Abstract
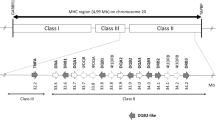
The Major Histocompatibility Complex (MHC) is a multigene family of outstanding polymorphism. MHC molecules bind antigenic peptides in the peptide-binding region (PBR) that consists of five binding pockets (P). In this study, we compared the genetic diversity of domestic pigs to that of the modern representatives of their wild ancestors, the wild boar, in two MHC loci, the oligomorphic DQA and the polymorphic DRB1. MHC nucleotide polymorphism was compared with the actual functional polymorphism in the PBR and the binding pockets P1, P4, P6, P7, and P9. The analysis of approximately 200 wild boars collected throughout Europe and 120 domestic pigs from four breeds (three pureblood, Pietrain, Leicoma, and Landrace, and one mixed Danbred) revealed that wild boars and domestic pigs share the same levels of nucleotide and amino acid polymorphism, allelic richness, and heterozygosity. Domestication did not appear to act as a bottleneck that would narrow MHC diversity. Although the pattern of polymorphism was uniform between the two loci, the magnitude of polymorphism was different. For both loci, most of the polymorphism was located in the PBR region and the presence of positive selection was supported by a statistically significant excess of nonsynonymous substitutions over synonymous substitutions in the PBR. P4 and P6 were the most polymorphic binding pockets. Functional polymorphism, i.e., the number and the distribution of pocket variants within and among populations, was significantly narrower than genetic polymorphism, indicative of a hierarchical action of selection pressures on MHC loci.


Similar content being viewed by others
References
Alexandri P, Triantafyllidis A, Papacostas S, Chatzinikos E, Platis P, Papageorgiou N, Larson G, Abatzopoulos T, Triantaphyllidis C (2012) The Balkans and the colonization of Europe: the post-glacial range expansion of the wild boar, Sus scrofa. J Biogeogr 39:713–732
Alves PC, Pinheiro I, Godinho R, Vocente J, Gortazar C, Scandura M (2010) Genetic diversity of wild boar populations and domestic pig breeds (Sus scrofa) in South-western Europe. Biol J Linn Soc Lon 101:797–822, Science 307(5715):1618–21
Ando A, Chardon P (2006) Gene organization and polymorphism of the swine major histocompatibility complex. Anim Sci J 77:127–137
Apollonio M, Randi E, Toso S (1988) The systematics of the wild boar (Sus scrofa L.) in Italy. Boll Zool 3:213–221
Axtner J, Sommer S (2007) Gene duplication, allelic diversity, selection processes and adaptive value of MHC class II DRB genes of the bank vole, Clethrionomys glareolus. Immunogenetics 5:417–426
Bak EJ, Ishii Y, Omatsu T, Kyuwa S, Hayasaka I, Yoshikawa Y (2005) Sequence analysis of the MHC class II DPB1 gene in chimpanzees (Pan troglodytes). Int J Immunogenet 3:187–192
Barbisan F, Savio C, Bertorelle G, Patarnello T, Congiu L (2009) Duplication polymorphism at MHC class II DRB1 locus in the wild boar (Sus scrofa). Immunogenetics 61:145–151
Beja-Pereira A, Caramelli D, Lalueza-Fox C, Vernesi C, Ferrand N, Casoli A, Goyache F, Royo LJ, Conti S, Lari M, Martini A, Ouragh L, Magid A, Atash A, Zsolnai A, Boscato P, Triantaphylidis C, Ploumi K, Sineo L, Mallegni F, Taberlet P, Erhardt G, Sampietro L, Bertranpetit J, Barbujani G, Luikart G, Bertorelle G (2006) The origin of European cattle: evidence from modern and ancient DNA. Proc Natl Acad Sci U S A 103:8113–8118
Belkhir K (1999) GENETIX, logiciel sous Windows pour la génétique des populations, Laboratoire Génome et Populations, CNRS UPR 9060. Universite´ de Montpellier II, Montpellier (France)
Bernatchez L, Landry C (2003) MHC studies in nonmodel vertebrates: what have we learned about natural selection in 15 years? J Evol Biol 16:363–377
Bondinas GP, Moustakas AK, Papadopoulos GK (2007) The spectrum of HLA-DQ and HLA-DR alleles, 2006: a listing correlating sequence and structure with function. Immunogenetics 59:539–553
Bos DH, Gopurenko D, Williams RN, Dewoody JA (2008) Inferring population history and demography using microsatellites, mitochondrial DNA, and major histocompatibility complex (MHC) genes. Evolution 62:1458–1468
