Abstract
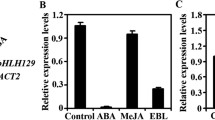
In our previous microarray analysis of NaCl-treated Arabidopsis roots, we identified a basic-helix-loop-helix (bHLH) transcription factor, bHLH92 (At5g43650), as one of the transcripts showing the greatest fold-increase in abundance upon NaCl exposure. Here, we characterize the role of bHLH92 in the context of abiotic stress physiology and hormone responses. We observed that bHLH92 transcript abundance increases in response to NaCl, dehydration, mannitol, and cold treatments, and compared these responses to those of two closely related genes: bHLH41 and bHLH42. The NaCl-inducibility of bHLH92 was only partially dependent on abscisic acid (ABA) biosynthesis and SALT OVERLY SENSITIVE2 (SOS2) pathways. As compared to WT, root elongation of bhlh92 mutants was more sensitive to mannitol, and these mutants also showed increased electrolyte leakage following NaCl treatments. Overexpression of bHLH92 moderately increased the tolerance to NaCl and osmotic stresses. Finally, we identified at least 19 putative downstream target genes of bHLH92 under NaCl treatment using an oligonucleotide microarray. Together these data show that bHLH92 functions in plant responses to osmotic stresses, although the net contribution of bHLH92-regulated genes to stress tolerance appears relatively limited in proportion to what might be expected from its transcript expression pattern.






Similar content being viewed by others
Abbreviations
- bHLH:
-
Basic helix-loop-helix
- ABA:
-
Abscisic acid
- abi:
-
ABA insensitive
- CBL:
-
Calcineurin B-like protein
- CIPK:
-
CBL-interacting protein kinase
- DAS:
-
Days after stratification
- LEA:
-
Late embryogenesis abundant
- MS:
-
Murashige and Skoog
- MV:
-
Methyl viologen
- OE:
-
Overexpression
- qRT–PCR:
-
Quantitative reverse transcription-polymerase chain reaction
- REL:
-
Relative electrolyte leakage
- ROS:
-
Reactive oxygen species
- SOS:
-
Salt overly sensitive
- TF:
-
Transcription factor
- WT:
-
Wild-type
References
Abe H, Urao T, Ito T, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 15:63–78
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK et al (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301:653–657
Anderson JP, Badruzsaufari E, Schenk PM, Manners JM, Desmond OJ, Ehlert C, Maclean DJ, Ebert PR, Kazan K (2004) Antagonistic interaction between abscisic acid and jasmonate-ethylene signaling pathways modulates defense gene expression and disease resistance in Arabidopsis. Plant Cell 16:3460–3479
Bailey PC, Martin C, Toledo-Ortiz G, Quail PH, Huq E, Heim MA, Jakoby M, Werber M, Weisshaar B (2003) Update on the basic helix-loop-helix transcription factor gene family in Arabidopsis thaliana. Plant Cell 15:2497–2502
Berglund AC, Sjolund E, Ostlund G, Sonnhammer ELL (2008) InParanoid 6: eukaryotic ortholog clusters with inparalogs. Nucleic Acids Res 36:D263–D266
Boter M, Ruiz-Rivero O, Abdeen A, Prat S (2004) Conserved MYC transcription factors play a key role in jasmonate signaling both in tomato and Arabidopsis. Genes Dev 18:1577–1591
Bray EA, Bailey-Serres J, Weretilnyk E (2000) Responses to abiotic stresses. In: Buchanan BB, Gruissem W, Jones RL (eds) Biochemistry and molecular biology of plants. American Society of Plant Physiologists, Rockville, MD, USA, pp 1158–1203
Chen WQ, Provart NJ, Glazebrook J, Katagiri F, Chang HS, Eulgem T, Mauch F, Luan S, Zou GZ, Whitham SA, Budworth PR, Tao Y, Xie ZY, Chen X, Lam S, Kreps JA, Harper JF, Si-Ammour A, Mauch-Mani B, Heinlein M, Kobayashi K, Hohn T, Dangl JL, Wang X, Zhu T (2002) Expression profile matrix of Arabidopsis transcription factor genes suggests their putative functions in response to environmental stresses. Plant Cell 14:559–574
Chinnusamy V, Ohta M, Kanrar S, Lee BH, Hong XH, Agarwal M, Zhu JK (2003) ICE1: a regulator of cold-induced transcriptome and freezing tolerance in Arabidopsis. Genes Dev 17:1043–1054
Chinnusamy V, Jagendorf A, Zhu JK (2005) Understanding and improving salt tolerance in plants. Crop Sci 45:437–448
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Czechowski T, Stitt M, Altmann T, Udvardi MK, Scheible W (2005) Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol 139:5–17
De Smet I, Zhang H, Inzé D, Beeckman T (2006) A novel role for abscisic acid emerges from underground. Trends Plant Sci 11:434–439
