: ρSub = − 0.98.
3.5 Thermochemical Rationalization of the Reaction Pathways
To understand the order of the hydrogen atom abstraction transition states we calculated the bond dissociation energies (BDEs) of various C–H and N–H bonds in the substrate. Often the strength of the C–H/N–H bond that is broken correlates with the barrier for hydrogen atom abstraction [71,73,73, 94,96,96]. Indeed for a series of substrates with known C–H bond strengths linear correlations between the natural logarithm of the rate constant with BDE was found. To calculate substrate BDE values, we took an isolated substrate molecule and also calculated the substrate with one hydrogen atom removed in a doublet spin state and took the relative energies between these structures and an isolated hydrogen atom. These DFT calculated BDEs are given in Fig. 8 for the substrate. As can be seen, in the gas-phase the weakest bond of the substrate is the C–H bond with a BDE1CH = 74.3 kcal mol−1, while the N–H bond is well stronger at BDE1NH = 80.7 kcal mol−1. Based on these relative BDE values; therefore, a regioselective C–H hydrogen atom abstraction would be expected rather than N–H abstraction. Consequently, the calculated BDE values are opposite of the hydrogen atom abstraction barriers shown above in Fig. 3 that predict regioselective N–H hydrogen atom abstraction instead. Clearly, the protein induces local perturbations to the active site pocket that change the relative C–H and N–H bond strengths. Previous work on nonheme iron dioxygenases showed that charged residues in the substrate binding pocket and active site, such as Lys residues, can donate positive charge into the binding pocket and affect local bonds strengths and selectivities dramatically [68, 97,99,99]. Moreover, studies using electric field effects showed that these external perturbations can influence electronic distributions as well as chemoselectivities of substrate activation by enzymes [100,102,102].
To put the energies in a better perspective, we also calculated the BDEOH for the strength of the O–H bond of the iron(III)-hydroxo complex that is formed after hydrogen atom abstraction. Using a minimal model of first-coordination sphere ligands to iron only, we calculate a ∆E + ZPE value for BDEOH,Fe(III) = 92.5 kcal mol−1. Technically, the exothermicity for hydrogen atom abstraction should equal the difference in energy between BDE1NH/BDE1CH and BDEOH,Fe(III) for initial hydrogen atom abstraction from the N–H or C–H bonds. This way, we find an exothermicity for hydrogen atom abstraction from the N–H bond of − 11.8 kcal mol−1, whereas for hydrogen atom abstraction from the C–H bond a value of − 18.2 kcal mol−1 is predicted. The value for N–H hydrogen atom abstraction exothermicity of − 11.8 kcal mol−1 is close in energy to the value obtained for 5IM1A of − 14.5 kcal mol−1. Consequently, this reaction step can be considered as hydrogen atom abstraction and follows the energies of the bond that are broken and formed in the process. The predicted hydrogen atom transfer from the C–H bond is only − 18.2 kcal mol−1, while 5IM1B is − 35.3 kcal mol−1 more exothermic than the reactant complex. Clearly, hydride transfer is more exothermic in this case than hydrogen atom transfer and consequently it is not observed in our model.
We also calculated hydride abstraction from the C–H bond of the substrate and find a BDE1HT = 34.8 kcal mol−1. In addition, we calculated the O–H bond formation as the BDEOH,Fe(II) for the energy to remove a hydride from an iron(II)-hydroxo complex. The BDEOH,Fe(II) for hydride release to form an iron(IV)-oxo from an iron(II)-hydroxo complex is calculated as 154.7 kcal mol−1. Therefore, hydride transfer will be highly exothermic and indeed during the energy landscape shown in Fig. 3 its exothermicity is well larger than hydrogen atom transfer from CABA substrate. The origin of the large exothermicity for hydride transfer is most likely that a stable double bond is formed in the product and the π-bond will add stability to the product and make this reaction more exothermic.