Bowen L, Aldridge BM, Gulland F, Woo J, Van Bonn W, DeLong R, Stott JL, Johnson ML (2002) Molecular characterization of expressed DQA and DQB genes in the California sea lion (Zalophus californianus). Immunogenetics 5:332–347
Brown JH, Jardetzky TS, Gorga JC, Stern LJ, Urban RG, Strominger JL, Wiley DC (1993) Three-dimensional structure of the human class II histocompatibility antigen HLA-DR1. Nature 364:33–39
Cárdenas C, Ortiz M, Balbín A, Villaveces JL, Patarroyo ME (2005) Allele effects in MHC–peptide interactions: a theoretical analysis of HLA-DRβ1*0101-HA and HLADRβ1*0401-HA complexes. Biochem Biophys Res Commun 330:1162–1167
Cárdenas C, Obregón M, Balbín A, Villaveces JL, Patarroyo ME (2007) Wave function analysis of MHC–peptide interactions. J Mol Graph Model 25:605–615
Chardon P, Renard C, Vaiman M (1999) The major histocompatibility complex in swine. Immunol Rev 167:179–192
Chen K, Baxter T, Muir WM, Groenen MA, Schook LB (2007) Genetic resources, genome mapping and evolutionary genomics of the pig (Sus scrofa). Int J Biol Sci 3:153–165
Clarke B, Kirby DR (1966) Maintenance of histocompatibility complex polymorphisms. Nature 211:999–1000
Edwards SV, Hedrick PW (1998) Evolution and ecology of MHC molecules: from genomics to sexual selection. Trends Ecol Evol 13:305–311
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial-DNA restriction data. Genetics 131:479–491
Fain MA, Zhao T, Kindt TJ (2001) Improved typing procedure for the polymorphic single-copy RLA-DQA gene of the rabbit reveals a new allele. Tissue Antigens 57:332–338
Fang M, Larson G, Soares Ribeiro H, Li N, Andersson L (2009) Contrasting mode of evolution at a coat color locus in wild and domestic pigs. PLoS Genet 5:e1000341
Giuffra E, Kijas JM, Amarger V, Carlborg O, Jeon JT, Andersson L (2000) The origin of the domestic pig: independent domestication and subsequent introgression. Genetics 154:1785–1791
Glover KA, Grimholt U, Bakke HG, Nilsen F, Storset A, Skaala O (2007) Major histocompatibility complex (MHC) variation and susceptibility to the sea louse Lepeophtheirus salmonis in Atlantic salmon Salmo salar. Dis Aquat Org 76:57–65
Goudet J (2001) FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). Available at: http://www2.unil.ch/popgen/softwares/fstat.html. Updated from Goudet 1995
Harf R, Sommer S (2005) Association between major histocompatibility complex class II DRB alleles and parasite load in the hairy-footed gerbil, Gerbillurus paeba, in the southern Kalahari. Mol Ecol 14:85–91
Hedrick PW (1999) Balancing selection and MHC. Genetica 104:3207–3214
Hedrick PW, Lee RN, Parker KM (2000) Major histocompatibility complex (MHC) variation in the endangered Mexican wolf and related canids. Heredity 85:617–624
Hedrick PW, Lee RN, Garrigan D (2002) Major histocompatibility complex variation in red wolves: evidence for common ancestry with coyotes and balancing selection. Mol Ecol 10:1905–1913
Hill AVS, Allsopp CEM, Kwiatkowski D, Anstey NM, Twumasi P, Rowe PA, Bennet S, Brewster D, McMichael AJ, Greenwood BM (1991) Common West African HLA antigens are associated with protection from severe malaria. Nature 352:595–600
Ho CS, Rochelle ES, Martens GW, Schook LB, Smith DM (2006) Characterization of swine leukocyte antigen polymorphism by sequence-based and PCR-SSP methods in Meishan pigs. Immunogenetics 58:873–882
Ho CS, Lunney JK, Lee J-H, Franzo-Romain MH, Martens GW, Rowland RRR, Smith DM (2010) Molecular characterization of swine leucocyte antigen class II genes in outbred pig populations. Anim Genet 41:428–432
Hughes AL, Yeager M (1998) Natural selection at major histocompatibility complex loci of vertebrates. Annu Rev Genet 32:415–435
Hughes A, Yeager M, Carrington M (1996) Peptide binding function and the paradox of HLA disease associations. Immunol Cell Biol 74:444–448
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian protein metabolism. Academic, New York, pp 21–123
Kanai TH, Tanioka Y, Tanigawa M, Matsumoto Y, Ueda S, Onodera T, Matsumoto Y (1999) Allelic diversity at class II DRB1 and DQB loci of the pig MHC (SLA). Immunogenetics 50:295–300
Kijas JHM, Wales R, Törnsten A, Chardon P, Mollet M, Andersson L (1998) Melanocortin receptor 1 (MC1R) mutations and coat color in pigs. Genetics 150:1177–1185
