Abstract
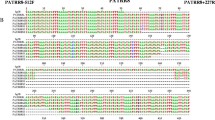
Although approximately 1 in 500 individuals carries a reciprocal translocation, little is known about the mechanisms that result in their formation. We analyzed the sequences surrounding the breakpoints in three unbalanced translocations of 1p and 9q, all of which were designated t(1;9)(p36.3;q34), to investigate the presence of sequence motifs that might mediate nonhomologous end joining (NHEJ). The breakpoint regions were unique in all individuals. Two of three translocations demonstrated insertions and duplications at the junctions, suggesting NHEJ in the formation of the rearrangements. No homology was identified in the breakpoint regions, further supporting NHEJ. We found translin motifs at the breakpoint junctions, suggesting the involvement of translin in the joining of the broken chromosome ends. We propose a model for balanced translocation formation in humans similar to transposition in bacteria, in which staggered nicks are repaired resulting in duplications and insertions at the translocation breakpoints.



Similar content being viewed by others
References
Abeysinghe SS, Chuzhanova N, Krawczak M, Ball EV, Cooper DN (2003) Translocation and gross deletion breakpoints in human inherited disease and cancer I: nucleotide composition and recombination-associated motifs. Hum Mutat 22:229–244
Aoki K, Suzuki K, Sugano T, Tasaka T, Nakahara K, Kuge O, Omori A, Kasai M (1995) A novel gene, translin, encodes a recombination hotspot binding protein associated with chromosomal translocations. Nat Genet 10:167–74
Bacolla A, Jaworski A, Larson JE, Jakupciak JP, Chuzhanova N, Abeysinghe SS, O’Connell CD, Cooper DN, Wells RD (2004) Breakpoints of gross deletions coincide with non-B DNA conformations. Proc Natl Acad Sci USA 101:14162–14167
Bacolla A, Wells RD (2004) Non-B DNA conformations, genomic rearrangements, and human disease. J Biol Chem 279:47411–47414
Badge RM, Yardley J, Jeffreys AJ, Armour JA (2000) Crossover breakpoint mapping identifies a subtelomeric hotspot for male meiotic recombination. Hum Mol Genet 9:1239–1244
Ballif BC, Gajecka M, Shaffer LG (2004a) Monosomy 1p36 breakpoints indicate repetitive DNA sequence elements may be involved in generating and/or stabilizing some terminal deletions. Chromosome Res 12:133–141
Ballif BC, Kashork CD, Shaffer LG (2000) FISHing for mechanisms of cytogenetically defined terminal deletions using chromosome-specific subtelomeric probes. Eur J Hum Genet 8:764–770
Ballif BC, Wakui K, Gajecka M, Shaffer LG (2004b) Translocation breakpoint mapping and sequence analysis in three monosomy 1p36 subjects with der(1)t(1;1)(p36;q44) suggest mechanisms for telomere capture in stabilizing de novo terminal rearrangements. Hum Genet 114:198–206
Ballif BC, Yu W, Shaw CA, Kashork CD, Shaffer LG (2003) Monosomy 1p36 breakpoint junctions suggest pre-meiotic breakage-fusion-bridge cycles are involved in generating terminal deletions. Hum Mol Genet 12:2153–2165
Benson G (1999) Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res 27:573–580
Bodrug SE, Holden JJ, Ray PN, Worton RG (1991) Molecular analysis of X-autosome translocations in females with Duchenne muscular dystrophy. Embo J 10:3931–3939
Bodrug SE, Ray PN, Gonzalez IL, Schmickel RD, Sylvester JE, Worton RG (1987) Molecular analysis of a constitutional X-autosome translocation in a female with muscular dystrophy. Science 237:1620–1624
Chalk JG, Barr FG, Mitchell CD (1997) Translin recognition site sequences flank chromosome translocation breakpoints in alveolar rhabdomyosarcoma cell lines. Oncogene 15:1199–1205
Champ PC, Maurice S, Vargason JM, Camp T, Ho PS (2004) Distributions of Z-DNA and nuclear factor I in human chromosome 22: a model for coupled transcriptional regulation. Nucleic Acids Res 32:6501–6510