Finally, we calculated the second hydrogen atom abstraction for pathways 1 and 2 and find a BDE2CH = 45.7 kcal mol−1 for hydrogen atom abstraction from the C–H bond after the N–H abstraction, while the reverse ordering of hydrogen atom abstractions has a second bond strength of BDE2NH = 71.6 kcal mol−1. Both of these second bond energies are lower than those for the first hydrogen atom abstraction and consequently, homolytic splitting of C–H/N–H bonds to form desaturated substrate should give a rate-determining first hydrogen atom abstraction step. However, due to the presence of polar groups in the substrate binding pocket, the reaction mechanism does not follow the thermodynamics of the reaction.
3.6 Reaction Mechanism with Lys 193 Removed from the Model
Crystallographic studies showed that the position of the Lys193 residue in the ScoE appears to be dependent on whether substrate is bound or not. Thus, in the crystal structure coordinates of the resting state protein without substrate bound (6DCH pdb file) the Lys193 side chain points away from the active site, whereas in the most recent substrate-bound structure (6XN6 pdb file [103]) it points into the active site. In particular, the Lys193 side chain in the substrate-bound structure is seen to form a hydrogen bonding interaction with the carboxylate group of (R)-3-((carboxymethyl)amino)butanoic acid (CABA) substrate. When we set-up our model we also reasoned it would be important for the substrate carboxylate groups to interact with polar residues and manually changed the position of Lys193 and moved it into the pocket to make the substrate-binding orientation to match the position in the substrate-bound structure.
To test how removal of the Lys side chain in the cluster model will affect structure and reactivity, we tested the hydrogen atom abstraction step in the first cycle with the model with Lys193 removed. To this end we took the optimized geometries of 5Re, 5TS1A and 5TS1B and removed the Lys from the structure and did a B3LYP/BS2 single point calculation. Figure 9 shows the relative energies of these three structures with Lys removed as compared to those at the same level of theory for the full system. As can be seen 5TS1A is raised in energy by a small amount (5TS1A,noLys is ∆E = 4.6 kcal mol−1 above reactants). By contrast, dramatic energy differences are seen for 5TS1B,noLys that is raised to 26.1 kcal mol−1 with Lys removed. As such, a charged residue in the substrate binding pocket and particularly a positively charged residue that is hydrogen bonding to the oxo group has a major influence on the hydrogen atom abstraction barrier. This is in analogy to the hygromycin biosynthesis enzyme, where an active site Lys residue was shown to affect rate constants and reaction selectivities by guiding the reaction to a specific product channel [99]. Furthermore, work on the hectochlorin biosynthesis enzyme identified an active site Glu residue that was shown to donate negative charge to an iron-bound halide and influence the regioselectivity of halogenation over hydroxylation in a nonheme iron enzyme by polarizing the Fe-Cl bond better [98].
4 Conclusions and Summary
In this work a computational study is presented on the isonitrile biosynthesis enzyme ScoE and various likely mechanisms for substrate activation have been studied. A large DFT cluster model incorporating the first and second coordination sphere of the enzyme has been created. The work shows that competitive C–H and N–H hydrogen atom abstraction pathways lead to desaturation with large exothermicity. Alternative substrate hydroxylation and decarboxylation pathways were also investigated but found to be well higher in energy than substrate desaturation and hence are unlikely to take place. Instead the system does not release desaturated substrate but binds another molecule of O2 and α-KG to initiate a second catalytic cycle with another iron(IV)-oxo species. The latter abstracts a hydrogen atom, which triggers release of CO2 to form isonitrile products with a barrier of 10 kcal mol−1. The results of this work are consistent with experimental observations as well as analogous computational studies on this and related enzymes. Our work; however, points to the involvement of an active site Lys residue that in the reaction mechanism donates positive charge into the substrate binding pocket and through charge-stabilization assists in the chemical process and enables a chemoselective substrate desaturation reaction.
5 Data Availability
Supporting Information is available containing raw data including absolute and relative energies, Cartesian coordinates of optimized geometries and tables with group spin densities and charges.
Supporting Information is available containing raw data including absolute and relative energies, Cartesian coordinates of optimized geometries and tables with group spin densities and charges.
