Abstract
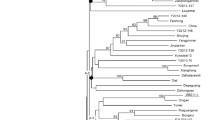
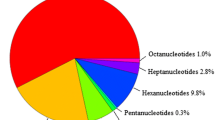
Spinach is a popular leafy green vegetable due to its nutritional composition. It contains high concentrations of vitamins A, E, C, and K, and folic acid. Development of genetic markers for spinach is important for diversity and breeding studies. In this work, Next Generation Sequencing (NGS) technology was used to develop genomic simple sequence repeat (SSR) markers. After cleaning and contig assembly, the sequence encompassed 2.5% of the 980 Mb spinach genome. The contigs were mined for SSRs. A total of 3852 SSRs were detected. Of these, 100 primer pairs were tested and 85% were found to yield clear, reproducible amplicons. These 85 markers were then applied to 48 spinach accessions from worldwide origins, resulting in 389 alleles with 89% polymorphism. The average gene diversity (GD) value of the markers (based on a GD calculation that ranges from 0 to 0.5) was 0.25. Our results demonstrated that the newly developed SSR markers are suitable for assessing genetic diversity and population structure of spinach germplasm. The markers also revealed clustering of the accessions based on geographical origin with clear separation of Far Eastern accessions which had the overall highest genetic diversity when compared with accessions from Persia, Turkey, Europe, and the USA. Thus, the SSR markers have good potential to provide valuable information for spinach breeding and germplasm management. Also they will be helpful for genome mapping and core collection establishment.



Similar content being viewed by others
References
Abuzayed M, El-Dabba N, Frary A, Doganla r S (2016) GDdom: an online tool for calculation of dominant marker gene diversity. Biochem Genet 43:1–3
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410
Arumuganathan K, Earle ED (1991) Nuclear DNA content of some important plant species. Plant Mol Biol Rep 9(3):208–218
Boswell VR (1949) Garden peas and spinach from the Middle East. Reprint of “Our Vegetable Travelers” Natl Geogr 96:2
Cardle L, Milbourne D, Macaulay M, Marshall D, Waugh R (2000) Computational and experimental characterization of physically clustered simple sequence repeats in plants. Genetics 156:847–854
Chitwood J, Shi A, Mou B, Evans M, Clark J, Motes D, Chen P, Hensley D (2016) Population structure and association analysis of bolting, plant height, and leaf erectness in spinach. HortScience 51(5):481–486
Dohm JC, Minoche AE, Holtgräwe D, Capella-Gutiérrez S, Zakrzewski F, Tafer H, Rupp O, Sörensen TR, Stracke R, Reinhardt R, Goesmann A, Kraft T, Schulz, Stadler PF, Schmidt T, Gabaldón T, Lehrach H, Weisshaar B, Himmelbauer H (2014) The genome of the recently domesticated crop plant sugar beet (Beta vulgaris). Nature 505(7484):546–549
Doyle JJ, Doyle JE (1990) Isolation of plant DNA from fresh tissue. Focus (12):13–15
Earl DA, Von Holdt BM (2012) Structure Harvester: a website and program for visualizing Structure output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Feng C, Bluhm BH, Correll JC (2015) Construction of a spinach bacterial artificial chromosome (BAC) library as a resource for gene identification and marker development. Plant Mol Biol Rep 33(6):1996–2005
Food and Agriculture Organization of the United Nati1ons, FAOSTAT (2013) http://www.fao.org/corp/statistics/en. Accessed 04 July 2016
Hu J, Mou B, Vick BA (2007) Genetic diversity of 38 spinach (Spinacia oleracea L.) germplasm accessions and 10 commercial hybrids assessed by TRAP markers. Genet Resour Crop Evol 54(8):1667–1674
Ito M, Ohmido N, Akiyama Y, Fukui K, Koba T (2000) Characterization of spinach chromosomes by condensation patterns and physical mapping of 5 S and 45 S rDNAs by FISH. J Am Soc Hortic Sci 125(1):59–62
Khattak JZK, Torp AM, Andersen SB (2006) A genetic linkage map of Spinacia oleracea and localization of a sex determination locus. Euphytica 148:311–318
Khattak JZK, Christiansen JL, Torp AM, Andersen SB (2007) Genic microsatellite markers for discrimination of spinach cultivars. Plant Breeding 126(4):454–456
Kuwahara K, Suzuki R, Ito Y, Mikami T, Onodera Y (2014) An analysis of genetic differentiation and geographical variation of spinach germplasm using SSR markers. Plant Genet Resour 12(02):185–190
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9(4):357–359
Lester GE, Makus DJ, Hodges DM, Jifon JL (2013) Summer (Subarctic) versus winter (Subtropic) production affects spinach (Spinacia oleracea L.) leaf bionutrients: Vitamins (C, E, Folate, K1, provitamin A), lutein, phenolics, and antioxidants. J Agric Food Chem 61(29):7019–7027
Ma J, Shi A, Mou B, Evans M, Clark JR, Motes D, Correll JC, Xiong H, Qin J, Chitwood J, Weng Y (2016) Association mapping of leaf traits in spinach (Spinacia oleracea L.). Plant Breeding 404:1–6
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17(1):10–12
Minoche AE, Dohm JC, Schneider J, Holtgräwe D, Viehöver P, Montfort M, Sörensen TR, Weisshaar B, Himmelbauer H (2015) Exploiting single-molecule transcript sequencing for eukaryotic gene prediction. Genome Biol 16(1):184
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70(12):3321–3323
Park YJ, Lee JK, Kim NS (2009) Simple sequence repeat polymorphisms (SSRPs) for evaluation of molecular diversity and germplasm classification of minor crops. Molecules 14(11):4546–4569
Perrier X, Jacquemoud-Collet JP (2006) DARwin software. http://darwin.cirad.fr
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Roldan-Ruiz I, Dendauw J, Bockstaele EV, Depicker A, Loose MD (2000) AFLP markers reveal high polymorphic rates in ryegrasses (Lolium spp.). Mol Breed 6:125–134
Shi A, Beiquan M, James CC (2016) Association analysis for oxalate concentration in spinach. Euphytica 212(1):17–28
Simpson JT, Wong K, Jackman SD, Schein JE, Jones SJ BL (2009) AbySS: a paralel assembler for short read sequence data. Genome Res 19(6):1117–1123
Yang XD, Tan HW, Zhu WM (2016) SpinachDB: a well-characterized genomic database for gene family classification and SNP information of spinach. PloS One 11(5):e0152706
Zalapa JE, Cuevas H, Zhu H, Steffan S, Senalik D, Zeldin E, McCown B, Harbut R, Simon P (2012) Using next-generation sequencing approaches to isolate simple sequence repeat (SSR) loci in the plant sciences. Am J Bot 99(2):193–208
Acknowledgements
This research was supported by funding from an Izmir Institute of Technology Scientific Research Project, IYTE-BAP2012-2014.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Data availability
Sequence data are available at the SRA database of NCBI (SRX2266012).
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Göl, Ş., Göktay, M., Allmer, J. et al. Newly developed SSR markers reveal genetic diversity and geographical clustering in spinach (Spinacia oleracea). Mol Genet Genomics 292, 847–855 (2017). https://doi.org/10.1007/s00438-017-1314-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-017-1314-4