Abstract
Rapid and sensitive detection of biomarkers enables monitoring patients’ health status and can enhance the early diagnosis of deadly diseases. In this work, we have developed a new colorimetric platform based on spherical nucleic acid (SNA) and G-quadruplex DNAzymes for the identification of specific miRNAs. The simple hybridization between the target miRNA and two capture probes (capture probe 1 located at AuNP surface and free capture probe 2) is the working principle of this biosensor. The hybridization and duplex formation among probes and miRNAs led to a significant decrease in the intensity of color change. A linear relationship between the decrease of colorimetric signal and the amount of target molecules was witnessed from 1 to 100 nM for miRNA-155. Using this method, we were able to detect concentrations of miRNA-155 as low as 0.7 nM. Furthermore, the proposed sensing platform can be utilized profitably to detect miRNA-155 in real human serum samples. We further investigated the applicability of the proposed method in a microfluidic system which displayed promising results.
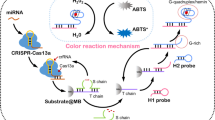
Graphical abstract
In this project, A G-quadruplex based SNAzyme was constructed to provide a fast and simple colorimetric method for miRNA detection. The SNAzyme actually employed as both target recognition element and catalytic nano labels for colorimetric detection.








Similar content being viewed by others
References
Borghei YS, Hosseini M, Dadmehr M, Hosseinkhani S, Ganjali MR, Sheikhnejad R (2016) Visual detection of cancer cells by colorimetric aptasensor based on aggregation of gold nanoparticles induced by DNA hybridization. Anal Chim Acta 904:92–97. https://doi.org/10.1016/j.aca.2015.11.026
Condrat CE, Thompson DC, Barbu MG, Bugnar OL, Boboc A, Cretoiu D, Suciu N, Cretoiu SM, Voinea SC (2020) miRNAs as biomarkers in disease: latest findings regarding their role in diagnosis and prognosis. Cells 9(2):276. https://doi.org/10.3390/cells9020276
Wang H, Peng R, Wang J, Qin Z, Xue L (2018) Circulating microRNAs as potential cancer biomarkers: the advantage and disadvantage. Clin Epigenetics 10:59. https://doi.org/10.1186/s13148-018-0492-1
Ying L, Du L, Zou R, Shi L, Zhang N, Jin J, Xu C, Zhang F, Zhu C, Wu J, Chen K, Huang M, Wu Y, Zhang Y, Zheng W, Pan X, Chen B, Lin A, Tam JKC, van Dam RM, Lai DTM, Chia KS, Zhou L, Too HP, Yu H, Mao W, Su D (2020) Development of a serum miRNA panel for detection of early stage non-small cell lung cancer. Proc Natl Acad Sci U S A 117(40):25036–25042. https://doi.org/10.1073/pnas.2006212117
Shahsavar K, Shokri E, Hosseini M (2020) A fluorescence-readout method for miRNA-155 detection with double-hairpin molecular beacon based on quadruplex DNA structure. Microchem J 158:105277. https://doi.org/10.1016/j.microc.2020.105277
Nemati F, Hosseini M (2021) A ratiometric fluorescence and colorimetric dual-mode assay for miRNA-155 based on Ce-decorated boron nitride nanosheets. Microchem J 168:106346. https://doi.org/10.1016/j.microc.2021.106346
Bian F, Sun L, Cai L, Wang Y, Zhao Y, Wang S, Zhou M (2019) Molybdenum disulfide-integrated photonic barcodes for tumor markers screening. Biosens Bioelectron 133:199–204. https://doi.org/10.1016/j.bios.2019.02.066
Wang L, Liu Z-J, Cao H-X, Liang G-X (2021) Ultrasensitive colorimetric miRNA detection based on magnetic 3D DNA walker and unmodified AuNPs. Sens Actuators, B Chem 337:129813. https://doi.org/10.1016/j.snb.2021.129813
Lee J, Kim Y-k, Lee S, Yoon S, Kim W-k (2019) Graphene oxide-based NET strategy for enhanced colorimetric sensing of miRNA. Sens Actuators, B Chem 282:861–867. https://doi.org/10.1016/j.snb.2018.11.149
Kong W, He L, Richards EJ, Challa S, Xu CX, Permuth-Wey J, Lancaster JM, Coppola D, Sellers TA, Djeu JY, Cheng JQ (2014) Upregulation of miRNA-155 promotes tumour angiogenesis by targeting VHL and is associated with poor prognosis and triple-negative breast cancer. Oncogene 33(6):679–689. https://doi.org/10.1038/onc.2012.636
Gurses HE, Hatipoğlu OF, Gunduz M, Gunduz E (2015) MicroRNAs as therapeutic targets in human breast cancer. In: Gunduz M (ed) A concise review of molecular pathology of breast cancer, Chapter 5. InTech, Croatia, pp 121–137. https://doi.org/10.5772/59681
Gillespie P, Ladame S, O’Hare D (2018) Molecular methods in electrochemical microRNA detection. Analyst 144(1):114–129. https://doi.org/10.1039/c8an01572d
Ye J, Xu M, Tian X, Cai S, Zeng S (2019) Research advances in the detection of miRNA. J Pharm Anal 9(4):217–226. https://doi.org/10.1016/j.jpha.2019.05.004
Wan Z, Umer M, Lobino M, Thiel D, Nguyen N-T, Trinchi A, Shiddiky MJA, Gao Y, Li Q (2020) Laser induced self-N-doped porous graphene as an electrochemical biosensor for femtomolar miRNA detection. Carbon 163:385–394. https://doi.org/10.1016/j.carbon.2020.03.043
