Abstract
Objective
To evaluate if a deep learning model can be used to characterise breast cancers on contrast-enhanced spectral mammography (CESM).
Methods
This retrospective mono-centric study included biopsy-proven invasive cancers with an enhancement on CESM. CESM images include low-energy images (LE) comparable to digital mammography and dual-energy subtracted images (DES) showing tumour angiogenesis. For each lesion, histologic type, tumour grade, estrogen receptor (ER) status, progesterone receptor (PR) status, HER-2 status, Ki-67 proliferation index, and the size of the invasive tumour were retrieved. The deep learning model used was a CheXNet-based model fine-tuned on CESM dataset. The area under the curve (AUC) of the receiver operating characteristic (ROC) curve was calculated for the different models: images by images and then by majority voting combining all the incidences for one tumour.
Results
In total, 447 invasive breast cancers detected on CESM with pathological evidence, in 389 patients, which represented 2460 images analysed, were included. Concerning the ER, the deep learning model on the DES images had an AUC of 0.83 with the image-by-image analysis and of 0.85 for the majority voting. For the triple-negative analysis, a high AUC was observable for all models, in particularity for the model on LE images with an AUC of 0.90 for the image-by-image analysis and 0.91 for the majority voting. The AUC for the other histoprognostic factors was lower.
Conclusion
Deep learning analysis on CESM has the potential to determine histoprognostic tumours makers, notably estrogen receptor status, and triple-negative receptor status.
Key Points
• A deep learning model developed for chest radiography was adapted by fine-tuning to be used on contrast-enhanced spectral mammography.
• The adapted models allowed to determine for invasive breast cancers the status of estrogen receptors and triple-negative receptors.
• Such models applied to contrast-enhanced spectral mammography could provide rapid prognostic and predictive information.
Similar content being viewed by others

Avoid common mistakes on your manuscript.
Introduction
Worldwide, breast cancer accounts for 11.6% of cancers of all genders and ages. 2.08 million new cases and 626,679 deaths are estimated in 2018.
Contrast-enhanced spectral mammography (CESM), also called contrast-enhanced dual-energy mammography (CEDM), is an emerging technique used to evaluate neovascularity of a breast lesion, usually associated with malignancy [1], by using an iodine-based contrast intravenous agent.
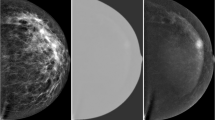
In this dual-energy breast imaging, a low-energy image (LE), comparable to conventional digital mammography (full-field digital mammography – FFDM) [2], is acquired and a recombined image (dual-energy subtracted DES) is obtained by subtracting both high- and low-energy images. The iodinated contrast material better absorbs high-energy X-photons, thus revealing regions of angiogenesis [3].
CESM has shown its superiority to FFDM in terms of sensitivity (92.7% vs 71.5%) and specificity (67.9% vs 51.8%) to detect primary breast cancers in dense breasts [4]. CESM has also a similar sensitivity and a slightly higher specificity compared to breast magnetic resonance imaging (MRI) which is considered the most sensitive technique for breast cancer detection (up to 100% sensitivity for both, with a specificity of 94% for CESM versus 88% for MRI in the studies) [5,6,7]. Moreover, compared to bMRI, CESM is more comfortable for the patients (position and faster acquisition time) but also less expensive [8, 9]. CESM is also efficient for the assessment of the extent of disease [10], for the evaluation of tumour response to neoadjuvant chemotherapy [11], for post-therapy monitoring [12], and as a problem-solving tool in association with other examinations [13].
According to the European Society of Breast Imaging (EUSOBI), CESM is currently recommended in case of bMRI contraindication or if bMRI is not available [14].
In parallel to the development of CESM, deep learning algorithm which attempts to learn multiple levels of representation of increasing complexity and abstraction [15] is a new powerful wave in all fields of radiology. Deep learning is a sub-domain of the machine learning (which is a sub-domain of artificial intelligence).
One of the latest examples of its use is in chest radiography, where a deep learning model, called CheXNeXt, can detect 14 clinically important pathologies [16]. On a validation set of 420 images for which the majority vote of 3 cardiothoracic speciality radiologists served as ground truth, the algorithm achieved performance equivalent to the practising radiologists on 10 pathologies, better on 1 pathology, and worse on 3 pathologies [16].