References
Schofield CJ, Zhang Z (1999) Structural and mechanistic studies on 2-oxoglutarate-dependent oxygenases and related enzymes. Curr Opin Struct Biol 9:722–731
Solomon EI, Brunold TC, Davis MI, Kemsley JN, Lee SK, Lehnert N, Neese F, Skulan AJ, Yang YS, Zhou J (2000) Geometric and electronic structure/function correlations in non-heme iron enzymes. Chem Rev 100:235–349
Costas M, Mehn MP, Jensen MP, Que L Jr (2004) Dioxygen activation at mononuclear nonheme iron active sites: enzymes, models, and intermediates. Chem Rev 104:939–986
Abu-Omar MM, Loaiza A, Hontzeas N (2005) Reaction mechanisms of mononuclear non-heme iron oxygenases. Chem Rev 105:2227–2252
Kovaleva EG, Lipscomb JD (2008) Versatility of biological non-heme Fe(II) centers in oxygen activation reactions. Nat Chem Biol 4:186–193
Bruijnincx PCA, van Koten G, Klein Gebbink RJM (2008) Mononuclear non-heme iron enzymes with the 2-His-1-carboxylate facial triad: recent developments in enzymology and modeling studies. Chem Soc Rev 37:2716–2744
de Visser SP, Kumar D (eds) (2011) Iron-containing enzymes: versatile catalysts of hydroxylation reactions in nature. Royal Society of Chemistry Publishing, Cambridge
White MD, Flashman E (2016) Catalytic strategies of the non-heme iron dependent oxygenases and their roles in plant biology. Curr Opin Chem Biol 31:126–135
O’Brien PJ (2006) Catalytic promiscuity and the divergent evolution of DNA repair enzymes. Chem Rev 106:720–752
Gorres K, Raines RT (2010) Prolyl 4-hydroxylase. Crit Rev Biochem Mol Biol 45:106–124
de Visser SP (2018) Mechanistic insight on the activity and substrate selectivity of nonheme iron dioxygenases. Chem Record 18:1501–1516
Borowski T, Blomberg MRA, Siegbahn PEM (2008) Reaction mechanism of apocarotenoid oxygenase (ACO): a DFT study. Chem Eur J 14:2264–2276
Sinnecker S, Svensen N, Barr EW, Ye S, Bollinger JM Jr, Neese F, Krebs C (2007) Spectroscopic and computational evaluation of the structure of the high-spin Fe(IV)-oxo intermediates in taurine:α-ketoglutarate dioxygenase from Escherichia coli and its His99Ala ligand variant. J Am Chem Soc 129:6168–6179
Godfrey E, Porro CS, de Visser SP (2008) Comparative quantum mechanics / molecular mechanics (QM/MM) and density functional theory calculations on the oxo-iron species of taurine/α-ketoglutarate dioxygenase. J Phys Chem A 112:2464–2468
Proshlyakov DA, Henshaw TF, Monterosso GR, Ryle MJ, Hausinger RP (2004) Direct detection of oxygen intermediates in the non-heme Fe enzyme taurine/α-ketoglutarate dioxygenase. J Am Chem Soc 126:1022–1023
Bollinger JM Jr, Price JC, Hoffart LM, Barr EW (2005) Krebs C (2005) Mechanism of taurine: α-ketoglutarate dioxygenase (TauD) from Escherichia coli. Eur J Inorg Chem 21:4245–4254
Galonić Fujimori D, Barr EW, Matthews ML, Koch GM, Yonce JR, Walsh CT, Bollinger JM Jr, Krebs C, Riggs-Gelasco PJ (2007) Spectroscopic evidence for a high-spin Br-Fe(IV)-oxo intermediate in the α-ketoglutarate-dependent halogenase CytC3 from Streptomyces. J Am Chem Soc 129:13408–13409
Light KM, Hangasky JA, Knapp MJ, Solomon EI (2013) Spectroscopic studies of the mononuclear non-heme FeII enzyme FIH: second-sphere contributions to reactivity. J Am Chem Soc 135:9665–9674
Dunham NP, Chang W-c, Mitchell AJ, Martinie RJ, Zhang B, Bergman JA, Rajakovich LJ, Wang B, Silakov A, Krebs C, Boal AK, Bollinger JM Jr (2018) Two distinct mechanisms for C-C desaturation by iron(II)- and 2-(oxo)glutarate-dependent oxygenases: importance of α-heteroatom assistance. J Am Chem Soc 140:7116–7126
