Abstract
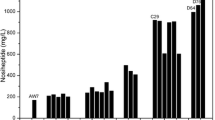
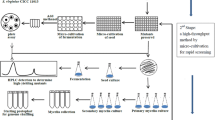
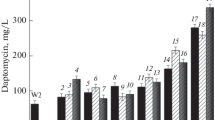
Avilamycin is one of EU-approved antimicrobial agents in feed industry to inhibit the growth of multidrug-resistant Gram-positive bacteria. Here, we applied a process of combining ribosome engineering and genome shuffling to achieve rapid improvement of avilamycin production in Streptomyces viridochromogenes AS 4.126. The starting mutant population was generated by 60Co γ-irradiation treatments of the spores. After five rounds of protoplast fusion with streptomycin-resistance screening, an improved recombinant E-219 was obtained and its yield of avilamycin reached 1.4 g/L, which was increased by 4.85-fold and 36.8-fold in comparison with that of the shuffling starter Co γ-316 and the ancestor AS 4.126. Furthermore, the mechanism for the improvement of shuffled strains was investigated. Recombinants with enhanced streptomycin resistance exhibited significantly higher avilamycin production and product resistance, probably due to the mutations in the ribosome protein S12. The morphological difference between the parent mutant and shuffled recombinant was observed in conidiospore, and hyphae pellets. The presence of genetic diversity among shuffled populations with varied avilamycin productivity was confirmed by randomly amplified polymorphic DNA analysis. In summary, our results demonstrated that genome shuffling combined with ribosome engineering was a powerful approach for molecular breeding of high-yield industrial strains.





Similar content being viewed by others
References
Aarestrup FM, Jensen LB (2000) Presence of variation in ribosomal protein L16 corresponding to susceptibility of enterococci to oligosaccharides (avilamycin and evernimicin). Antimicrob Agents Chemother 44:3425–3427
Adrio JL, Demain AL (2006) Genetic improvement of processes yielding microbial products. FEMS Microbiol Rev 30:187–214
Buzzetti F, Eisenberg F, Grant HN, Keller-Schierlein W, Voser W, Zāhner H (1968) Avilamycin. Experimentia 24:320–324
Chen JM, Xu LT (1991) Analysis of antibiotic industry, 2nd edn. Pharmaceutical Science, Beijing, pp 109–139
El-Fendy MM, EL-Bondkly AM (2011) Genome shuffling of marine derived bacterium Nocardia sp. ALAA 2000 for improved ayamycin production. Antonie van Leeuwenhoek 99:773–780
Foster DR, Rybak MJ (1999) Pharmacologic and bacteriologic properties of SCH-27899 (Ziracin), an investigational antibiotic from the everninomicin family. Pharmacotherapy 19:1111–1117
Hida H, Yamada T, Yamady Y (2007) Genome shuffling of Streptomyces sp U121 for improved production of hydroxycitric acid. Appl Microbiol Biotechnol 73:1387–1393
Hopwood DA, Bibb MJ, Chater KF, Kieser T, Bruton CJ, Kieser HM, Lydiate DJ, Smith CP, del Ward JM, Schrempf H (1985) Genetic manipulation of Streptomyces: a laboratory manual. John Innes Foundation, Norwich
Hosaka T, Ohnishi-Kameyama M, Muramatsu H, Murakami K, Tsurumi Y, Kodani S, Yoshida M, Fujie A, Ochi K (2009) Antibacterial discovery in actinomycetes strains with mutations in RNA polymerase or ribosomal protein S12. Nat Biotechnol 27:462–464
Hosoya Y, Okamoto S, Muramatsu H, Ochi K (1998) Acquisition of certain streptomycin-resistant (str) mutations enhances antibiotic production in bacteria. Antimicrob Agents Chemother 42:2041–2047
Hu H, Ochi K (2001) Novel approach for improving the productivity of antibiotic-producing strains by inducing combined resistant mutations. Appl Environ Microbiol 67:1885–1892
Jia B, Jin ZH, Lei YL, Mei LH, Li NH (2006) Improved production of pristinamycin coupled with an adsorbent resin in fermentation by Streptomyces pristinaespiralis. Biotechnol Lett 28:1811–1815