Chennathukuzhi VM, Kurihara Y, Bray JD, Hecht NB (2001) Trax (translin-associated factor X), a primarily cytoplasmic protein, inhibits the binding of TB-RBP (translin) to RNA. J Biol Chem 276:13256–13263
Cho YS, Iguchi N, Yang J, Handel MA, Hecht NB (2005) Meiotic mRNA and non-coding RNA targets of the RNA-binding protein translin (TSN) in mouse testis. Biol Reprod
Chuzhanova N, Abeysinghe SS, Krawczak M, Cooper DN (2003) Translocation and gross deletion breakpoints in human inherited disease and cancer II: Potential involvement of repetitive sequence elements in secondary structure formation between DNA ends. Hum Mutat 22:245–251
Collins A, Frezal J, Teague J, Morton NE (1996) A metric map of humans: 23,500 loci in 850 bands. Proc Natl Acad Sci USA 93:14771–14775
Erdemir T, Bilican B, Oncel D, Goding CR, Yavuzer U (2002) DNA damage-dependent interaction of the nuclear matrix protein C1D with translin-associated factor X (TRAX). J Cell Sci 115:207–216
Gajecka M, Glotzbach CD, Jarmuz M, Ballif BC, Shaffer LG (2006) Identification of cryptic imbalance in phenotypically normal and abnormal translocation carriers. Eur J Hum Genet (in press)
Haider S, Matsumoto R, Kurosawa N, Wakui K, Fukushima Y, Isobe M (2006) Molecular characterization of a novel translocation t(5;14)(q21;q32) in a patient with congenital abnormalities. J Hum Genet 51:335–340
Heilstedt HA, Ballif BC, Howard LA, Lewis RA, Stal S, Kashork CD, Bacino CA, Shapira SK, Shaffer LG (2003) Physical map of 1p36, placement of breakpoints in monosomy 1p36, and clinical characterization of the syndrome. Am J Hum Genet 72:1200–1212
Ho PS, Ellison MJ, Quigley GJ, Rich A (1986) A computer aided thermodynamic approach for predicting the formation of Z-DNA in naturally occurring sequences. Embo J 5:2737–2744
Hosaka T, Kanoe H, Nakayama T, Murakami H, Yamamoto H, Nakamata T, Tsuboyama T, Oka M, Kasai M, Sasaki MS, Nakamura T, Toguchida J (2000) Translin binds to the sequences adjacent to the breakpoints of the TLS and CHOP genes in liposarcomas with translocation t(12;6). Oncogene 19:5821–5825
Inoue K, Osaka H, Thurston VC, Clarke JT, Yoneyama A, Rosenbarker L, Bird TD, Hodes ME, Shaffer LG, Lupski JR (2002) Genomic rearrangements resulting in PLP1 deletion occur by nonhomologous end joining and cause different dysmyelinating phenotypes in males and females. Am J Hum Genet 71:838–853
Ishida R, Okado H, Sato H, Shionoiri C, Aoki K, Kasai M (2002) A role for the octameric ring protein, translin, in mitotic cell division. FEBS Lett 525:105–110
Jacob E, Pucshansky L, Zeruya E, Baran N, Manor H (2004) The human protein translin specifically binds single-stranded microsatellite repeats, d(GT)n, and G-strand telomeric repeats, d(TTAGGG)n: a study of the binding parameters. J Mol Biol 344:939–950
Jeggo PA (1998) DNA breakage and repair. Adv Genet 38:185–218
Jurka J (2000) Repbase update: a database and an electronic journal of repetitive elements. Trends Genet 16:418–420
Kanoe H, Nakayama T, Hosaka T, Murakami H, Yamamoto H, Nakashima Y, Tsuboyama T, Nakamura T, Ron D, Sasaki MS, Toguchida J (1999) Characteristics of genomic breakpoints in TLS-CHOP translocations in liposarcomas suggest the involvement of translin and topoisomerase II in the process of translocation. Oncogene 18:721–729
Kasai M, Matsuzaki T, Katayanagi K, Omori A, Maziarz RT, Strominger JL, Aoki K, Suzuki K (1997) The translin ring specifically recognizes DNA ends at recombination hot spots in the human genome. J Biol Chem 272:11402–11407
Liang F, Han M, Romanienko PJ, Jasin M (1998) Homology-directed repair is a major double-strand break repair pathway in mammalian cells. Proc Natl Acad Sci USA 95:5172–5177
