Abstract
Background
Seven patients with chronic hepatitis C virus (HCV) infection were followed up for 10 years in order to analyze the molecular evolution of the HCV nonstructural 5A (NS5A) gene.
Methods
Serum samples were obtained from seven patients with anti-HCV who were HCV-RNA positive. The HCV NS5A region, including the interferon sensitivity-determining region (ISDR), protein kinase R binding domain (PKR-BD), and V3 was amplified and cloned, and then sequenced. The nucleotide sequences of the region at the beginning and at the end of the 10 years were aligned with the CLUSTAL W program, version 1.8, and the corresponding amino-acid sequences were deduced.
Results
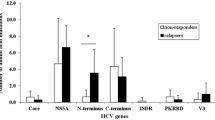
The three serine residues at positions 2197, 2201, and 2204, suggested to be important for hyperphosphorylation of NS5A, were highly conserved in different patients and were within the quasispecies of each patient. The wild-type ISDR or minimally mutated strain was dominant in all the patients and the number of quasispecies decreased over time.
Conclusions
Our findings showed that the functional domains of NS5A were generally conserved over an interval of 10 years. Quasispecies distribution was variable and distinctly clustered over time during the natural course of HCV infection. These results may contribute to better understanding of HCV natural infection.
Similar content being viewed by others
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Fan, W., Zhu, W., Wei, L. et al. Nonstructural 5A gene variability of hepatitis C virus (HCV) during a 10-year follow up. J Gastroenterol 40, 43–51 (2005). https://doi.org/10.1007/s00535-004-1446-2
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00535-004-1446-2