Concerning mammography, an international study has already shown that a deep learning model could be used successfully for the screening of primary breast cancers on FFDM. A combination of human and machine results had a performance equivalent to that of the traditional double-reading process but reduces the workload of the second reader by 88% [17]. More works are however needed for use in clinical practice [18].
As CESM is a recent technique, the contribution of deep learning to its interpretation has, to our knowledge, not been studied yet. In addition, following the EUSOBI recommendations, CESM is rather used as a second line after FFDM screening in place of bMRI.
Thus, rather than differentiating between benign and malignant in primary screening, CESM could be used to determine prognostic factors (as a histologic grade, Ki67 proliferation index, cell receptors) in case of cancer to adapt the treatment, as a “deep-radiomics” tool [19]. Most prognostic factors are determined histologically. However, it is possible that the histological phenotype can also be identified in X-ray imaging [20], notably through deep learning analysis.
However, as CESM is a new technique, available data is still scarce. In such a situation, it is possible to use transfer learning with fine-tuning to adapt a deep learning model, learned on a large dataset (e.g., more than 100,000 images) to a new task with fewer images (e.g., less than 1000 images). Transfer learning from CheXNeXt has already been used on FFDM to detect malignant lesions, with the best accuracy of 90.38% [21]. More recently, transfer learning from CheXNeXt has been used to detect COVID-19 infection (COVID-CXNet) with an accuracy of 94.20% [22].
The aim of this study was to evaluate if a deep learning model can be used to characterise breast malignant lesions on CESM.
Materials and methods
Data set
This was a retrospective mono-centric study (Henri Becquerel Cancer Center, Rouen Normandy, France) approved by the local ethics committee. Patients were informed that their anonymised images could be used for research and gave their consent. Indications for CESM in our institution were the assessment of the extent of invasive disease (all invasive cancers, excluding in situ cancers), the evaluation of tumour response to neoadjuvant chemotherapy, post-therapy monitoring, and as a problem-solving tool. CESM was performed also in case of contraindications to performing bMRI or if bMRI was not available.
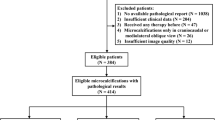
All CESM images performed between August 2017 and January 2019 were retrieved. Only biopsy-proven invasive cancers with an enhancement on CESM were included. CESM with benign lesions or borderline lesions or non-enhanced malignant lesions were not included. The exclusion criteria were as follows: chemotherapy prior to CESM, lack of histological data, the presence of artefacts (no possible visual analysis), a non-contributory examination (a lesion with very low enhancement, or a masking background enhancement making reliable segmentation of the lesion impossible) (Fig. 1).
For each lesion, its size (maximum diameter) was recorded. The breast density was evaluated according to the BI-RADS lexicon and the background parenchymal enhancement was scaled from 0 to 3 (0 = no enhancement, 1 = minimal enhancement, 2 = moderate enhancement, and 3 = marked enhancement). The type of enhancement (mass or non-mass enhancement) and the biopsy markers were retrieved.
In the case of multi-focal disease, the lesion with the maximum diameter was selected and the other lesions could also be selected if they were well separated with clear limits in at least one incidence.
Histopathology
The histological characteristics were based on analysis of the surgical specimen or percutaneous biopsy if the definitive histologic analysis was not available or if the patient had received neoadjuvant chemotherapy.
For each lesion, histological type, Nottingham histological score (histologic grade), estrogen receptor (ER) status, progesterone receptor (PR) status, HER-2 status, Ki-67 proliferation index, and the size of the invasive tumour were retrieved.
According to current guidelines in France, the status was considered positive when the hormonal receptors (ER and PR) had a value ≥ 10% [23]. According to the GEFPICS recommendation, HER2 was considered positive if IHC results are 3 + or 2 + amplified on ISH (in situ hybridization) [24]. Similarly, the Ki 67 was considered high if it was ≥ 20% [25]. Triple-negative status, a known poor prognostic factor [26], was considered for a combination of ER-negative, PR-negative, and HER2-negative.
CESM image acquisition
CESM was performed on mammography system (Angiologic Stephanix, Hologic®) using an automated single-shot intravenous injection of iodinated contrast-agent (Iomeron 350 mg/mL, Bracco®) with the dose of 1.5 mL/kg body weight with a flow rate of 3 mL/s followed by a saline flush, after eliminating contraindications. CESM image protocol acquisition is described in supplemental data.
