Abstract
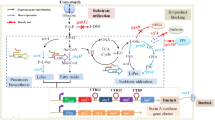
The genes encoding an alcohol dehydrogenase, Baeyer–Villiger monooxygenase and an esterase from P. fluorescens DSM 50106, which seemed to be metabolically connected based on the sequence of the corresponding open reading frames, were cloned into one vector (pABE) and functionally expressed in Escherichia coli. Overall expression levels were quite low, however, using whole cells of E. coli JM109 pABE expressing the three recombinant enzymes, conversion of secondary alcohols (Cn) to the corresponding primary alcohols (Cn−2) and acetic acid via ketone and ester was possible. In this way, 2-decanol was almost completely converted within 20 h at 30°C. Thus, it could be shown that the three enzymes are metabolically connected and that they are most probably involved in alkane degradation via sub-terminal oxidation of the acyclic aliphatic hydrocarbons.





Similar content being viewed by others
References
Abdel-Megeed A (2004) Psychrophilic degradation of long chain alkanes. Dissertation, Technical University Hamburg-Harburg, Germany
Ashraf W, Mihdhir A, Murell JC (1994) Bacterial oxidation of propane. FEMS Microbiol Lett 122:1–6
Atlas RM (1981) Microbial degradation of petroleum hydrocarbons: an environmental perspective. Microbiol Rev 45:180–209
Britton LN (1984) Microbial degradation of aliphatic hydrocarbons. In: Gibson DT (ed) Microbial degradation of organic compounds, vol. 13. Marcel Dekker, New York, pp 89–129
Forney FW, Markovetz AJ (1970) Subterminal oxidation of aliphatic hydrocarbons. J Bacteriol 102:281–282
Hildebrandt P, Musidlowska A, Bornscheuer UT (2002) Cloning, functional expression and biochemical characterization of a stereoselective alcohol dehydrogenase from Pseudomonas fluorescens DSM50106. Appl Microbiol Biotechnol 59:483–487
Hundt K, Wagner M, Becher D, Hammer E, Schauer F (1998) Effect of selected environmental factors on degradation and mineralization of biaryl compounds by the bacterium Ralstonia pickettii in soil and compost. Chemosphere 36:2321–2335
Khalameyzer V, Fischer I, Bornscheuer UT, Altenbuchner J (1999) Screening, nucleotide sequence and biochemical characterization of an esterase from Pseudomonas fluorescens with high activity toward lactones. Appl Environ Microbiol 65:477–482
Kirschner A, Altenbuchner J, Bornscheuer UT (2007) Cloning, expression and characterization of a Baeyer–Villiger monooxygenase from Pseudomonas fluorescens DSM 50106 in E. coli. Appl Microbiol Biotechnol 73:1065–1072
Laemmli UK (1970) Cleavage of structural proteins during assembly of the head of bacteriophage T4. Nature 27:680–685
Leahy JG, Colwell RR (1990) Microbial degradation of hydrocarbons in the environment. Microbiol Rev 54:305–315
Margesin R, Schinner F (1997) Bioremediation of diesel-oil-contaminated alpine soils at low temperatures. Appl Microbiol Biotechnol 47:462–468
Margesin R, Schninner T (2001) Biodegradation and bioremediation of hydrocarbons in extreme environments. Appl Microbiol Biotechnol 56:650–663
Stumpp T, Wilms B, Altenbucher J (2000) Ein neues, l-Rhamnose-induzierbares Expressionssystem für Escherichia coli. Biospektrum 1.00:33–36
Urlacher V, Schmid RD (2006) Recent advances in oxygenase-catalyzed biotransformations. Curr Opin Chem Biol 10:156–161
van Beilen JB, Li Z, Duetz WA, Smits THM, Witholt B (2003) Diversity of alkane hydroxylase systems in the environment. Oil Gas Sci Technol Rev IFP 58:427–440
Whyte LG, Hawari J, Zhou E, Bourbonnière L, Inniss WE, Greer CW (1998) Biodegradation of variable-chain-length alkanes at low temperatures by a psychrophilic Rhodococcus sp. Appl Environ Microbiol 64:2578–2584
Acknowledgment
We thank the Fonds der Chemischen Industrie (Frankfurt, Germany) and the Studienstiftung des deutschen Volkes (Bonn, Germany) for stipends to Anett Kirschner.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kirschner, A., Altenbuchner, J. & Bornscheuer, U.T. Design of a secondary alcohol degradation pathway from Pseudomonas fluorescens DSM 50106 in an engineered Escherichia coli . Appl Microbiol Biotechnol 75, 1095–1101 (2007). https://doi.org/10.1007/s00253-007-0902-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-007-0902-3