Knapp JE, Chandlee JM (1996) RNA/DNA mini-prep from a single sample of orchid tissue. Biotechniques 21:54–56
Manteca A, Alvarez R, Salazar N, Yagüe P, Sanchez J (2008) Mycelium differentiation and antibiotic production in submerged cultures of Streptomyces coelicolor. Appl Environ Microbiol 74:3877–3886
Miguélez EM, Hardisson C, Manzanal MB (1999) Hyphal death during colony development in Streptomyces antibioticus: morphological evidence for the existence of a process of cell deletion in a multicellular prokaryote. J Cell Biol 145:515–525
Nakashio S, Iwasawa H, Dun FY, Kanemitsu K, Shimada J (1995) Everninomicin, a new oligosaccharide antibiotic: its antimicrobial activity, post-antibiotic effect and synergistic bactericidal activity. Drug Exp Clin Res 21:7–16
Ochi K (2007) From microbial differentiation to ribosome engineering. Biosci Biotechnol Biochem 71:1373–1386
Ochi K, Okamoto S, Tozawa Y, Inaoka T, Hosaka T, Xu J, Kurosawa K (2004) Ribosome engineering and secondary metabolite production. Adv Appl Microbiol 56:155–184
Patnaik R, Louie S, Gavrilovic V, Perry K, Stemmer WP, Ryan CM, del Cardayré SB (2002) Genome shuffling of Lactobacillus for improved acid tolerance. Nat Biotechnol 20:707–712
Shima J, Hesketh A, Okamoto S, Kawamoto S, Ochi K (1996) Induction of actinorhodin production by rpsL (encoding ribosomal protein S12) mutations that confer streptomycin resistance in Streptomyces lividans and Streptomyces coelicor A3(2). J Bacteriol 178:7276–7284
Sneath PH, Sokal RR (1973) The principals and practice of numerical classification. Freeman, San Francisco
Stemmer WP (2002) Molecular breeding of genes, pathways and genomes by DNA shuffling. J Mol Catal B: Enzym 19–20:3–12
Tamehiro N, Hosaka T, Xu J, Hu HF, Otake N, Ochi K (2003) Innovative approach for improvement of an antibiotic-overproducing industrial strain of Streptomyces albus. Appl Environ Microbiol 69:6412–6417
Wang G, Hosaka T, Ochi K (2008) Dramatic activation of antibiotic production in Streptomyces coelicolor by cumulative drug resistance mutations. Appl Environ Microbiol 74:2834–2840
Wright DE (1979) The orthosomycins, a new family of antibiotics. Tetrahedron 35:1207–1237
Xu B, Jin ZH, Wang HZ, Jin QC, Jin X, Cen PL (2008) Evolution of Streptomyces pristinaespiralis for resistance and production of pristinamycin by genome shuffling. Appl Microbiol Biotechnol 80:261–267
Yin XH, Zabriskie TM (2006) The enduracidin biosynthetic gene cluster from Streptomyces fungicidicus. Microbiol 152:2969–2983
Zhang YX, Perry K, Vinci VA, Powell K, Stemmer WP, del Cardayré SB (2002) Genome shuffling leads to rapid phenotypic improvement in bacteria. Nature 415:644–646
Zhu CH, Lu FP, He YN, Han ZL, Du LX (2007) Regulation of avilamycin biosynthesis in Streptomyces viridochromogenes: effects of glucose, ammonium ion, and inorganic phosphate. Appl Microbiol Biotechnol 73:1031–1038
Acknowledgments
We thank Dr. Fumin Zhang (Rensselaer Polytechnic Institute, NY, USA) for critical reading of the manuscript. The work was financially supported by the National Nature and Science Foundation of China (3117175) and Nature and Science Foundation of Zhejiang Province (Y3100609).
Conflict of interests
The authors declare that they have no conflict of interests.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 3074 kb)
Rights and permissions
About this article
Cite this article
Lv, XA., Jin, YY., Li, YD. et al. Genome shuffling of Streptomyces viridochromogenes for improved production of avilamycin. Appl Microbiol Biotechnol 97, 641–648 (2013). https://doi.org/10.1007/s00253-012-4322-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-012-4322-7