Huang R, Liao Y, Zhou X, Fu Y, Xing D (2017) Multiplexed detection of microRNA biomarkers from tumor cells and tissues with a homogeneous nano-photon switch. Sens Actuators, B Chem 247:505–513. https://doi.org/10.1016/j.snb.2017.03.055
Fiammengo R (2017) Can nanotechnology improve cancer diagnosis through miRNA detection? Biomark Med 11(1):69–86. https://doi.org/10.2217/bmm-2016-0195
Cutler JI, Auyeung E, Mirkin CA (2012) Spherical nucleic acids. J Am Chem Soc 134(3):1376–1391. https://doi.org/10.1021/ja209351u
Sun Y, Shi L, Wang Q, Mi L, Li T (2019) Spherical nucleic acid enzyme (SNAzyme) boosted chemiluminescence miRNA imaging using a smartphone. Anal Chem 91(5):3652–3658. https://doi.org/10.1021/acs.analchem.8b05696
Sun Y, Wang Q, Mi L, Shi L, Li T (2019) Target-induced payload amplification for spherical nucleic acid enzyme (SNAzyme)-catalyzed electrochemiluminescence detection of circulating microRNAs. Anal Chem 91(20):12948–12953. https://doi.org/10.1021/acs.analchem.9b03001
Mirkin CA, Letsinger RL, Mucic RC, Storhoff JJ (1996) A DNA-based method for rationally assembling nanoparticles into macroscopic materials. Nature 382(6592):607–609. https://doi.org/10.1038/382607a0
Li Z, Jin R, Mirkin CA, Letsinger RL (2002) Multiple thiol-anchor capped DNA-gold nanoparticle conjugates. Nucleic Acids Res 30(7):1558–1562. https://doi.org/10.1093/nar/30.7.1558
Sato K, Hosokawa K, Maeda M (2003) Rapid aggregation of gold nanoparticles induced by non-cross-linking DNA hybridization. J Am Chem Soc 125(27):8102–8103. https://doi.org/10.1021/ja034876s
Karami A, Hasani M, Azizi Jalilian F, Ezati R (2021) Conventional PCR assisted single-component assembly of spherical nucleic acids for simple colorimetric detection of SARS-CoV-2. Sens Actuators B Chem 328:128971. https://doi.org/10.1016/j.snb.2020.128971
Frens G (1973) Controlled nucleation for the regulation of the particle size in monodisperse gold suspensions. Nat Phys Sci 241(105):20–22. https://doi.org/10.1038/physci241020a0
Shahsavar K, Hosseini M, Shokri E, Ganjali MR, Ju H (2017) A sensitive colorimetric aptasensor with a triple-helix molecular switch based on peroxidase-like activity of a DNAzyme for ATP detection. Anal Methods 9(32):4726–4731. https://doi.org/10.1039/c7ay01381g
Naderi M, Hosseini M, Ganjali MR (2018) Naked-eye detection of potassium ions in a novel gold nanoparticle aggregation-based aptasensor. Spectrochim Acta A Mol Biomol Spectrosc 195:75–83. https://doi.org/10.1016/j.saa.2018.01.051
Shahsavar K, Hosseini M, Shokri E, Xu G (2021) New insight into G-quadruplexes; diagnosis application in cancer. Anal Biochem 620:114149. https://doi.org/10.1016/j.ab.2021.114149
Dehghani Z, Hosseini M, Mohammadnejad J, Bakhshi B, Rezayan AH (2018) Colorimetric aptasensor for Campylobacter jejuni cells by exploiting the peroxidase like activity of Au@Pd nanoparticles. Mikrochim Acta 185(10):448. https://doi.org/10.1007/s00604-018-2976-2
Hosseini M, Khabbaz H, Dadmehr M, Ganjali MR, Mohamadnejad J (2015) Aptamer-based colorimetric and chemiluminescence detection of aflatoxin B1 in foods samples. Acta Chim Slov 62(3):721–728. https://doi.org/10.17344/acsi.2015.1358
Liu J, Lu Y (2006) Preparation of aptamer-linked gold nanoparticle purple aggregates for colorimetric sensing of analytes. Nat Protoc 1(1):246–252. https://doi.org/10.1038/nprot.2006.38
Liu B, Liu J (2017) Freezing directed construction of bio/nano interfaces: reagentless conjugation, denser spherical nucleic acids, and better nanoflares. J Am Chem Soc 139(28):9471–9474. https://doi.org/10.1021/jacs.7b04885
Lytton-Jean AK, Mirkin CA (2005) A thermodynamic investigation into the binding properties of DNA functionalized gold nanoparticle probes and molecular fluorophore probes. J Am Chem Soc 127(37):12754–12755. https://doi.org/10.1021/ja052255o
Shah P, Zhu X, Li CZ (2013) Development of paper-based analytical kit for point-of-care testing. Expert Rev Mol Diagn 13(1):83–91. https://doi.org/10.1586/erm.12.130
Acknowledgements
The authors are grateful to the Research Council of University of Tehran for the financial support of this work.
Author information
Authors and Affiliations
Contributions
Kosar Shahsavar: conceptualization, methodology, writing—original draft; Morteza Hosseini: resources, supervision, funding acquisition, and review and editing; Ehsan Shokri: validation, writing—review and editing.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shahsavar, K., Shokri, E. & Hosseini, M. Sensitive colorimetric detection of miRNA-155 via G-quadruplex DNAzyme decorated spherical nucleic acid. Microchim Acta 189, 357 (2022). https://doi.org/10.1007/s00604-022-05455-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-022-05455-7