Deak KI, Malamy J (2005) Osmotic regulation of root system architecture. Plant J 43:17–28
Finkelstein RR, Rock CD (2002) Abscisic acid biosynthesis and response. In: Somerville CR, Meyerowitz EM (eds) The Arabidopsis book. American Society of Plant Biologists, Rockville, MD
Gilmour SJ, Artus NN, Thomashow MF (1992) cDNA sequence analysis and expression of two cold-regulated genes of Arabidopsis thaliana. Plant Mol Biol 18:13–21
Gong ZZ, Koiwa H, Cushman MA, Ray A, Bufford D, Kore-eda S, Matsumoto TK, Zhu JH, Cushman JC, Bressan RA, Hasegawa PM (2001) Genes that are uniquely stress regulated in salt overly sensitive (sos) mutants. Plant Physiol 126:363–375
Hanano S, Stracke R, Jakoby M, Merkle T, Domagalska MA, Weisshaar B, Davis SJ (2008) A systematic survey in Arabidopsis thaliana of transcription factors that modulate circadian parameters. BMC Genom 9:182
Hasegawa PM, Bressan RA, Zhu JK, Bohnert HJ (2000) Plant cellular and molecular responses to high salinity. Ann Rev Plant Physiol Plant Mol Biol 51:463–499
Heim MA, Jakoby M, Werber M, Martin C, Weisshaar B, Bailey PC (2003) The basic helix-loop-helix transcription factor family in plants: a genome-wide study of protein structure and functional diversity. Mol Biol Evol 20:735–747
Jiang Y-Q, Deyholos MK (2006) Comprehensive transcriptional profiling of NaCl-stressed Arabidopsis roots reveals novel classes of responsive genes. BMC Plant Biol 6:25
Jiang Y-Q, Deyholos MK (2009) Functional characterization of Arabidopsis NaCl-inducible WRKY25 and WRKY33 transcription factors in abiotic stress. Plant Mol Biol 69:91–105
Jiang Y, Yang B, Harris NS, Deyholos MK (2007) Comparative proteomic analysis of NaCl stress-responsive proteins in Arabidopsis roots. J Exp Bot 58:3591–3607
Kamei A, Seki M, Umezawa T, Ishida J, Satou M, Akiyama K, Zhu JK, Shinozaki K (2005) Analysis of gene expression profiles in Arabidopsis salt overly sensitive mutants sos2-1 and sos3-1. Plant Cell Environ 28:1267–1275
Kim J, Kim HY (2006) Functional analysis of a calcium-binding transcription factor involved in plant NaCl stress signaling. FEBS Lett 80:5251–5256
Kiyosue T, Yamaguchishinozaki K, Shinozaki K (1994) Characterization of two cDNAs (ERD10 and ERD14) corresponding to genes that respond rapidly to dehydration stress in Arabidopsis thaliana. Plant Cell Physiol 35:225–231
Knight H, Trewavas AJ, Knight MR (1997) Calcium signalling in Arabidopsis thaliana responding to drought and salinity. Plant J 12:1067–1078
Kreps JA, Wu YJ, Chang HS, Zhu T, Wang X, Harper JF (2002) Transcriptome changes for Arabidopsis in response to salt, osmotic, and cold stress. Plant Physiol 130:2129–2141
Kumari M, Taylor GJ, Deyholos MK (2008) Transcriptomic responses to aluminum stress in roots of Arabidopsis thaliana. Mol Genet Genomics 279:339–357
Kurkela S, Franck M (1990) Cloning and characterization of a cold- and ABA-inducible Arabidopsis gene. Plant Mol Biol 15:137–144
Lee BH, Henderson DA, Zhu JK (2005) The Arabidopsis cold-responsive transcriptome and its regulation by ICE1. Plant Cell 17:3155–3175
Li HM, Sun JQ, Xu YX, Jiang HL, Wu XY, Li CY (2007) The bHLH-type transcription factor AtAIB positively regulates ABA response in Arabidopsis. Plant Mol Biol 65:655–665
Lorenzo O, Chico JM, Sanchez-Serrano JJ, Solano R (2004) JASMONATE-INSENSITIVE1 encodes a MYC transcription factor essential to discriminate between different jasmonate regulated defense responses in Arabidopsis. Plant Cell 16:1938–1950
Ma SS, Gong QQ, Bohnert HJ (2006) Dissecting salt stress pathways. J Exp Bot 57:1097–1107
Miller G, Shulaev V, Mittler R (2008) Reactive oxygen signaling and abiotic stress. Physiol Plant 133:481–489
Montiel G, Gantet P, Jay-Allemand C, Breton C (2004) Transcription factor networks. Pathways to the knowledge of root development. Plant Physiol 136:3478–3485
Munns R (2005) Genes and salt tolerance: bringing them together. New Phytol 167:645–663
Murre C, McCaw PS, Baltimore D (1989) A new DNA binding and dimerization motif in immunoglobulin enhancer binding, daughterless, MyoD and myc proteins. Cell 56:777–783
Nesi N, Debeaujon I, Jond C, Pelletier G, Caboche M, Lepiniec L (2000) The TT8 gene encodes a basic helix-loop-helix domain protein required for expression of DFR and BAN genes in Arabidopsis siliques. Plant Cell 12:1863–1878