Image pre-processing
Patient images were extracted from the PACS (Picture Archiving and Communication System) and anonymised. Manual segmentation of the images was performed by a radiologist using an in-house plugin for the ImageJ software [27]. All available breast incidences were considered. In the case of a visible lesion, the regions of interest (ROI) of the lesions were segmented as the smallest rectangular ROI encompassing the entire lesion [28] on the DES images, the subtracted images corresponding to the vascularisation. The segmented ROI was then propagated on the low-energy images, corresponding to an image similar to FFDM. Images were normalised to an 8bit (0–255) images. Images were then cropped from a 2560 × 3328 array to 1024 × 1024 array with the ROI lesion at the centre. In the case of a ROI lesion larger than 1024 in at least one axis (e.g., in case of large breast masses), the ROI lesion was proportionally resized to a 1024 × 1024 array. Image augmentation is utilised in order to prevent over-fitting by using orthogonal (90°, 180°, and 270°) rotations. Images were down-sized to 320 × 320 array to prevent resource exhaustion and decrease GPU RAM usage.
Deep learning analysis
Dataset was separated randomly on training (2 × 787 images, 64%), validation (2 × 195 images, 16%), and testing (2 × 248 images, 20%) datasets. Images of one patient could only be in one of these datasets, even if multiple lesions were segmented. Frequencies of the different issues were equalised between the datasets.
The deep learning model used was a CheXNet-based model [29], fine-tuned on CESM dataset. CheXNet is based on a densely connected convolutional network with 121 layers (DenseNet-121 [30]) that takes a chest X-ray image as input and outputs the probability of a pathology. The CheXNet model was trained on a dataset containing 112,120 frontal-view chest X-ray images individually labelled with up to 14 different thoracics [29]. This model, specialised in the analysis of thoracic radiographs, can be adapted by fine-tuning, a partial relearning on a new database, in order to analyse new pathologies, such as COVID, but also other examinations, notably mammography [21, 22]. For the fine-tuning, similar to the one performed in [22], Adam optimiser was used with a learning rate of 0.0001 and 10 epochs. A fully connected layer followed by a dropout layer with a 0.2 dropping rate to prevent overfitting was used. The architecture of the model and fine-tuning performed is presented in Fig. 2.
Three models were trained. The first one on the DES images only, the second one on the LE images only, and the third one on both DES and LE images. Images were analysed independently of the incidence. A majority voting, combining all the incidences for one tumour, was also performed for the three models. The learning of the models was done using python 2.7.17, tensorflow 2.4.1, and keras 2.4.3.
Statistical analysis
The receiver operating characteristic (ROC) curve and area under (AUC) of the ROC curve were calculated to assess the ability of the CNN to determine tumour grade (1 and 2 vs. 3), hormone receptor status (ER positive vs. ER negative, PR positive vs. PR negative), Her2 (Her2 positive vs. Her2 negative), and Ki 67 (high or not), and triple negative vs. other.
The model was evaluated image by the image on DES and LE images separately, and then for all images simultaneously. Then results were calculated by majority voting (including all the incidences for one tumour) on the DES, LE images, and for the combination of LE and DES images.
We computed overall classification accuracy, as well as sensitivity, specificity, from the results image by image on DES and LE images separately, and then for all images simultaneously.
Statistical analysis was performed with python 2.7.17.
Results
Patient population and lesion characteristics
A total of 447 malignant lesions in 389 patients (mean age, 61.7 ± 12.6 years; age range, 27–91) were detected on CESM with pathological evidence, which ultimately represents 2460 images analysed.
Patient demographics and imaging characteristics are provided in Table 1. The median size of enhancing lesions was 28.3 mm ± 23.0 (range 5.0–130.0). A description of the types of lesions and their histoprognostic factors are given in Table 2.
There were no significant differences between the training, the validation, and the testing sets in terms of histological grade, estrogen receptor (ER) status, progesterone receptor (PR) status, HER-2 status, Ki-67 proliferation index, or triple negative.
Deep learning analysis
Figures 3 and 4 illustrate the performance of the CNN algorithm area under the receiver operating characteristic curve for the analysis image by image and by majority voting respectively.