Borowski T, Bassan A, Siegbahn PEM (2004) Mechanism of dioxygen activation in 2-oxoglutarate-dependent enzymes: a hybrid DFT study. Chem Eur J 10:1031–1041
de Visser SP (2007) Can the peroxosuccinate complex in the catalytic cycle of taurine/α-ketoglutarate dioxygenase (TauD) act as an alternative oxidant? Chem Commun 43:171–173
Wójcik A, Radoń M, Borowski T (2016) Mechanism of O2 activation by α-ketoglutarate dependent oxygenases revisited. A quantum chemical study. J Phys Chem A 120:1261–1274
Latifi R, Minnick JL, Quesne MG, de Visser SP, Tahsini L (2020) Computational studies of DNA base repair mechanisms by nonheme iron dioxygenases: selective epoxidation and hydroxylation pathways. Dalton Trans 49:4266–4276
Tchesnokov EP, Faponle AS, Davies CG, Quesne MG, Turner R, Fellner M, Souness RJ, Wilbanks SM, de Visser SP, Jameson GNL (2016) An iron-oxygen intermediate formed during the catalytic cycle of cysteine dioxygenase. Chem Commun 52:8814–8817
Geng C, Ye S, Neese F (2010) Analysis of reaction channels for alkane hydroxylation by nonheme iron(IV)–oxo xomplexes. Angew Chem Int Ed 49:5717–5720
de Visser SP (2006) Differences in and comparison of the catalytic properties of heme and non-heme enzymes with a central oxo-iron group. Angew Chem Int Ed 45:1790–1793
Chen H, Lai W, Yao J, Shaik S (2011) Perferryl FeV–oxo non-heme complexes: do they have high-spin or low-spin ground states? J Chem Theory Comput 7:3049–3053
Bushnell EAC, Fortowsky GB, Gauld JW (2012) Model iron–oxo species and the oxidation of imidazole: insights into the mechanism of OvoA and EgtB? Inorg Chem 51:13351–13356
Kulik HJ, Drennan CL (2012) Substrate placement influences reactivity in non-heme Fe(II) halogenases and hydroxylases. J Biol Chem 288:11233–11241
Quesne MG, Latifi R, Gonzalez-Ovalle LE, Kumar D, de Visser SP (2014) Quantum mechanics/molecular mechanics study on the oxygen binding and substrate hydroxylation step in AlkB repair enzymes. Chem Eur J 20:435–446
Álvarez-Barcia S, Kästner J (2017) Atom tunneling in the hydroxylation process of taurine/α-ketoglutarate dioxygenase identified by quantum mechanics/molecular mechanics simulations. J Phys Chem B 121:5347–5354
Timmins A, Saint-André M, de Visser SP (2017) Understanding how prolyl-4-hydroxylase structure steers a ferryl oxidant toward scission of a strong C-H bond. J Am Chem Soc 139:9855–9866
Manna RN, Malakar T, Jana B, Paul A (2018) Unraveling the crucial role of single active water molecule in the oxidative cleavage of aliphatic C−C bond of 2,4′-dihydroxy acetophenone catalyzed by 2,4′-dihydroxyacetophenone dioxygenase enzyme: a quantum mechanics/molecular mechanics investigation. ACS Catal 8:10043–10050
Iyer SR, Chaplin VD, Knapp MJ, Solomon EI (2018) O2 activation by nonheme FeII α-ketoglutarate-dependent enzyme variants: elucidating the role of the facial triad carboxylate in FIH. J Am Chem Soc 140:11777–11783
Xue J, Lu J, Lai W (2019) Mechanistic insights into the non-heme 2-oxoglutarate-dependent ethylene-forming enzyme: selectivity of ethylene-formation versus L-Arg hydroxylation. Phys Chem Chem Phys 21:9957–9968
Chaturvedi SS, Ramanan R, Lehnert N, Schofield CJ, Karabencheva-Christova TG, Christov CZ (2020) Catalysis by the non-heme iron(II) histone demethylase PHF8 involves iron center rearrangement and conformational modulation of substrate orientation. ACS Catal 10:1195–1209