Kjǿglum S, Larsen S, Bakke HG, Grimholt U (2006) How specific MHC class I and class II combinations affect disease resistance against infectious salmon anaemia in Atlantic salmon (Salmo salar). Fish Shellfish Immunol 21:431–441
Klein J (1986) Natural history of the major histocompatibility complex. Wiley, New York
Klein J, Sato A, Nikolaidis N (2007) MHC, TSP, and the origin of species: From immunogenetics to evolutionary genetics. Annu Rev Genet 41:281–304
Klein J, Sato A, Nagl S, O'hUigin C (1998) Molecular transspecies polymorphism. Ann Rev Ecol Syst 29:1–21
Köppel C, Knopf L, Ryser M-P, Miserez R, Thür B, Stärk KDC (2007) Serosurveillance for selected infectious disease agents in wild boars (Sus scrofa) and outdoor pigs in Switzerland. Eur J Wildl Res 53:212–220
Koutsogiannouli EA, Moutou KA, Sarafidou T, Stamatis C, Mamuris Z (2009a) Detection of hybrids between wild boars (Sus scrofa scrofa) and domestic pigs (Sus scrofa f. domestica) in Greece, using the PCR-RFLP method on melanocortin-1 receptor (MC1R) mutations. Mamm Biol 75:69–73
Koutsogiannouli EA, Moutou KA, Sarafidou T, Stamatis C, Spyrou V, Mamuris Z (2009b) Major histocompatibility complex variation at class II DQA locus in the brown hare (Lepus europaeus). Mol Ecol 18:4631–4649
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Larson G, Dobney K, Albarella U, Fang M, Matisoo-Smith E, Robins J, Lowden S, Finlayson H, Brand T, Willerslev E, Rowley-Conwy P, Andersson L, Cooper A (2005) Worldwide phylogeography of wild boar reveals multiple centers of pig domestication. Science 307:1618–1621
Lohm J, Grahn M, Langefors A, Andersen O, Storset A, von Schantz T (2002) Experimental evidence for major histocompatibility complex—allele-specific resistance to a bacterial infection. Proc R Soc London Ser B-Biol Sci 269:2029–2033
Luetkemeier ES, Malhi RS, Beever JE, Schook LB (2009) Diversification of porcine MHC class II genes: evidence for selective advantage. Immunogenetics 2:119–129
Massei G, Genov P (2000) Il Cinghiale. Calderini Edagricole, Bologna
McFarland BJ, Beeson C (2002) Binding interactions between peptides and proteins of the class II major histocompatibility complex. Med Res Rev 22:168–203
Meyer D, Thomson G (2001) How selection shapes variation of the human major histocompatibility complex: a review. Ann Hum Genet 65:1–26
Murray BW, Michaud R, Bradley N, White BN (1999) Allelic and haplotype variation of major histocompatibility complex class II DRB1 and DQB loci in the St Lawrence beluga (Delphinapterus leucas). Mol Ecol 8:1127–1139
Musolf K, Meyer-Lucht Y, Sommer S (2004) Evolution of MHC-DRB class II polymorphism in the genus Apodemus and a comparison of DRB sequences within the family Muridae (Mammalia: Rodentia). Immunogenetics 56:420–426
Nei M, Gojobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3(4):18–426
Oppelt C, Wutzler R, von Holst D (2010) Characterisation of MHC class II DRB genes in the northern tree shrew (Tupaia belangeri). Immunogenetics 9:613–622
Otting N, de Groot NG, Doxiadis GGM, Bontrop RE (2002) Extensive Mhc-DQB variation in humans and non-human primate species. Immunogenetics 54:230–239
Ottova E, Simkova A, Martin JF, Goüy de Bellocq J, Gelnara M, Allienne J-F, Morand S (2005) Evolution and trans-species polymorphism of MHC class IIb genes in cyprinid fish. Fish Shellfish Immunol 18:199–222
Paterson S, Wilson K, Pemberton JM (1998) Major histocompatibility complex variation associated with juvenile survival and parasite resistance in a large unmanaged ungulate population (Ovis aries L.). Proc Natl Acad Sci U S A 95:3714–3719
Pokorny I, Sharma R, Goyal SP, Mishra S, Tiedemann R (2010) MHC class I and MHC class II DRB gene variability in wild and captive Bengal tigers (Panthera tigris tigris). Immunogenetics 10:667–679
Potts WK, Manning CJ, Wakeland EK (1991) Mating patterns in seminatural populations of mice influenced by MHC genotype. Nature 352:619–621
Rammensee HG, Fried T, Stevanovic S (1995) MHC ligands and peptide motifs: first listing. Immunogenetics 41:178–228
Scandura M, Iacolina L, Crestanello B, Pecchioli E, Di Benedetto MF, Russo V, Davoli R, Apollonio M, Bertorelle G (2008) Ancient vs. recent processes as factors shaping the genetic variation of the European wild boar: are the effects of the last glaciation still detectable? Mol Ecol 7:1745–1762