Concerning the ER, the deep learning model on the DES images had an AUC of 0.83 with the image-by-image analysis and of 0.85 for the majority voting. These results were slightly higher than those obtained for the LE images (0.73 and 0.72, respectively) but comparable to those obtained when DES and LE images were combined (0.85 and 0.86, respectively). For the triple-negative analysis, a high AUC was observable for all models, in particularity for the model on LE images with an AUC of 0.90 for the image-by-image analysis and 0.91 for the majority voting.
The AUCs for the other histoprognostic factors were lower; for example, when DES and LE images were combined, AUC = 0.65 for the diagnosis of the HER2 status or AUC = 0.62 to predict grade 3.
The sensitivity, specificity, and accuracy of the deep learning model image by the image on DES and LE images separately, and then for all images simultaneously for each category are provided in Table 3.
The best results were observed with the determination of the ER status on DES images with a sensitivity, a specificity, and an accuracy of 82% and with the determination of the triple-negative status on LE images with a sensitivity of 83%, a specificity of 85%, and an accuracy of 85%.
To illustrate a decision taken by the deep learning model, a representation of a gradCAM (a heat map showing the part of the image where the algorithm is focused for its decision) of the models for a triple-negative cancer is shown in Fig. 5.
Discussion
The aim of this retrospective study was to define if a deep learning model based on transfer learning can be used to characterise breast malignant lesions on CESM. Our results demonstrated a good performance to characterise estrogen receptor (ER) status and to differentiate triple-negative breast cancers (TNBC).
The characterisation of tumours is essential to define the best management and choose the different therapeutics. The analysis by deep learning allows the study of an entire tumour and of each detected lesion quickly and non-invasively. This model could serve as a virtual biopsy and could also help radiologists target their biopsy to areas of interest.
Previous studies on CESM and machine learning have focused on the development of CAD systems to classify breast masses as benign or malignant.
Patel et al. used a machine learning algorithm based on SVM (support vector machine) classification and added radiomics analysis to show the diagnostic performance of computer-assisted contrast-enhanced spectral mammography (CAD-CESM) on 50 lesions described on the CESM. Their results showed that this CAD-CESM can provide an overall diagnostic accuracy of 90% in predicting the malignancy of a lesion and that it could help radiologists’ interpretation, mainly by reducing the number of false-positive results [31]. Massafra et al. evaluated on 58 ROIs from 53 patients a CAD based on a radiomics analysis with 464 features of different kinds (such as points and corners of interest, textural and statistical features) on native or filtered images. After a principal component analysis (PCA), dimension reduction technique, and the use of random forest classifier, they obtained a model able to predict benign/malignant ROIs with median values for sensitivity and specificity of 88.37% and 100% [32]. These results improved on previous results obtained by the same team with median values of sensitivity and specificity of 87.5% and 91.7%, respectively [33].
In the study by Danala et al., they developed a CAD scheme of CESM images to classify breast masses. They demonstrated that CESM images could improve the accuracy of mass segmentation of a CAD and classify 111 lesions as benign or malignant using perceptron-based multilayer machine learning. Their results also showed that LE and DES images contain valuable additional information and that the combination of these two types of images significantly improved CAD lesion classification performance [34].
Perek et al. [35] compared two CNN methods, fine-tuning a pre-trained network and fully training a convolutional neural network, for the classification of CESM breast mass as benign or malignant on 129 randomly selected breast lesions and obtained the following results using a multimodal network with 100% sensitivity and 66% specificity.
Gao et al. [36] evaluated a deep-CNN using 49 CESM cases employed to extract novel features from LE, recombined or “virtual” recombined images to classify the cases as benign vs. cancer. They proved that the model performance can achieve an accuracy of 0.85 and an AUC of 0.84 using LE imaging, and they have shown to improve the accuracy of 0.89 with an AUC of 0.91 using both LE and DES imaging.
There are no studies to date on the potential of deep learning to characterise malignant lesions. In addition, previous studies on machine learning and CESM were conducted on a smaller number of cases.
In a study by La Forgia et al. performed on 68 lesions from 52 patients, seven features (histogram and texture parameters) were extracted from each original ROI from both LE and DES images or 14 features per ROI. With a multivariate linear discriminant analysis, they found that for the tasks to differentiate ER status, PR status, Ki67, grade, TNBC status, and HER status, AUC values of 83.79%, 75.50%, 84.80%, 79.85%, 76.80%, and 90.89%, respectively should be achieved.