Clarke-Pearson MF, Brady SF (2008) Paerucumarin, a new metabolite produced by the pvc gene cluster from Pseudomonas aeruginosa. J Bacteriol 190:6927–6930
Micallef ML, Sharma D, Bunn BM, Gerwick L, Viswanathan R, Moffitt MC (2014) Comparative analysis of hapalindole, ambiguine and welwitindolinone gene clusters and reconstitution of indole-isonitrile biosynthesis from cyanobacteria. BMC Microbiol 14:213
Zhang X, Evanno L, Poupon E (2020) Biosynthetic routes to natural isocyanides. Eur J Org Chem 2020:1919–1929
Schäfer RJB, Monaco MR, Li M, Tirla A, Rivera-Fuentes P, Wennemers H (2019) J Am Chem Soc 141:18644–18648
Chen Y, Wu KL, Tang J, Loredo A, Clements J, Pei J, Peng Z, Gupta R, Fang X, Xiao H (2019) Addition of isocyanide-containing amino acids to the genetic code for protein labeling and activation. ACS Chem Biol 14:2793–2799
Harris NC, Sato M, Herman NA, Twigg F, Cai W, Liu J, Zhu X, Downey J, Khalaf R, Martin J, Koshino H, Zhang W (2017) Biosynthesis of isonitrile lipopeptides by conserved nonribosomal peptide synthetase gene clusters in Actinobacteria. Proc Natl Acad Sci USA 114:7025–7030
Chen T-Y, Chen J, Tang Y, Zhou J, Guo Y, Chang W-c (2021) Current understanding toward isonitrile group biosynthesis and mechanism. Chin J Chem 39:463–472
Dose B, Niehs SP, Scherlach K, Shahda S, Flórez LV, Kaltenpoth M, Hertweck C (2021) Biosynthesis of sinapigladioside, an antifungal isothiocyanate from Burkholderia symbionts. ChemBioChem. https://doi.org/10.1002/cbic.202100089
Harris NC, Born DA, Cai W, Huang Y, Martin J, Khalaf R, Drennan CL, Zhang W (2018) Isonitrile formation by a non-heme iron(II)-dependent oxidase/decarboxylase. Angew Chem Int Ed 57:9707–9710
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28:235–242
Chen T-Y, Chen J, Tang Y, Zhou J, Guo Y, Chang Wc (2020) Pathway from N-alkylglycine to alkylisonitrile catalyzed by iron(II) and 2-oxoglutarate-dependent oxygenases. Angew Chem Int Ed 59:7367–7371
Li H, Liu Y (2020) Mechanistic investigation of isonitrile formation catalyzed by the nonheme iron/α-KG-dependent decarboxylase (ScoE). ACS Catal 10:2942–2957
Quesne MG, Borowski T, de Visser SP (2016) Quantum mechanics/molecular mechanics modelling of enzymatic processes: caveats and breakthroughs. Chem Eur J 22:2562–2581
Pickl M, Kurakin S, Cantú Reinhard FG, Schmid P, Pöcheim A, Winkler CK, Kroutil W, de Visser SP, Faber K (2019) Mechanistic studies of fatty acid activation by CYP152 peroxygenases reveal unexpected desaturase activity. ACS Catal 9:565–577
Mubarak MQE, Gérard EF, Blanford CF, Hay S, de Visser SP (2020) How do vanadium chloroperoxidases generate hypochlorite from hydrogen peroxide and chloride? A computational study. ACS Catal 10:14067–14079
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera–a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Ghafoor S, Mansha A, de Visser SP (2019) Selective hydrogen atom abstraction from dihydroflavonol by a nonheme iron center is the key step in the enzymatic flavonol synthesis and avoids byproducts. J Am Chem Soc 141:20278–20292
Louka S, Barry SM, Heyes DJ, Mubarak MQE, Ali HS, Alkhalaf LM, Munro AW, Scrutton NS, Challis GL, de Visser SP (2020) The catalytic mechanism of aromatic nitration by cytochrome P450 TxtE: involvement of a ferric-peroxynitrite intermediate. J Am Chem Soc 142:15764–15779