Ni M, Tepperman JM, Quail PH (1998) PIF3, a phytochrome-interacting factor necessary for normal photoinduced signal transduction, is a novel basic helix-loop-helix protein. Cell 95:657–667
Orozco-Ca’rdenas ML, Ryan CA (2002) Nitric oxide negatively modulates wound signaling in tomato plants. Plant Physiol 130:487–493
Passardi F, Cosio C, Penel C, Dunand C (2005) Peroxidases have more functions than a Swiss army knife. Plant Cell Rep 24:255–265
Payne CT, Zhang F, Lloyd AM (2000) GL3 encodes a bHLH protein that regulates trichome development in Arabidopsis through interaction with GL1 and TTG1. Genetics 156:1349–1362
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29:e45
Robinson KA, Koepke JI, Kharodawala M, Lopes JM (2000) A network of yeast basic helix-loop-helix interactions. Nucleic Acids Res 28:4460–4466
Sambrook J, Russel DW (2001) Molecular cloning: a Laboratory Manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Seki M, Ishida J, Narusaka M, Fujita M, Nanjo T, Umezawa T, Kamiya A, Nakajima M, Enju A, Sakurai T, Satou M, Akiyama K, Yamaguchi-Shinozaki K, Carninci P, Kawai J, Hayashizaki Y, Shinozaki K (2002a) Monitoring the expression pattern of around 7,000 Arabidopsis genes under ABA treatments using a full-length cDNA microarray. Funct Integr Genom 2:282–291
Seki M, Narusaka M, Ishida J, Nanjo T, Fujita M, Oono Y, Kamiya A, Nakajima M, Enju A, Sakurai T, Satou M, Akiyama K, Taji T, Yamaguchi-Shinozaki K, Carninci P, Kawai J, Hayashizaki Y, Shinozaki K (2002b) Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J 31:279–292
Sessions A, Burke E, Presting G, Aux G, McElver J, Patton D, Dietrich B, Ho P, Bacwaden J, Ko C, Clarke JD, Cotton D, Bullis D, Snell J, Miguel T, Hutchison D, Kimmerly B, Mitzel T, Katagiri F, Glazebrook J, Law M, Goff SA (2002) A high-throughput Arabidopsis reverse genetics system. Plant Cell 14:2985–2994
Singh KB, Foley RC, Onate-Sanchez L (2002) Transcription factors in plant defense and stress responses. Curr Opin Plant Biol 5:430–436
Toledo-Ortiz G, Huq E, Quail PH (2003) The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell 15:1749–1770
Tracy FE, Gilliham M, Dodd AN, Webb AA, Tester M (2008) NaCl-induced changes in cytosolic free Ca2+ in Arabidopsis thaliana are heterogeneous and modified by external ionic composition. Plant Cell Environ 31:1063–1073
Welin BV, Olson A, Palva ET (1995) Structure and organization of two closely related low-temperature-induced dhn/lea/rab-like genes in Arabidopsis thaliana L. Heynh. Plant Mol Biol 29:391–395
Xiong LM, Zhu JK (2003) Regulation of abscisic acid biosynthesis. Plant Physiol 133:29–36
Xiong LM, Schumaker KS, Zhu JK (2002) Cell signaling during cold, drought, and salt stress. Plant Cell 14:S165–S183
Yamaguchi-Shinozaki K, Shinozaki K (1993) Characterization of the expression of a desiccation-responsive rd29 gene of Arabidopsis thaliana and analysis of its promoter in transgenic plants. Mol Gen Genet 236:331–340
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Ann Rev Plant Biol 57:781–803
Yang L, Zheng B, Mao C, Qi X, Liu F, Wu P (2004) Analysis of transcripts that are differentially expressed in three sectors of the rice root system under water deficit. Mol Genet Genom 272:433–442
Zhu J-K (2001) Cell signaling under salt, water and cold stresses. Curr Opin Plant Biol 4:401–406
Acknowledgments
We would like to thank ABRC for providing the T-DNA insertion and mutant seeds. We are also grateful to Dr. Anthony Cornish and Troy Locke in MBSU for help in microarray and qRT-PCR, to Dr. Enrico Scarpella for help in confocal microscopy, to Mary DePauw for technical assistance, and to Manjeet Kumari for developing Arabidopsis hydroponics system in our lab. The project was funded by a NSERC (Natural Sciences and Engineering Research Council) Discovery grant and Alberta Ingenuity awards to M.K.D.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Ronne.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jiang, Y., Yang, B. & Deyholos, M.K. Functional characterization of the Arabidopsis bHLH92 transcription factor in abiotic stress. Mol Genet Genomics 282, 503–516 (2009). https://doi.org/10.1007/s00438-009-0481-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-009-0481-3