Mansouri MR, Carlsson B, Davey E, Nordenskjold A, Wester T, Anneren G, Lackgren G, Dahl N (2006) Molecular genetic analysis of a de novo balanced translocation t(6;17)(p21.31;q11.2) associated with hypospadias and anorectal malformation. Hum Genet 119:162–168
McMullan TW, Crolla JA, Gregory SG, Carter NP, Cooper RA, Howell GR, Robinson DO (2002) A candidate gene for congenital bilateral isolated ptosis identified by molecular analysis of a de novo balanced translocation. Hum Genet 110:244–250
Millar JK, Wilson-Annan JC, Anderson S, Christie S, Taylor MS, Semple CA, Devon RS, Clair DM, Muir WJ, Blackwood DH, Porteous DJ (2000) Disruption of two novel genes by a translocation co-segregating with Schizophrenia. Hum Mol Genet 9:1415–1423
Moore JK, Haber JE (1996) Cell cycle and genetic requirements of two pathways of nonhomologous end-joining repair of double-strand breaks in Saccharomyces cerevisiae. Mol Cell Biol 16:2164–2173
Paces J, Zika R, Paces V, Pavlicek A, Clay O, Bernardi G (2004) Representing GC variation along eukaryotic chromosomes. Gene 333:135–141
Rice P, Longden I, Bleasby A (2000) EMBOSS: the European molecular biology open software suite. Trends Genet 16:276–277
Saccone S, De Sario A, Della Valle G, Bernardi G (1992) The highest gene concentrations in the human genome are in telomeric bands of metaphase chromosomes. Proc Natl Acad Sci USA 89:4913–4917
Schule B, Albalwi M, Northrop E, Francis DI, Rowell M, Slater HR, Gardner RJ, Francke U (2005) Molecular breakpoint cloning and gene expression studies of a novel translocation t(4;15)(q27;q11.2) associated with Prader-Willi syndrome. BMC Med Genet 6:18
Shapiro JA (1979) Molecular model for the transposition and replication of bacteriophage Mu and other transposable elements. Proc Natl Acad Sci USA 76:1933–1937
Shaw CJ, Lupski JR (2004) Implications of human genome architecture for rearrangement-based disorders: the genomic basis of disease. Hum Mol Genet 13 Spec No 1: R57–R64
Vinogradov AE (2003) DNA helix: the importance of being GC-rich. Nucleic Acids Res 31:1838–1844
Visser R, Shimokawa O, Harada N, Kinoshita A, Ohta T, Niikawa N, Matsumoto N (2005) Identification of a 3.0-kb major recombination hotspot in patients with sotos syndrome who carry a common 1.9-Mb microdeletion. Am J Hum Genet 76:52–67
Yang S, Cho YS, Chennathukuzhi VM, Underkoffler LA, Loomes K, Hecht NB (2004) Translin-associated factor X is post-transcriptionally regulated by its partner protein TB-RBP, and both are essential for normal cell proliferation. J Biol Chem 279:12605–12614
Yoshiura K, Machida J, Daack-Hirsch S, Patil SR, Ashworth LK, Hecht JT, Murray JC (1998) Characterization of a novel gene disrupted by a balanced chromosomal translocation t(2;19)(q11.2;q13.3) in a family with cleft lip and palate. Genomics 54:231–240
Yu X, Gabriel A (2004) Reciprocal translocations in Saccharomyces cerevisiae formed by nonhomologous end joining. Genetics 166:741–751
Acknowledgments
We thank A. Theisen (Washington State University, Spokane, WA, USA) for his critical editing of the manuscript, Dr. P. Shing Ho (Oregon State University, Corvallis, OR, USA) for his help with the ZHUNT program and Z-DNA prediction, and Drs. J. Lupski (Baylor College of Medicine, Houston, TX, USA) and B. Morrow (Albert Einstein College of Medicine, Bronx, NY) for helpful discussions. This work was supported in part from a National Institutes of Health PO1 (HD39420).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gajecka, M., Pavlicek, A., Glotzbach, C.D. et al. Identification of sequence motifs at the breakpoint junctions in three t(1;9)(p36.3;q34) and delineation of mechanisms involved in generating balanced translocations. Hum Genet 120, 519–526 (2006). https://doi.org/10.1007/s00439-006-0222-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-006-0222-1