In another study performed by Marino et al., which also deals with CESM and artificial intelligence, the authors used another area of machine learning and radiomics analysis with the extraction of tumour features followed by a machine learning classification, to characterise breast lesions. They showed in their retrospective study of 103 breast cancers that radiomics analysis with CESM was able to differentiate invasive tumours from non-invasive and define their hormone receptor status and tumour grade [37].
We found comparable results for the determination of ER status but not for the PR status and the grade. However, we found that our deep learning model was able to determine the TNBC status. For TNBC, the results observed can be related to the characteristics of these tumours that present a specific appearance in imaging (an irregular non-calcified mass with ill-defined or spiculated margins on mammography, and a hypoechoic or complex mass with an irregular shape and non-circumscribed margins on ultrasound [38] and a rim enhancement on DCE-MRI for example) [39]. Interestingly, in accordance with the radiological semiology described, the models developed in this study seem to take their decision on the ring for the DES images and on the ill-defined or spiculated margins for the LE images in case showed in Fig. 5. The quick detection before any surgery of TNBC is very interesting for this type of aggressive tumour for which patients can benefit from neoadjuvant chemotherapy in some cases. However, the results obtained with the LE images, comparable to the FFDM, and with the DES images were globally similar without any obvious improvement of the results by combining them with the majority voting system. This could call into question the interest of injection for this indication, although tumour localisation and segmentation are greatly facilitated by injection.
Our study has several limitations. First, it is a retrospective study mono-centric with a relatively small number of patients for deep learning analysis although we used data augmentation and transfer learning with fine-tuning to overcome this limitation.
In our study, all the examinations were performed on the same mammography machine, which made it possible to obtain standardised images, but the results may not be applicable in other centres using other equipment.
Finally, we did not take into account lesions that do not enhancement on the CESM and are only visible on the low-energy image, which may induce a selection bias in our study.
Conclusion
In conclusion, our results demonstrate that deep learning analysis on CESM has the potential to determine histoprognostic tumour makers, notably estrogen receptor status and triple-negative receptor status, which could provide rapid prognostic and predictive information.
Abbreviations
- AUC:
-
Area under the curve
- CC:
-
Craniocaudal
- CEDM:
-
Contrast-enhanced dual-energy mammography
- CESM:
-
Contrast-enhanced spectral mammography
- DES:
-
Dual-energy subtracted
- ER:
-
Estrogen receptor
- FFDM:
-
Full-field digital mammography
- LE:
-
Low-energy image
- LM:
-
Lateromedial
- MLO:
-
Media-lateral oblique
- MRI:
-
Magnetic resonance imaging
- PACS:
-
Picture Archiving and Communication System
- PR:
-
Progesterone receptor
- ROC:
-
Receiver operating characteristic
- ROI:
-
Region of interest
- TNBC:
-
Triple-negative breast cancers
References
Jochelson M (2014) Contrast-enhanced digital mammography. Radiol Clin North Am 52:609–616. https://doi.org/10.1016/j.rcl.2013.12.004
Lalji UC, Jeukens CRLPN, Houben I et al (2015) Evaluation of low-energy contrast-enhanced spectral mammography images by comparing them to full-field digital mammography using EUREF image quality criteria. Eur Radiol 25:2813–2820. https://doi.org/10.1007/s00330-015-3695-2
Dromain C, Balleyguier C, Adler G et al (2009) Contrast-enhanced digital mammography. Eur J Radiol 69:34–42. https://doi.org/10.1016/j.ejrad.2008.07.035
Cheung Y-C, Lin Y-C, Wan Y-L et al (2014) Diagnostic performance of dual-energy contrast-enhanced subtracted mammography in dense breasts compared to mammography alone: interobserver blind-reading analysis. Eur Radiol 24:2394–2403. https://doi.org/10.1007/s00330-014-3271-1