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Montgomery JA Jr, Peralta JE, Ogliaro F, Bearpark M, Heyd JJ, Brothers E, Kudin KN, Staroverov VN, Kobayashi R, Normand R, Raghavachari K, Rendell A, Burant JC, Iyengar SS, Tomasi J, Cossi M, Rega N, Millam JM, Klene M, Knox JE, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Martin RL, Morokuma K, Zakrzewski VG, Voth GA, Salvador P, Dannenberg JJ, Dapprich S, Daniels AD, Farkas Ö, Foresman JB, Ortiz JV, Cioslowski J, Fox DJ (2009) Gaussian 09, Revision D.01. Gaussian Inc, Wallingford
Ali HS, Henchman RH, de Visser SP (2021) What determines the selectivity of arginine dihydroxylation by the nonheme iron enzyme OrfP? Chem Eur J 27:1795–1809
Han SB, Ali HS, de Visser SP (2021) Glutarate hydroxylation by the carbon starvation-induced protein D. A computational study into the stereo- and regioselectivity of the reaction. Inorg Chem 60:4800–4815
Becke AD (1993) Density-functional thermochemistry. III. The role of exact exchange. J Chem Phys 98:5648–5652
Lee C, Yang W, Parr RG (1988) Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys Rev B 37:785–789
Hay PJ, Wadt WR (1985) Ab Initio effective core potentials for molecular calculations. Potentials for the transition metal atoms Sc to Hg. J Chem Phys 82:270–283
Hehre WJ, Ditchfield R, Pople JA (1972) Self-consistent molecular-orbital methods. IX. An extended Gaussian-type basis for molecular-orbital studies of organic molecules. J Chem Phys 56:2257–2262
Tomasi J, Mennucci B, Cammi R (2005) Quantum mechanical continuum solvation models. Chem Rev 105:2999–3093
Grimme S, Antony J, Ehrlich S, Krieg H (2010) A consistent and accurate ab initio parametrization of density functional dispersion correction (DFT-D) for the 94 elements H-Pu. J Chem Phys 132:154104
Faponle AS, Seebeck FP, de Visser SP (2017) Sulfoxide synthase versus cysteine dioxygenase reactivity in a nonheme iron enzyme. J Am Chem Soc 139:9259–9270
Lin YT, Stańczak A, Manchev Y, Straganz GD, de Visser SP (2020) Can a mononuclear iron(III)-superoxo active site catalyze the decarboxylation of dodecanoic acid in UndA to produce biofuels? Chem Eur J 26:2233–2242
Lin YT, Ali HS, de Visser SP (2021) Electrostatic perturbations from the protein affect C-H bond strengths of the substrate and enable negative catalysis in the TmpA biosynthesis enzyme. Chem Eur J. https://doi.org/10.1002/chem.202100791 (in press)
Ali HS, Henchman RH, de Visser SP (2020) Lignin biodegradation by a cytochrome P450 enzyme: a computational study into syringol activation by GcoA. Chem Eur J 26:13093–13102
Ali HS, Henchman RH, Warwicker J, de Visser SP (2021) How do electrostatic perturbations of the protein affect the bifurcation pathways of substrate hydroxylation versus desaturation in the nonheme iron-dependent viomycin biosynthesis enzyme? J Phys Chem A 125:1720–1737
Faponle AS, Quesne MG, de Visser SP (2016) Origin of the regioselective fatty acid hydroxylation versus decarboxylation by a cytochrome P450 peroxygenase: what drives the reaction to biofuel production? Chem Eur J 22:5478–5483
Chowdhury AS, Ali HS, Faponle AS, de Visser SP (2020) How external perturbations affect the chemoselectivity of substrate activation by cytochrome P450 OleTJE. Phys Chem Chem Phys 22:27178–27190
Shaik S, Kumar D, de Visser SP (2008) A valence bond modeling of trends in hydrogen abstraction barriers and transition states of hydroxylation reactions catalyzed by cytochrome P450 enzymes. J Am Chem Soc 130:10128–10140
