Abstract
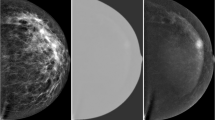
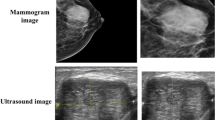
Contrast-enhanced digital mammography (CEDM) is a promising imaging modality in breast cancer diagnosis. This study aims to investigate how to optimally develop a computer-aided diagnosis (CAD) scheme of CEDM images to classify breast masses. A CEDM dataset of 111 patients was assembled, which includes 33 benign and 78 malignant cases. Each CEDM includes two types of images namely, low energy (LE) and dual-energy subtracted (DES) images. A CAD scheme was applied to segment mass regions depicting on LE and DES images separately. Optimal segmentation results generated from DES images were also mapped to LE images or vice versa. After computing image features, multilayer perceptron based machine learning classifiers that integrate with a correlation-based feature subset evaluator and leave-one-case-out cross-validation method were built to classify mass regions. When applying CAD to DES and LE images with original segmentation, areas under ROC curves (AUC) were 0.759 ± 0.053 and 0.753 ± 0.047, respectively. After mapping the mass regions optimally segmented on DES images to LE images, AUC significantly increased to 0.848 ± 0.038 (p < 0.01). Study demonstrated that DES images eliminated overlapping effect of dense breast tissue, which helps improve mass segmentation accuracy. The study demonstrated that applying a novel approach to optimally map mass region segmented from DES images to LE images enabled CAD to yield significantly improved performance.








Similar content being viewed by others
References
Aghaei, F., M. Tan, A. B. Hollingsworth, and B. Zheng. Applying a new quantitative global breast MRI feature analysis scheme to assess tumor response to chemotherapy. J. Magn. Reson. Imaging 44:1099–1106, 2016.
Baltzer, P. A. T., M. Benndorf, M. Dietzel, M. Gajda, I. B. Runnebaum, and W. A. Kaiser. False-positive findings at contrast-enhanced breast MRI: a BI-RADS descriptor study. Am. J. Roentgenol. 194:1658–1663, 2010.
Berg, W. A., Z. Zhang, D. Lehrer, and R. A. Jong. Detection of breast cancer with addition of annual screening ultrasound or a single screening MRI to mammography in women with elevated breast cancer risk. J. Am. Med. Assoc. 307:1394–1404, 2012.
Brodersen, J., and V. Siersma. Long-term psychosocial consequences of false-positive mammography screening. Ann. Fam. Med. 11:106–115, 2013.
Carney, P. A., D. L. Miglioretti, B. C. Yankaskas, K. Kerlikowske, R. Rosenberg, C. M. Rutter, B. M. Geller, L. A. Abraham, S. H. Taplin, M. M. Dignan, and R. Gary Cutter. Individual and combined effects of age, breast density, and hormone replacement therapy use on the accuracy of screening mammography. Ann. Intern. Med. 138:168–175, 2003.
Chawla, N. V., K. W. Bowyer, L. O. Hall, and W. P. Kegelmeyer. SMOTE: synthetic minority over-sampling technique. J. Artif. Intell. Res. 16:321–357, 2002.
Danala, G., T. Thai, C. C. Gunderson, K. M. Moxley, K. Moore, R. S. Mannel, H. Liu, B. Zheng, and Y. Qiu. Applying quantitative CT image feature analysis to predict response of ovarian cancer patients to chemotherapy. Acad. Radiol. 24:1233–1239, 2017.
Dromain, C., C. Balleyguier, S. Muller, M.-C. Mathieu, F. Rochard, P. Opolon, and R. Sigal. Evaluation of tumor angiogenesis of breast carcinoma using contrast-enhanced digital mammography. Am. J. Roentgenol. 187:W528–W537, 2006.
Eltonsy, N. H., G. D. Tourassi, A. S. Elmaghraby, and S. Member. A concentric morphology model for the detection of masses in mammography. IEEE Trans. Med. Imaging 26:880–889, 2007.
Gur, D., J. Stalder, and L. A. Hardesty. CAD performance on sequentially ascertained mammographic examinations of masses: an assessment. Radiology 233:418–423, 2004.
Kriege, M., C. T. M. Brekelmans, C. Boetes, and P. E. Besnard. Efficacy of MRI and mammography for breast-cancer screening in women with a familial or genetic predisposition. N. Engl. J. Med. 351:427–437, 2004.
Laya, M. B., E. B. Larson, S. H. Taplin, and E. White. Effect of estrogen replacement therapy on the specificity and sensitivity of screening mammography. J. Natl. Cancer Inst. 88:643–649, 1996.