Jochelson MS, Dershaw DD, Sung JS et al (2013) Bilateral contrast-enhanced dual-energy digital mammography: feasibility and comparison with conventional digital mammography and MR imaging in women with known breast carcinoma. Radiology 266:743–751. https://doi.org/10.1148/radiol.12121084
Fallenberg EM, Schmitzberger FF, Amer H et al (2017) Contrast-enhanced spectral mammography vs. mammography and MRI – clinical performance in a multi-reader evaluation. Eur Radiol 27:2752–2764. https://doi.org/10.1007/s00330-016-4650-6
Li L, Roth R, Germaine P et al (2017) Contrast-enhanced spectral mammography (CESM) versus breast magnetic resonance imaging (MRI): a retrospective comparison in 66 breast lesions. Diagn Interv Imaging 98:113–123. https://doi.org/10.1016/j.diii.2016.08.013
Hobbs MM, Taylor DB, Buzynski S, Peake RE (2015) Contrast-enhanced spectral mammography (CESM) and contrast enhanced MRI (CEMRI): patient preferences and tolerance: CESM and CEMRI preferences and tolerance. J Med Imaging Radiat Oncol 59:300–305. https://doi.org/10.1111/1754-9485.12296
Patel BK, Gray RJ, Pockaj BA (2017) Potential cost savings of contrast-enhanced digital mammography. AJR Am J Roentgenol 208:W231–W237. https://doi.org/10.2214/AJR.16.17239
Lee-Felker SA, Tekchandani L, Thomas M et al (2017) Newly diagnosed breast cancer: comparison of contrast-enhanced spectral mammography and breast MR imaging in the evaluation of extent of disease. Radiology 285:389–400. https://doi.org/10.1148/radiol.2017161592
Iotti V, Ravaioli S, Vacondio R et al (2017) Contrast-enhanced spectral mammography in neoadjuvant chemotherapy monitoring: a comparison with breast magnetic resonance imaging. Breast Cancer Res 19:106. https://doi.org/10.1186/s13058-017-0899-1
Badr S, Laurent N, Régis C et al (2014) Dual-energy contrast-enhanced digital mammography in routine clinical practice in 2013. Diagn Interv Imaging 95:245–258. https://doi.org/10.1016/j.diii.2013.10.002
Dromain C, Vietti-Violi N, Meuwly JY (2019) Angiomammography: a review of current evidences. Diagn Interv Imaging 100:593–605. https://doi.org/10.1016/j.diii.2019.01.011
Sardanelli F, Fallenberg EM, Clauser P et al (2017) Mammography: an update of the EUSOBI recommendations on information for women. Insights Imaging 8:11–18. https://doi.org/10.1007/s13244-016-0531-4
LeCun Y, Bengio Y, Hinton G (2015) Deep learning. Nature 521:436–444. https://doi.org/10.1038/nature14539
Rajpurkar P, Irvin J, Ball RL et al (2018) Deep learning for chest radiograph diagnosis: a retrospective comparison of the CheXNeXt algorithm to practicing radiologists. PLoS Med 15:e1002686. https://doi.org/10.1371/journal.pmed.1002686
McKinney SM, Sieniek M, Godbole V et al (2020) International evaluation of an AI system for breast cancer screening. Nature 577: https://doi.org/10.1038/s41586-019-1799-6
Geras KJ, Mann RM, Moy L (2019) Artificial intelligence for mammography and digital breast tomosynthesis: current concepts and future perspectives. Radiology 293:246–259. https://doi.org/10.1148/radiol.2019182627
Parekh VS, Jacobs MA (2019) Deep learning and radiomics in precision medicine. Expert Rev Precis Med Drug Dev 4:59–72. https://doi.org/10.1080/23808993.2019.1585805
Boisserie-Lacroix M, Hurtevent-Labrot G, Ferron S et al (2013) Correlation between imaging and molecular classification of breast cancers. Diagn Interv Imaging 94:1069–1080. https://doi.org/10.1016/j.diii.2013.04.010
Pardamean B, Cenggoro TW, Rahutomo R et al (2018) Transfer learning from chest X-ray pre-trained convolutional neural network for learning mammogram data. Procedia Computer Science 135:400–407. https://doi.org/10.1016/j.procs.2018.08.190
Haghanifar A, Majdabadi MM, Choi Y, et al (2020) COVID-CXNet: Detecting COVID-19 in Frontal Chest X-ray Images using Deep Learning. arXiv:200613807 [cs, eess]
Yi M, Huo L, Koenig KB et al (2014) Which threshold for ER positivity? a retrospective study based on 9639 patients. Ann Oncol 25:1004–1011. https://doi.org/10.1093/annonc/mdu053