Latifi R, Bagherzadeh M, de Visser SP (2009) Origin of the correlation of the rate constant of substrate hydroxylation by nonheme iron(IV)-oxo complexes with the bond-dissociation energy of the C-H bond of the substrate. Chem Eur J 15:6651–6662
de Visser SP, Kumar D, Cohen S, Shacham R, Shaik S (2004) A predictive pattern of computed barriers for C-H hydroxylation by compound I of cytochrome P450. J Am Chem Soc 126:8362–8363
Kumar D, Latifi R, Kumar S, Rybak-Akimova EV, Sainna MA, de Visser SP (2013) Rationalization of the barrier height for para-Z-styrene epoxidation by iron(IV)-oxo porphyrins with variable axial ligands. Inorg Chem 52:7968–7979
de Visser SP, Tan LS (2008) Is the bound substrate in nitric oxide synthase protonated or neutral and what is the active oxidant that performs substrate hydroxylation? J Am Chem Soc 130:12961–12974
de Visser SP (2006) Propene activation by the oxo-iron active species of taurine/α-ketoglutarate dioxygenase (TauD) enzyme. How does the catalysis compare to heme-enzymes? J Am Chem Soc 128:9813–9824
Decker A, Rohde J-U, Klinker EJ, Wong SD, Que L Jr, Solomon EI (2007) Spectroscopic and quantum chemical studies on low-spin FeIV=O complexes: Fe-O bonding and its contributions to reactivity. J Am Chem Soc 129:15983–15996
Hirao H, Li F, Que L Jr, Morokuma K (2011) Theoretical study of the mechanism of oxoiron (IV) formation from H2O2 and a nonheme iron (II) complex: O−O cleavage involving proton-coupled electron transfer. Inorg Chem 50:6637–6648
Ye S, Geng CY, Shaik S, Neese F (2013) Electronic structure analysis of multistate reactivity in transition metal catalyzed reactions: the case of C−H bond activation by non-heme iron(IV)−oxo cores. Phys Chem Chem Phys 15:8017–8030
Hirao H, Kumar D, Que L Jr, Shaik S (2006) Two-state reactivity in alkane hydroxylation by non-heme iron-oxo complexes. J Am Chem Soc 128:8590–8606
Yeh S, Neese F (2011) Nonheme oxo-iron (IV) intermediates form an oxyl radical upon approaching the C-H bond activation transition state. Proc Natl Acad Sci 108:1228–1233
de Visser SP (2006) What factors influence the ratio of C-H hydroxylation versus C=C epoxidation by a nonheme cytochrome P450 biomimetic? J Am Chem Soc 128:15809–15818
Ogliaro F, Harris N, Cohen S, Filatov M, de Visser SP, Shaik S (2000) A model “rebound” mechanism of hydroxylation by cytochrome P450: stepwise and effectively concerted pathways, and their reactivity patterns. J Am Chem Soc 122:8977–8989
Kamachi T, Yoshizawa K (2003) A theoretical study on the mechanism of camphor hydroxylation by compound I of cytochrome P450. J Am Chem Soc 125:4652–4661
Kumar D, de Visser SP, Shaik S (2004) Oxygen economy of cytochrome P450: what is the origin of the mixed functionality as a dehydrogenase–oxidase enzyme compared with its normal function? J Am Chem Soc 126:5072–5073
Kumar D, Tahsini L, de Visser SP, Kang HY, Kim SJ, Nam W (2009) The effect of porphyrin ligands on the regioselective dehydrogenation versus epoxidation of olefins by oxoiron(IV) mimics of cytochrome P450. J Phys Chem A 113:11713–11722
Ji L, Faponle AS, Quesne MG, Sainna MA, Zhang J, Franke A, Kumar D, van Eldik R, Liu W, de Visser SP (2015) Drug metabolism by cytochrome P450 enzymes: What distinguishes the pathways leading to substrate hydroxylation over desaturation? Chem Eur J 21:9083–9092