Scandura M, Iacolina L, Apollonio M (2011) Genetic diversity in the European wild boar Sus scrofa: phylogeography, population structure and wild X domestic hybridization. Mammal Rev 41:125–137
Schad J, Ganzhorn JU, Sommer S (2005) Parasite burden and constitution of major histocompatibility complex in the Malagasy mouse lemur, Microcebus murinus. Evolution 2:439–450
Seddon JM, Baverstock PR (2000) Evolutionary lineages of RT1.Ba in the Australian Rattus. Mol Biol Evol 17:768–772
Seddon JM, Ellegren H (2002) MHC class II genes in European wolves: a comparison with dogs. Immunogenetics 7:490–500
Slade RW, McCallum HI (1992) Overdominant vs. frequency-dependent selection at MHC loci. Genetics 132:861–864
Smith TP, Rohrer GA, Alexander LJ, Troyer DL, Kirby-Dobbels KR, Janzen MA, Cornwell DL, Louis CF, Schook LB, Beattie CW (1995) Directed integration of the physical and genetic linkage maps of swine chromosome 7 reveals that the SLA spans the centromere. Genome Res 5:259–271
Smith TP, Lunney JK, Ho CS, Martens GW, Ando A, Lee JH, Schook L, Renard C, Chardon P (2005) Nomenclature for factors of the swine leucocyte antigen class II system. Tissue Antigens 66:623–639
Surridge AK, van der Loo W, Abrentes J, Carneiro M, Hewitt GM, Esteves PJ (2008) Diversity and evolutionary history of the MHC DQA gene in leporids. Immunogenetics 60:515–525
Takahata N (1993) Allelic genealogy and human evolution. Mol Biol Evol 10:2–22
Takahata N, Nei M (1990) Allelic genealogy under overdominant and frequency-dependent selection and polymorphism of major histocompatibility complex loci. Genetics 124:967–978
Takahata N, Satta Y (1997) Evolution of the primate lineage leading to modern humans: phylogenetic and demographic inferences from DNA sequences. Proc Natl Acad Sci U S A 94:4811–4815
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tennessen JA (2005) Molecular evolution of animal antimicrobial peptides: widespread moderate positive selection. J Evol Biol 18:1387–1394
Thong LM, Choi H, Kwon O-J, Kim J-H, Kim Y-B, Oh J-W, Seo K, Yeom S-C, Lee W-J, Park C (2011) Systematic analysis of swine leucocyte antigen-DRB1 nucleotide polymorphisms using genomic DNA-based high-resolution genotyping and identification of new alleles. Tissue Antigens 77:572–583
Van Eijk MJT, Stewart-Haynes JA, Beever JE, Fernando RL, Lewin HA (1992) Development of persistent lymphocytosis in cattle in closely associated with DRB2. Immunogenetics 37:64–68
Vernesi C, Crestanello B, Pecchioli E, Tartari D, Caramelli D, Hauffe H, Bertorelle G (2003) The genetic impact of demographic decline and reintroduction in the wild boar (Sus scrofa): a microsatellite analysis. Mol Ecol 3:585–595
Vilà C, Seddon J, Ellegren H (2005) Genes of domestic mammals augmented by backcrossing with wild ancestors. Trends Genet 21:214–218
Xu A, Van Eijk MJT, Park C, Lewin HA (1993) Polymorphism in BoLA-DRB3 exon 2 correlates with resistance to persistent lymphocytosis caused by bovine leukaemia virus. J Immunol 151:6977–6985
Yamazaki K, Boyse EA, Mike V, Thaler HT, Mathieson BJ, Abbott J, Boyse J, Zayas ZA, Thomas L (1976) Control of mating preferences in mice by genes in the major histocompatibility complex. J Exp Med 114:1324–1335
Acknowledgments
The authors would like to thank the hunting associations for their help in collecting samples from all over Greece as well as Dr. Chak-Sum (Sam) Ho, Chair of ISAG/IUIS-VIC SLA Nomenclature Committee for his valuable suggestions in allele assignment. Laboratory work was financed by the Postgraduate Courses “Biotechnology—Quality Assessment in Nutrition and the Environment” and “Applications of Molecular Biology—Genetics—Diagnostic Biomarkers” of the Department of Biochemistry and Biotechnology, University of Thessaly, Greece.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Moutou, K.A., Koutsogiannouli, E.A., Stamatis, C. et al. Domestication does not narrow MHC diversity in Sus scrofa . Immunogenetics 65, 195–209 (2013). https://doi.org/10.1007/s00251-012-0671-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-012-0671-8