Penault-Llorca F, Vincent-Salomon A, MacGrogan G et al (2014) 2014 update of the GEFPICS’ recommendations for HER2 status determination in breast cancers in France. Ann Pathol 34:352–365. https://doi.org/10.1016/j.annpat.2014.08.018
Gnant M, Thomssen C, Harbeck N (2015) St. Gallen/Vienna 2015: A brief summary of the consensus discussion. Breast Care (Basel) 10:124–130. https://doi.org/10.1159/000430488
Yamamoto Y, Iwase H (2010) Clinicopathological features and treatment strategy for triple-negative breast cancer. Int J Clin Oncol 15:341–351. https://doi.org/10.1007/s10147-010-0106-1
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Meth 9:671–675. https://doi.org/10.1038/nmeth.2089
Zhou J, Zhang Y, Chang K-T et al (2020) Diagnosis of benign and malignant breast lesions on DCE-MRI by using radiomics and deep learning with consideration of peritumor tissue. J Magn Reson Imaging 51:798–809. https://doi.org/10.1002/jmri.26981
Rajpurkar P, Irvin J, Zhu K et al (2017) CheXNet: radiologistlevel pneumonia detection on chest X-rays with deep learning. arXiv:1711.05225
Huang G, Liu Z, Van Der Maaten L, Weinberger KQ (2017) Densely connected convolutional networks. In: 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). pp 2261–2269
Patel BK, Ranjbar S, Wu T et al (2018) Computer-aided diagnosis of contrast-enhanced spectral mammography: a feasibility study. Eur J Radiol 98:207–213. https://doi.org/10.1016/j.ejrad.2017.11.024
Massafra R, Bove S, Lorusso V et al (2021) Radiomic feature reduction approach to predict breast cancer by contrast-enhanced spectral mammography images. Diagnostics 11:684. https://doi.org/10.3390/diagnostics11040684
Fanizzi A, Losurdo L, Basile TMA et al (2019) Fully automated support system for diagnosis of breast cancer in contrast-enhanced spectral mammography images. J Clin Med 8:E891. https://doi.org/10.3390/jcm8060891
Danala G, Patel B, Aghaei F et al (2018) Classification of breast masses using a computer-aided diagnosis scheme of contrast enhanced digital mammograms. Ann Biomed Eng 46:1419–1431. https://doi.org/10.1007/s10439-018-2044-4
Perek S, Kiryati N, Zimmerman-Moreno G et al (2019) Classification of contrast-enhanced spectral mammography (CESM) images. Int J Comput Assist Radiol Surg 14:249–257. https://doi.org/10.1007/s11548-018-1876-6
Gao F, Wu T, Li J et al (2018) SD-CNN: a shallow-deep CNN for improved breast cancer diagnosis. Comput Med Imaging Graph 70:53–62. https://doi.org/10.1016/j.compmedimag.2018.09.004
Marino MA, Pinker K, Leithner D et al (2020) Contrast-enhanced mammography and radiomics analysis for noninvasive breast cancer characterization: initial results. Mol Imaging Biol 22:780–787. https://doi.org/10.1007/s11307-019-01423-5
Krizmanich-Conniff KM, Paramagul C, Patterson SK et al (2012) Triple receptor–negative breast cancer: imaging and clinical characteristics. AJR Am J Roentgenol 199:458–464. https://doi.org/10.2214/AJR.10.6096
Youk JH, Son EJ, Chung J et al (2012) Triple-negative invasive breast cancer on dynamic contrast-enhanced and diffusion-weighted MR imaging: comparison with other breast cancer subtypes. Eur Radiol 22:1724–1734. https://doi.org/10.1007/s00330-012-2425-2
Funding
The authors state that this work has not received any funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Pierre Decazes.
Conflict of interest
The authors of this manuscript declare no relationships with any companies, whose products or services may be related to the subject matter of the article.
Statistics and biometry
One of the authors has significant statistical expertise.
Informed consent
Written informed consent was obtained from all subjects (patients) in this study.
Ethical approval
Institutional Review Board approval was obtained.
Methodology
• Retrospective.
• Diagnostic or prognostic study.
• Performed at one institution.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dominique, C., Callonnec, F., Berghian, A. et al. Deep learning analysis of contrast-enhanced spectral mammography to determine histoprognostic factors of malignant breast tumours. Eur Radiol 32, 4834–4844 (2022). https://doi.org/10.1007/s00330-022-08538-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-022-08538-4