Shaik S, Cohen S, de Visser SP, Sharma PK, Kumar D, Kozuch S, Ogliaro F, Danovich D (2004) The “rebound controversy”: an overview and theoretical modeling of the rebound step in C-H hydroxylation by cytochrome P450. Eur J Inorg Chem 2004:207–226
Cummins DC, Alvarado JG, Zaragoza JPT, Mubarak MQE, Lin YT, de Visser SP, Goldberg DP (2020) Hydroxyl transfer to carbon radicals by Mn(OH) versus Fe(OH) corrole complexes. Inorg Chem 59:16053–16064
Shaik S, Kumar D, de Visser SP, Altun A, Thiel W (2005) Theoretical perspective on the structure and mechanism of cytochrome P450 enzymes. Chem Rev 105:2279–2328
Hirao H, Que L Jr, Nam W, Shaik S (2008) A Two-state reactivity rationale for counterintuitive axial ligand effects on the C-H activation reactivity of nonheme FeIV=O oxidants. Chem Eur J 14:1740–1756
Sorokin AB (2013) Phthalocyanine metal complexes in catalysis. Chem Rev 113:8152–8191
Huang X, Groves JT (2018) Oxygen activation and radical transformations in heme proteins and metalloporphyrins. Chem Rev 118:2491–2553
Friedrich LE (1983) The two hydrogen-oxygen bond-dissociation energies of hydroquinone. J Org Chem 48:3851–3852
Bordwell FG, Cheng JP (1991) Substituent effects on the stabilities of phenoxyl radicals and the acidities of phenoxyl radical. J Am Chem Soc 113:1736–1743
Mayer JM (1998) Hydrogen atom abstraction by metal−oxo complexes: understanding the analogy with organic radical reactions. Acc Chem Res 31:441–450
de Visser SP (2020) Second-coordination sphere effects on selectivity and specificity of heme and nonheme iron enzymes. Chem Eur J 26:5308–5327
Timmins A, Fowler NJ, Warwicker J, Straganz GD, de Visser SP (2018) Does substrate positioning affect the selectivity and reactivity in the hectochlorin biosynthesis halogenase?”. Front Chem 6:513. https://doi.org/10.3389/fchem.2018.00513
Ali HS, Henchman RH, de Visser SP (2021) Mechanism of oxidative ring-closure as part of the hygromycin biosynthesis step by a nonheme iron dioxygenase. ChemCatChem. https://doi.org/10.1002/cctc.202100393 (in press)
Shaik S, de Visser SP, Kumar D (2004) External electric field will control the selectivity of enzymatic-like bond activations. J Am Chem Soc 126:11746–11749
Stuyver T, De Proft F, Geerlings P, Shaik S (2020) How do local reactivity descriptors shape the potential energy surface associated with chemical reactions? The valence bond delocalization perspective. J Am Chem Soc 142:10102–10113
de Visser SP, Lin YT, Ali HS, Bagha UK, Mukherjee G, Sastri CV (2021) Negative catalysis or non-Bell-Evans-Polanyi reactivity by metalloenzymes: examples from mononuclear heme and non-heme iron oxygenases. Coord Chem Rev 439:213914
Jonnalagadda R, Del Rio FA, Cai W, Mehmood R, Narayanamoorthy M, Ren C, Zaragoza JPJ, Kulik HJ, Zhang W, Drennan CL (2021) Biochemical and crystallographic investigations into isonitrile formation by a nonheme iron-dependent oxidase/decarboxylase. J Biol Chem 296:100231
Funding
The Punjab Education Endowment Fund (PEEF) is acknowledged for a PhD studentship to HAS. SG thanks the Higher Education Commission of Pakistan for a travel grant.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ali, H.S., Ghafoor, S. & de Visser, S.P. Density Functional Theory Study into the Reaction Mechanism of Isonitrile Biosynthesis by the Nonheme Iron Enzyme ScoE. Top Catal 65, 528–543 (2022). https://doi.org/10.1007/s11244-021-01460-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11244-021-01460-x