Lee, A. Y., D. J. Wisner, S. Aminololama-Shakeri, V. A. Arasu, S. A. Feig, J. Hargreaves, H. Ojeda-Fournier, L. W. Bassett, C. J. Wells, J. De Guzman, C. I. Flowers, J. E. Campbell, S. L. Elson, H. Retallack, and B. N. Joe. Inter-reader variability in the use of BI-RADS descriptors for suspicious findings on diagnostic mammography: a multi-institution study of 10 academic radiologists. Acad. Radiol. 24:60–66, 2017.
Li, Q., and K. Doi. Reduction of bias and variance for evaluation of computer-aided diagnostic schemes. Med. Phys. 33:868–875, 2006.
Lu, W., Z. Li, and J. Chu. A novel computer-aided diagnosis system for breast MRI based on feature selection and ensemble learning. Comput. Biol. Med. 83:157–165, 2017.
Mandelson, T. M. Breast density as a predictor of breast cancer risk. J. Natl. Cancer Inst. 12:1081–1087, 2000.
Oliver, A., J. Freixenet, J. Martí, E. Pérez, J. Pont, E. R. E. Denton, and R. Zwiggelaar. A review of automatic mass detection and segmentation in mammographic images. Med. Image Anal. 14:87–110, 2010.
Patel, B. K., S. Ranjbar, T. Wu, B. A. Pockaj, J. Li, N. Zhang, M. Lobbes, B. Zhang, and J. R. Mitchell. Computer-aided diagnosis of contrast-enhanced spectral mammography: a feasibility study. Eur. J. Radiol. 98:207–213, 2018.
Peer, P. G. M., A. L. M. Verbeek, H. Straatman, J. H. C. L. Hendriks, and R. Holland. Age-specific sensitivities of mammographic screening for breast cancer. Breast Cancer Res. 38:153–160, 1996.
Qiu, Y., S. Yan, R. R. Gundreddy, Y. Wang, S. Cheng, H. Liu, and B. Zheng. A new approach to develop computer-aided diagnosis scheme of breast mass classification using deep learning technology. J. Xray. Sci. Technol. 25:751–763, 2017.
Rebecca, A. H., K. Kerlikowske, C. I. Flowers, B. C. Yankaskas, Z. Weiwei, and D. L. Miglioretti. Cumulative probability of false-positive recall or biopsy recommendation after 10 years of screening mammography. Ann. Intern. Med. 155:481–492, 2011.
Saeys, Y., I. Inza, and P. Larranaga. A review of feature selection techniques in bioinformatics. Bioinformatics 23:2507–2517, 2007.
Tan, M., F. Aghaei, Y. Wang, and B. Zheng. Developing a new case based computer-aided detection scheme and an adaptive cueing method to improve performance in detecting mammographic lesions. Phys. Med. Biol. 62:358–376, 2017.
Tan, M., B. Zheng, J. K. Leader, and D. Gur. Association between changes in mammographic image features and risk for near-term breast cancer development. IEEE Trans. Med. Imaging 35:1719–1728, 2016.
Wang, Y., F. Aghaei, A. Zarafshani, Y. Qiu, W. Qian, and B. Zheng. Computer-aided classification of mammographic masses using visually sensitive image features. J. Xray. Sci. Technol. 25:171–186, 2017.
Wang, X., D. Lederman, J. Tan, X. H. Wang, and B. Zheng. Computerized prediction of risk for developing breast cancer based on bilateral mammographic breast tissue asymmetry. Med. Eng. Phys. 27:934–942, 2011.
Weaver, D. L., R. D. Rosenberg, W. E. Barlow, L. Ichikawa, P. A. Carney, K. Kerlikowske, D. S. Buist, B. M. Geller, C. R. Key, S. J. Maygarden, and R. Ballard-Barbash. Pathologic findings from the breast cancer surveillance consortium. Cancer 106(4):732–742, 2006.
Witten, I. H., E. Frank, and M. A. Hall. Data Mining: Practical Machine Learning Tools and Techniques (3rd ed.). Amsterdam: Elsevier, 2011.
Zheng, B., J. H. Sumkin, M. L. Zuley, D. Lederman, X. Wang, and D. Gur. Computer-aided detection of breast masses depicted on full-field digital mammograms: a performance assessment. Br. J. Radiol. 85:e153–e161, 2012.
Acknowledgment
This work is supported in part by Grant R01 CA197150 from the National Cancer Institute, National Institutes of Health, USA.
Author information
Authors and Affiliations
Corresponding author
Additional information
Associate Editor Ender A Finol oversaw the review of this article.
Rights and permissions
About this article
Cite this article
Danala, G., Patel, B., Aghaei, F. et al. Classification of Breast Masses Using a Computer-Aided Diagnosis Scheme of Contrast Enhanced Digital Mammograms. Ann Biomed Eng 46, 1419–1431 (2018). https://doi.org/10.1007/s10439-018-2044-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10439-018-2044-4